Biology:Delta-aminolevulinic acid dehydratase
 Generic protein structure example |
| porphobilinogen synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 DALA dehydratase | |||||||||
| Identifiers | |||||||||
| EC number | 4.2.1.24 | ||||||||
| CAS number | 9036-37-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Delta-aminolevulinic acid dehydratase | |
|---|---|
| Identifiers | |
| Symbol | ALAD |
| NCBI gene | 210 |
| HGNC | 395 |
| OMIM | 125270 |
| RefSeq | NM_001003945 |
| UniProt | P13716 |
| Other data | |
| EC number | 4.2.1.24 |
| Locus | Chr. 9 q32 |
| ALAD | |||||||||
|---|---|---|---|---|---|---|---|---|---|
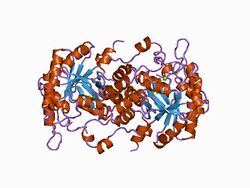 high resolution crystal structure of a mg2-dependent 5-aminolevulinic acid dehydratase | |||||||||
| Identifiers | |||||||||
| Symbol | ALAD | ||||||||
| Pfam | PF00490 | ||||||||
| Pfam clan | CL0036 | ||||||||
| InterPro | IPR001731 | ||||||||
| PROSITE | PDOC00153 | ||||||||
| SCOP2 | 1aw5 / SCOPe / SUPFAM | ||||||||
| |||||||||
Aminolevulinic acid dehydratase (porphobilinogen synthase, or ALA dehydratase, or aminolevulinate dehydratase) is an enzyme (EC 4.2.1.24) that in humans is encoded by the ALAD gene.[1][2] Porphobilinogen synthase (or ALA dehydratase, or aminolevulinate dehydratase) synthesizes porphobilinogen through the asymmetric condensation of two molecules of aminolevulinic acid. All natural tetrapyrroles, including hemes, chlorophylls and vitamin B12, share porphobilinogen as a common precursor. Porphobilinogen synthase is the prototype morpheein.[3]
Function
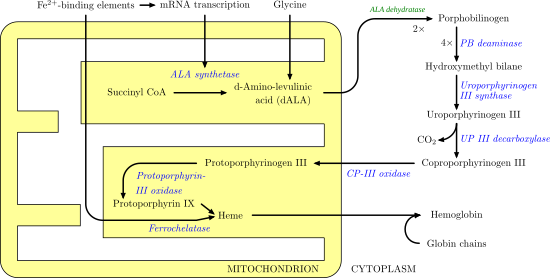
It catalyzes the following reaction, the second step of the biosynthesis of porphyrin:
- 2 5-Aminolevulinic acid porphobilinogen + 2 H2O
It therefore catalyzes the condensation of 2 molecules of 5-aminolevulinate to form porphobilinogen (a precursor of heme, cytochromes and other hemoproteins). This reaction is the first common step in the biosynthesis of all biological tetrapyrroles. Zinc is essential for enzymatic activity.
Structure
The structural basis for allosteric regulation of Porphobilinogen synthase (PBGS) is modulation of a quaternary structure equilibrium between octamer and hexamer (via dimers), which is represented schematically as 6mer* ↔ 2mer* ↔ 2mer ↔ 8mer. The * represents a reorientation between two domains of each subunit that occurs in the dissociated state because it is sterically forbidden in the larger multimers.[3]
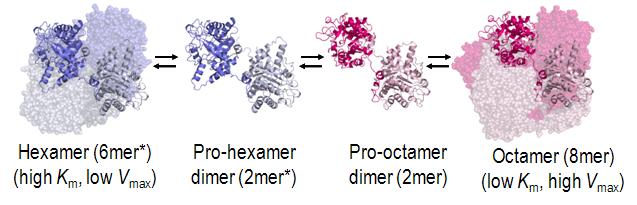
PBGS is encoded by a single gene and each PBGS multimer is composed of multiple copies of the same protein. Each PBGS subunit consists of a ~300 residue αβ-barrel domain, which houses the enzyme's active site in its center, and a >25 residue N-terminal arm domain. Allosteric regulation of PBGS can be described in terms of the orientation of the αβ-barrel domain with respect to the N-terminal arm domain.
Each N-terminal arm has up to two interactions with other subunits in a PBGS multimer. One of these interactions helps to stabilize a "closed" conformation of the active site lid. The other interaction restricts solvent access from the other end of the αβ-barrel.
In the inactive multimeric state, the N-terminal arm domain is not involved in the lid-stabilizing interaction, and in the crystal structure of the inactive assembly, the active site lid is disordered.
