Biology:Enolase
| phosphopyruvate hydratase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
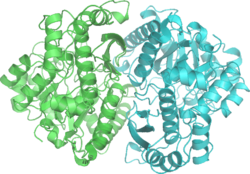 Yeast enolase dimer.[1] | |||||||||
| Identifiers | |||||||||
| EC number | 4.2.1.11 | ||||||||
| CAS number | 9014-08-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Enolase, N-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
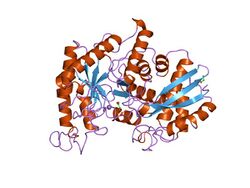 x-ray structure and catalytic mechanism of lobster enolase | |||||||||
| Identifiers | |||||||||
| Symbol | Enolase_N | ||||||||
| Pfam | PF03952 | ||||||||
| Pfam clan | CL0227 | ||||||||
| InterPro | IPR020811 | ||||||||
| PROSITE | PDOC00148 | ||||||||
| SCOP2 | 1els / SCOPe / SUPFAM | ||||||||
| |||||||||
| Enolase | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Enolase | ||||||||||
| Pfam | PF00113 | ||||||||||
| InterPro | IPR000941 | ||||||||||
| PROSITE | PDOC00148 | ||||||||||
| |||||||||||
Phosphopyruvate hydratase, usually known as enolase, is a metalloenzyme (EC 4.2.1.11) that catalyses the conversion of 2-phosphoglycerate (2-PG) to phosphoenolpyruvate (PEP), the ninth and penultimate step of glycolysis. The chemical reaction is:
- 2-phospho-D-glycerate phosphoenolpyruvate + H2O
Phosphopyruvate hydratase belongs to the family of lyases, specifically the hydro-lyases, which cleave carbon-oxygen bonds. The systematic name of this enzyme is 2-phospho-D-glycerate hydro-lyase (phosphoenolpyruvate-forming).
The reaction is reversible, depending on environmental concentrations of substrates.[3] The optimum pH for the human enzyme is 6.5.[4] Enolase is present in all tissues and organisms capable of glycolysis or fermentation. The enzyme was discovered by Lohmann and Meyerhof in 1934,[5] and has since been isolated from a variety of sources including human muscle and erythrocytes.[4] In humans, deficiency of ENO1 is linked to hereditary haemolytic anemia, while ENO3 deficiency is linked to glycogen storage disease type XIII.
Isozymes
In humans there are three subunits of enolase, α, β, and γ, each encoded by a separate gene that can combine to form five different isoenzymes: αα, αβ, αγ, ββ, and γγ.[3][6] Three of these isoenzymes (all homodimers) are more commonly found in adult human cells than the others:
- αα or non-neuronal enolase (NNE). Also known as enolase 1. Found in a variety of tissues, including liver, brain, kidney, spleen, adipose. It is present at some level in all normal human cells.
- ββ or muscle-specific enolase (MSE). Also known as enolase 3. This enzyme is largely restricted to muscle where it is present at very high levels in muscle.
- γγ or neuron-specific enolase (NSE). Also known as enolase 2. Expressed at very high levels in neurons and neural tissues, where it can account for as much as 3% of total soluble protein. It is expressed at much lower levels in most mammalian cells.
When present in the same cell, different isozymes readily form heterodimers.[citation needed]
Structure
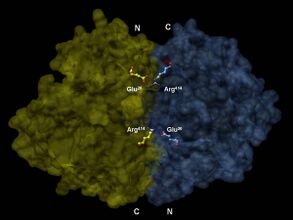
Enolase is a member of the large enolase superfamily. It has a molecular weight of 82,000–100,000 daltons depending on the isoform.[3][4] In human alpha enolase, the two subunits are antiparallel in orientation so that Glu20 of one subunit forms an ionic bond with Arg414 of the other subunit.[3] Each subunit has two distinct domains. The smaller N-terminal domain consists of three α-helices and four β-sheets.[3][6] The larger C-terminal domain starts with two β-sheets followed by two α-helices and ends with a barrel composed of alternating β-sheets and α-helices arranged so that the β-beta sheets are surrounded by the α-helices.[3][6] The enzyme's compact, globular structure results from significant hydrophobic interactions between these two domains.
Enolase is a highly conserved enzyme with five active-site residues being especially important for activity. When compared to wild-type enolase, a mutant enolase that differs at either the Glu168, Glu211, Lys345, or Lys396 residue has an activity level that is cut by a factor of 105.[3] Also, changes affecting His159 leave the mutant with only 0.01% of its catalytic activity.[3] An integral part of enolase are two Mg2+ cofactors in the active site, which serve to stabilize negative charges in the substrate.[3][6]
Recently, moonlighting functions of several enolases, such as interaction with plasminogen, have brought interest to the enzymes' catalytic loops and their structural diversity.[7][8]
-
3-D depiction of enolase dimer in antiparallel orientation. One dimer’s N-terminal Glu20 forms an ionic bond with the other’s C-terminal Arg414 to stabilize the enzyme’s quaternary structure.
-
Active site of enolase in the middle of the C-terminal domain’s barrel. Depicted are two Mg2+ cofactors and five highly conserved residues imperative for proper catalytic function: His159, Glu168, Glu211, Lys345, Lys396.
Mechanism
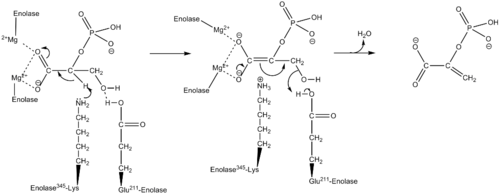
Using isotopic probes, the overall mechanism for converting 2-PG to PEP is proposed to be an E1cB elimination reaction involving a carbanion intermediate.[9] The following detailed mechanism is based on studies of crystal structure and kinetics.[3][10][11][12][13][14][15] When the substrate, 2-phosphoglycerate, binds to α-enolase, its carboxyl group coordinates with two magnesium ion cofactors in the active site. This stabilizes the negative charge on the deprotonated oxygen while increasing the acidity of the alpha hydrogen. Enolase's Lys345 deprotonates the alpha hydrogen, and the resulting negative charge is stabilized by resonance to the carboxylate oxygen and by the magnesium ion cofactors. Following the creation of the carbanion intermediate, the hydroxide on C3 is eliminated as water with the help of Glu211, and PEP is formed.
Additionally, conformational changes occur within the enzyme that aid catalysis. In human α-enolase, the substrate is rotated into position upon binding to the enzyme due to interactions with the two catalytic magnesium ions, Gln167, and Lys396. Movements of loops Ser36 to His43, Ser158 to Gly162, and Asp255 to Asn256 allow Ser39 to coordinate with Mg2+ and close off the active site. In addition to coordination with the catalytic magnesium ions, the pKa of the substrate's alpha hydrogen is also lowered due to protonation of the phosphoryl group by His159 and its proximity to Arg374. Arg374 also causes Lys345 in the active site to become deprotonated, which primes Lys345 for its role in the mechanism.
Diagnostic uses
In recent medical experiments, enolase concentrations have been sampled in an attempt to diagnose certain conditions and their severity. For example, higher concentrations of enolase in cerebrospinal fluid more strongly correlated to low-grade astrocytoma than did other enzymes tested (aldolase, pyruvate kinase, creatine kinase, and lactate dehydrogenase).[16] The same study showed that the fastest rate of tumor growth occurred in patients with the highest levels of CSF enolase. Increased levels of enolase have also been identified in patients who have suffered a recent myocardial infarction or cerebrovascular accident. It has been inferred that levels of CSF neuron-specific enolase, serum NSE, and creatine kinase (type BB) are indicative in the prognostic assessment of cardiac arrest victims.[17] Other studies have focused on the prognostic value of NSE values in cerebrovascular accident victims.[18]
Autoantibodies to alpha-enolase are associated with rheumatoid arthritis[19] and the rare syndrome called Hashimoto's encephalopathy.[20]
Inhibitors