Allosteric regulators
As a nearly universal enzyme with a highly conserved active site, PBGS would not be a prime target for the development of antimicrobials and/or herbicides. To the contrary, allosteric sites can be much more phylogenetically variable than active sites, thus presenting more drug development opportunities.[3]
Phylogenetic variation in PBGS allostery leads to the framing of discussion of PBGS allosteric regulation in terms of intrinsic and extrinsic factors.
Intrinsic allosteric regulators
Magnesium
The allosteric magnesium ion lies at the highly hydrated interface of two pro-octamer dimers. It appears to be easily dissociable, and it has been shown that hexamers accumulate when magnesium is removed in vitro.[4]
pH
Though it is not common to consider hydronium ions as allosteric regulators, in the case of PBGS, side chain protonation at locations other than the active site has been shown to affect the quaternary structure equilibrium, and thus to affect the rate of its catalyzed reaction as well.
Extrinsic allosteric regulators
Small molecule hexamer stabilization
Inspection of the PBGS 6mer* reveals a surface cavity that is not present in the 8mer. Small molecule binding to this phylogenetically variable cavity has been proposed to stabilize 6mer* of the targeted PBGS and consequently inhibit activity.
Such allosteric regulators are known as morphlocks because they lock PBGS in a specific morpheein form (6mer*).[5]
Lead poisoning
ALAD enzymatic activity is inhibited by lead, beginning at blood lead levels that were once considered to be safe (<10 μg/dL) and continuing to correlate negatively across the range from 5 to 95 μg/dL.[6] Inhibition of ALAD by lead leads to anemia primarily because it both inhibits heme synthesis and shortens the lifespan of circulating red blood cells, but also by stimulating the excessive production of the hormone erythropoietin, leading to inadequate maturation of red cells from their progenitors. A defect in the ALAD structural gene can cause increased sensitivity to lead poisoning and acute hepatic porphyria. Alternatively spliced transcript variants encoding different isoforms have been identified.[7]
Deficiency
A deficiency of porphobilinogen synthase is usually acquired (rather than hereditary) and can be caused by heavy metal poisoning, especially lead poisoning, as the enzyme is very susceptible to inhibition by heavy metals.[8]
Hereditary insufficiency of porphobilinogen synthase is called porphobilinogen synthase (or ALA dehydratase) deficiency poprhyria. It is an extremely rare cause of porphyria,[9] with less than 10 cases ever reported.[10] All disease associated protein variants favor hexamer formation relative to the wild type human enzyme.[9]
 |
PBGS as the prototype morpheein
The morpheein model of allostery exemplified by PBGS adds an additional layer of understanding to potential mechanisms for regulation of protein function and complements the increased focus that the protein science community is placing on protein structure dynamics.[3]
This model illustrates how the dynamics of phenomena such as alternate protein conformations, alternate oligomeric states, and transient protein-protein interactions can be harnessed for allosteric regulation of catalytic activity.
References
- ↑ "delta-Aminolevulinatedehydrase: synteny with ABO-AK1-ORM (and assignment to chromosome 9)". Clinical Genetics 23 (2): 150–4. February 1983. doi:10.1111/j.1399-0004.1983.tb01864.x. PMID 6839527.
- ↑ "Assignment of the human gene for delta aminolevulinate dehydrase to chromosome 9 by somatic cell hybridization and specific enzyme immunoassay". Annals of Human Genetics 48 (2): 153–9. May 1984. doi:10.1111/j.1469-1809.1984.tb01010.x. PMID 6378062.
- ↑ 3.0 3.1 3.2 3.3 "Allostery and the dynamic oligomerization of porphobilinogen synthase". Archives of Biochemistry and Biophysics 519 (2): 144–53. March 2012. doi:10.1016/j.abb.2011.10.010. PMID 22037356.
- ↑ "Control of tetrapyrrole biosynthesis by alternate quaternary forms of porphobilinogen synthase". Nature Structural Biology 10 (9): 757–63. September 2003. doi:10.1038/nsb963. PMID 12897770.
- ↑ "Expanding the Concepts in Protein Structure-Function Relationships and Enzyme Kinetics: Teaching using Morpheeins". Biochemistry and Molecular Biology Education 36 (4): 274–283. 2008. doi:10.1002/bmb.20211. PMID 19578473.
- ↑ Toxicological Profile for Lead. Atlanta, GA: Agency for Toxic Substances and Disease Registry (US). August 2007. pp. 22, 30. http://www.atsdr.cdc.gov/toxprofiles/tp13.pdf. Retrieved 22 November 2015.
- ↑ "Entrez Gene: ALAD aminolevulinate, delta-, dehydratase". https://www.ncbi.nlm.nih.gov/sites/entrez?Db=gene&Cmd=ShowDetailView&TermToSearch=210.