Small-molecule inhibitors of enolase have been synthesized as chemical probes (substrate-analogues) of the catalytic mechanism of the enzyme and more recently, have been investigated as potential treatments for cancer and infectious diseases.[21][22] Most inhibitors have metal chelating properties and bind to enzyme by interactions with the structural Magnesium Atom Mg(A).[23][24] The most potent of these is phosphonoacetohydroxamate,[24] which in its unprotonated form has pM affinity for the enzyme. It has structural similarity to the presumed catalytic intermediate, between PEP and 2-PG. Attempts have been made to use this inhibitor as an anti-trypanosome drug,[25] and more recently, as an anti-cancer agent, specifically, in glioblastoma that are enolase-deficient due to homozygous deletion of the ENO1 gene as part of the 1p36 tumor suppressor locus (synthetic lethality).[26] A natural product phosphonate antibiotic, SF2312 (CAS 107729-45-3), which is active against gram positive and negative bacteria especially under anaerobic conditions,[27] is a high potency inhibitor of Enolase 4zcw that binds in manner similar to phoshphonoacetohydroxamate 4za0.[28] SF2312 inhibits Enolase activity in both eukaryotic and prokaryotic origin,[29] reflecting the strong evolutionary conservation of Enolase and the ancient origin of the glycolysis pathway. SF2312 is a chiral molecule with only the 3S-enantiomer showing Enolase inhibitory activity and biological activity against bacteria.[30] More recently, a derivative of SF2312, termed HEX, and a prodrug thereoff, POMHEX, were shown to exert anti-neoplastic activity against ENO1-deleted glioma in a pre-clinical intracranial orthotopic mouse model.[31] An allosteric binder, ENOblock[22] was initially described as an inhibitor of Enolase, but subsequently shown not to actually inhibit the enzyme, but rather, interfere with the Enolase in vitro enzymatic assay.[32] ENOblock was found to alter the cellular localization of enolase, influencing its secondary, non-glycolytic functions, such as transcription regulation.[33] Subsequent analysis using a commercial assay also indicated that ENOblock can inhibit enolase activity in biological contexts, such as cells and animal tissues.[33] Methylglyoxal has also been described as an inhibitor of human enolase.[34]
Active site transition state analogue Enolase inhibitors have been explored pre-clinically for the treatment of various microbial pathogens, as well as in precision oncology for tumors with 1p36 homozygous deletions, that lack ENO1.[31][35][36][37][38][39][40]
Fluoride is a known competitor of enolase's substrate 2-PG. Fluoride can form a complex with magnesium and phosphate, which binds in the active site instead of 2-PG.[4] One study found that fluoride could inhibit bacterial enolase in vitro.[41] The Enolase inhibitory activity of Fluoride anion may contribute to the anti-cavity effect of fluoride toothpaste, by limiting lactic acid (a product of glycolysis, which requires Enolase) production.
References
- ↑ PDB: 2ONE; "Mechanism of enolase: the crystal structure of asymmetric dimer enolase-2-phospho-D-glycerate/enolase-phosphoenolpyruvate at 2.0 A resolution". Biochemistry 36 (41): 12526–12534. October 1997. doi:10.1021/bi9712450. PMID 9376357.
- ↑ PDB: 2XSX; "Crystal structure of human beta enolase ENOB". Protein Data Bank. 2010.
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 "Multifunctional alpha-enolase: its role in diseases". Cellular and Molecular Life Sciences 58 (7): 902–920. June 2001. doi:10.1007/pl00000910. PMID 11497239.
- ↑ 4.0 4.1 4.2 4.3 "Purification and properties of enolase of human erythrocytes". International Journal of Biochemistry 5 (11–12): 845–852. November 1974. doi:10.1016/0020-711X(74)90119-0.
- ↑ Lohman, K; Meyerhof, O (1934). "Über die enzymatische umwandlung von phosphoglyzerinsäure in brenztraubensäure und phosphorsäure" (in de). Biochemische Zeitschrift 273: 60–72.
- ↑ 6.0 6.1 6.2 6.3 "Molecular structure of the human muscle-specific enolase gene (ENO3)". The Biochemical Journal 275 ( Pt 2) (Pt 2): 427–433. April 1991. doi:10.1042/bj2750427. PMID 1840492.
- ↑ "Plasmin(ogen)-binding alpha-enolase from Streptococcus pneumoniae: crystal structure and evaluation of plasmin(ogen)-binding sites". Journal of Molecular Biology 343 (4): 997–1005. October 2004. doi:10.1016/j.jmb.2004.08.088. PMID 15476816.
- ↑ "Crystal structure of an efficacious gonococcal adherence inhibitor: an enolase from Lactobacillus gasseri". FEBS Letters 588 (14): 2212–2216. June 2014. doi:10.1016/j.febslet.2014.05.020. PMID 24859038.
- ↑ "Isotopic probes of the enolase reaction mechanism". J Biol Chem 240 (14): 4586–93. 1971. doi:10.1016/S0021-9258(18)62051-4.
- ↑ "Toward identification of acid/base catalysts in the active site of enolase: comparison of the properties of K345A, E168Q, and E211Q variants". Biochemistry 35 (5): 1692–1699. February 1996. doi:10.1021/bi952186y. PMID 8634301.
- ↑ "Structural and mechanistic studies of enolase". Current Opinion in Structural Biology 6 (6): 736–743. December 1996. doi:10.1016/S0959-440X(96)80002-9. PMID 8994873.
- ↑ "Octahedral coordination at the high-affinity metal site in enolase: crystallographic analysis of the MgII--enzyme complex from yeast at 1.9 A resolution". Biochemistry 34 (13): 4325–4330. April 1995. doi:10.1021/bi00013a022. PMID 7703246.
- ↑ "Chelation of serine 39 to Mg2+ latches a gate at the active site of enolase: structure of the bis(Mg2+) complex of yeast enolase and the intermediate analog phosphonoacetohydroxamate at 2.1-A resolution". Biochemistry 33 (31): 9333–9342. August 1994. doi:10.1021/bi00197a038. PMID 8049235.
- ↑ "A carboxylate oxygen of the substrate bridges the magnesium ions at the active site of enolase: structure of the yeast enzyme complexed with the equilibrium mixture of 2-phosphoglycerate and phosphoenolpyruvate at 1.8 A resolution". Biochemistry 35 (14): 4349–4358. April 1996. doi:10.1021/bi952859c. PMID 8605183.
- ↑ "X-ray structure and catalytic mechanism of lobster enolase". Biochemistry 34 (39): 12513–12523. October 1995. doi:10.1021/bi00039a005. PMID 7547999.
- ↑ "Levels of enolase and other enzymes in the cerebrospinal fluid as indices of pathological change". Journal of Neurology, Neurosurgery, and Psychiatry 44 (12): 1129–1135. December 1981. doi:10.1136/jnnp.44.12.1129. PMID 7334408.
- ↑ "Neurological outcome after out-of-hospital cardiac arrest. Prediction by cerebrospinal fluid enzyme analysis". Archives of Neurology 46 (7): 753–756. July 1989. doi:10.1001/archneur.1989.00520430047015. PMID 2742544.
- ↑ "Cerebrospinal fluid enolase in stroke". Journal of Neurology, Neurosurgery, and Psychiatry 47 (7): 724–729. July 1984. doi:10.1136/jnnp.47.7.724. PMID 6747647.
- ↑ "Antibodies to citrullinated alpha-enolase peptide 1 are specific for rheumatoid arthritis and cross-react with bacterial enolase". Arthritis and Rheumatism 58 (10): 3009–3019. October 2008. doi:10.1002/art.23936. PMID 18821669.
- ↑ "Autoantibodies against the amino terminal of alpha-enolase are a useful diagnostic marker of Hashimoto's encephalopathy". Journal of Neuroimmunology 162 (1–2): 130–136. May 2005. doi:10.1016/j.jneuroim.2005.02.004. PMID 15833368.
- ↑ "Reaction intermediate analogues for enolase". Biochemistry 23 (12): 2779–2786. June 1984. doi:10.1021/bi00307a038. PMID 6380574.
- ↑ 22.0 22.1 "A unique small molecule inhibitor of enolase clarifies its role in fundamental biological processes". ACS Chemical Biology 8 (6): 1271–1282. 2 April 2013. doi:10.1021/cb300687k. PMID 23547795.
- ↑ "Structure of the bis divalent cation complex with phosphonoacetohydroxamate at the active site of enolase". Biochemistry 31 (31): 7166–7173. August 1992. doi:10.1021/bi00146a020. PMID 1322695.
- ↑ 24.0 24.1 "Catalytic metal ion binding in enolase: the crystal structure of an enolase-Mn2+-phosphonoacetohydroxamate complex at 2.4-A resolution". Biochemistry 33 (20): 6295–6300. May 1994. doi:10.1021/bi00186a032. PMID 8193144.
- ↑ "Passenger deletions generate therapeutic vulnerabilities in cancer". Nature 488 (7411): 337–342. August 2012. doi:10.1038/nature11331. PMID 22895339. Bibcode: 2012Natur.488..337M.
- ↑ "Studies on a new phosphonic acid antibiotic, SF-2312.". Sci Rep Meiji Seika Kaisha. 25: 12–17. 1986.
- ↑ "SF2312 is a natural phosphonate inhibitor of enolase". Nature Chemical Biology 12 (12): 1053–1058. December 2016. doi:10.1038/nchembio.2195. PMID 27723749.
- ↑ "Functional and structural basis of E. coli enolase inhibition by SF2312: a mimic of the carbanion intermediate". Scientific Reports 9 (1): 17106. November 2019. doi:10.1038/s41598-019-53301-3. PMID 31745118. Bibcode: 2019NatSR...917106K.