- ↑ ALA dehydratase reaction, from NetBiochem at the University of Utah. Last modified 1/5/95
- ↑ 9.0 9.1 "ALAD porphyria is a conformational disease". American Journal of Human Genetics 80 (2): 329–37. February 2007. doi:10.1086/511444. PMID 17236137.
- ↑ Overview of the Porphyrias at The Porphyrias Consortium (a part of NIH Rare Diseases Clinical Research Network (RDCRN)) Retrieved June 2011
External links
- Human ALAD genome location and ALAD gene details page in the UCSC Genome Browser.
- delta-Aminolevulinic+Acid+Dehydratase at the US National Library of Medicine Medical Subject Headings (MeSH)
- http://www.omim.org/entry/125270?search=pbgs&highlight=pbgs
Further reading
- "Metal-induced alterations of delta-aminolevulinic acid dehydratase". Annals of the New York Academy of Sciences 514: 41–7. 1988. doi:10.1111/j.1749-6632.1987.tb48759.x. PMID 3327436.
- "The porphobilinogen synthase catalyzed reaction mechanism". Bioorganic Chemistry 32 (5): 316–25. October 2004. doi:10.1016/j.bioorg.2004.05.010. PMID 15381398.
- "Comparison of in vivo effect of inorganic lead and cadmium on glutathione reductase system and delta-aminolevulinate dehydratase in human erythrocytes". British Journal of Industrial Medicine 32 (3): 181–92. August 1975. doi:10.1136/oem.32.3.181. PMID 1156566.
- "Cloning and expression of the defective genes from a patient with delta-aminolevulinate dehydratase porphyria". The Journal of Clinical Investigation 89 (5): 1431–7. May 1992. doi:10.1172/JCI115732. PMID 1569184.
- "Treatment of Haemophilus aphrophilus endocarditis with ciprofloxacin". The Journal of Infection 24 (3): 317–20. May 1992. doi:10.1016/S0163-4453(05)80037-4. PMID 1602151.
- "RsaI polymorphism in the human delta-aminolevulinate dehydratase gene at 9q34". Nucleic Acids Research 19 (15): 4307. August 1991. doi:10.1093/nar/19.15.4307-a. PMID 1678509.
- "Molecular characterization of the human delta-aminolevulinate dehydratase 2 (ALAD2) allele: implications for molecular screening of individuals for genetic susceptibility to lead poisoning". American Journal of Human Genetics 49 (4): 757–63. October 1991. PMID 1716854.
- "delta-Aminolevulinate dehydratase deficient porphyria: identification of the molecular lesions in a severely affected homozygote". American Journal of Human Genetics 49 (1): 167–74. July 1991. PMID 2063868.
- "Human delta-aminolevulinate dehydratase: chromosomal localization to 9q34 by in situ hybridization". Human Genetics 76 (3): 236–9. July 1987. doi:10.1007/BF00283614. PMID 3036687.
- "Identification of lysine at the active site of human 5-aminolaevulinate dehydratase". The Biochemical Journal 236 (2): 447–51. June 1986. doi:10.1042/bj2360447. PMID 3092810.
- "Human delta-aminolevulinate dehydratase: nucleotide sequence of a full-length cDNA clone". Proceedings of the National Academy of Sciences of the United States of America 83 (20): 7703–7. October 1986. doi:10.1073/pnas.83.20.7703. PMID 3463993. Bibcode: 1986PNAS...83.7703W.
- "Molecular cloning of a cDNA for human delta-aminolevulinate dehydratase". Gene 43 (1–2): 123–30. 1986. doi:10.1016/0378-1119(86)90015-6. PMID 3758678.
- "Acute hepatic porphyria syndrome with porphobilinogen synthase defect". The International Journal of Biochemistry 12 (5–6): 823–6. 1981. doi:10.1016/0020-711X(80)90170-6. PMID 7450139.
- "Human delta-aminolevulinate dehydratase (ALAD) gene: structure and alternative splicing of the erythroid and housekeeping mRNAs". Genomics 19 (2): 242–8. January 1994. doi:10.1006/geno.1994.1054. PMID 8188255.
- "A novel mutation of delta-aminolaevulinate dehydratase in a healthy child with 12% erythrocyte enzyme activity". British Journal of Haematology 106 (4): 931–7. September 1999. doi:10.1046/j.1365-2141.1999.01647.x. PMID 10519994.
- "Novel molecular defects of the delta-aminolevulinate dehydratase gene in a patient with inherited acute hepatic porphyria". Hepatology 31 (3): 704–8. March 2000. doi:10.1002/hep.510310321. PMID 10706561.
- "Mechanistic basis for suicide inactivation of porphobilinogen synthase by 4,7-dioxosebacic acid, an inhibitor that shows dramatic species selectivity". Biochemistry 40 (28): 8227–36. July 2001. doi:10.1021/bi010656k. PMID 11444968.
 |