- ↑ "The 3S Enantiomer Drives Enolase Inhibitory Activity in SF2312 and Its Analogues". Molecules 24 (13): 2510. July 2019. doi:10.3390/molecules24132510. PMID 31324042.
- ↑ 31.0 31.1 "An enolase inhibitor for the targeted treatment of ENO1-deleted cancers". Nature Metabolism 2 (12): 1413–1426. December 2020. doi:10.1038/s42255-020-00313-3. PMID 33230295.
- ↑ "ENOblock Does Not Inhibit the Activity of the Glycolytic Enzyme Enolase". PLOS ONE 11 (12): e0168739. 28 December 2016. doi:10.1371/journal.pone.0168739. PMID 28030597. Bibcode: 2016PLoSO..1168739S.
- ↑ 33.0 33.1 "ENOblock, a unique small molecule inhibitor of the non-glycolytic functions of enolase, alleviates the symptoms of type 2 diabetes". Scientific Reports 7: 44186. March 2017. doi:10.1038/srep44186. PMID 28272459. Bibcode: 2017NatSR...744186C.
- ↑ "Inhibition of human muscle-specific enolase by methylglyoxal and irreversible formation of advanced glycation end products". Journal of Enzyme Inhibition and Medicinal Chemistry 24 (2): 356–364. April 2009. doi:10.1080/14756360802187679. PMID 18830874.
- ↑ "Targeting Host Glycolysis as a Strategy for Antimalarial Development". Frontiers in Cellular and Infection Microbiology 11: 730413. 16 September 2021. doi:10.3389/fcimb.2021.730413. PMID 34604112.
- ↑ "Structure-guided microbial targeting of antistaphylococcal prodrugs". eLife 10: e66657. July 2021. doi:10.7554/eLife.66657. PMID 34279224.
- ↑ "Antimicrobial Prodrug Activation by the Staphylococcal Glyoxalase GloB". ACS Infectious Diseases 6 (11): 3064–3075. November 2020. doi:10.1021/acsinfecdis.0c00582. PMID 33118347.
- ↑ "Enhancer Reprogramming Confers Dependence on Glycolysis and IGF Signaling in KMT2D Mutant Melanoma". Cell Reports 33 (3): 108293. October 2020. doi:10.1016/j.celrep.2020.108293. PMID 33086062.
- ↑ "Functional and structural basis of E. coli enolase inhibition by SF2312: a mimic of the carbanion intermediate". Scientific Reports 9 (1): 17106. November 2019. doi:10.1038/s41598-019-53301-3. PMID 31745118. Bibcode: 2019NatSR...917106K.
- ↑ "The 3S Enantiomer Drives Enolase Inhibitory Activity in SF2312 and Its Analogues". Molecules 24 (13): 2510. July 2019. doi:10.3390/molecules24132510. PMID 31324042.
- ↑ "Isolation, characterization, and inhibition kinetics of enolase from Streptococcus rattus FA-1". Infection and Immunity 58 (4): 1043–1047. April 1990. doi:10.1128/IAI.58.4.1043-1047.1990. PMID 2318530.
Further reading
- "The isolation and characterization of rabbit muscle enolase". The Journal of Biological Chemistry 236 (12): 3227–3231. December 1961. doi:10.1016/S0021-9258(18)94000-7. PMID 13908561.
- Boyer, P.D., Lardy, H. and Myrback, K. (Eds.), The Enzymes, 2nd ed., vol. 5, Academic Press, New York, 1961, p. 471-494.
- "A Purification of Brewers' and Bakers' Yeast Enolase Yielding a Single Active Component". The Journal of Biological Chemistry 239 (8): 2464–2468. August 1964. doi:10.1016/S0021-9258(18)93875-5. PMID 14235523.
External links
- Enolase at the US National Library of Medicine Medical Subject Headings (MeSH)
{{Navbox
| name = Glycolysis enzymes | title = Metabolism: carbohydrate metabolism: [[Biology:Glycoglycolysis/gluconeogenesis enzymes | state = autocollapse | listclass = hlist
| group1 = Glycolysis
| list1 =
| |
| group2 = Gluconeogenesis only| list2 =
| to oxaloacetate: | |
|---|---|
| from lactate (Cori cycle): | |
| from alanine (Alanine cycle): | |
| from glycerol: |
| group4 = Regulatory | list4 =
}}
 |