Biology:Protein-glutamate O-methyltransferase
| protein-glutamate O-methyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 2.1.1.80 | ||||||||
| CAS number | 9055-09-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| CheR methyltransferase, all-alpha domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
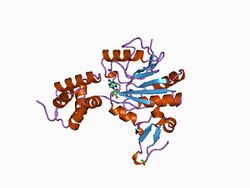 chemotaxis receptor recognition by protein methyltransferase cher | |||||||||
| Identifiers | |||||||||
| Symbol | CheR_N | ||||||||
| Pfam | PF03705 | ||||||||
| InterPro | IPR022641 | ||||||||
| SCOP2 | 1af7 / SCOPe / SUPFAM | ||||||||
| |||||||||
| CheR methyltransferase, SAM binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 chemotaxis receptor recognition by protein methyltransferase cher | |||||||||
| Identifiers | |||||||||
| Symbol | CheR | ||||||||
| Pfam | PF01739 | ||||||||
| Pfam clan | CL0063 | ||||||||
| InterPro | IPR022642 | ||||||||
| SCOP2 | 1af7 / SCOPe / SUPFAM | ||||||||
| |||||||||
In enzymology, a protein-glutamate O-methyltransferase (EC 2.1.1.80) is an enzyme that catalyzes the chemical reaction
- S-adenosyl-L-methionine + protein L-glutamate [math]\displaystyle{ \rightleftharpoons }[/math] S-adenosyl-L-homocysteine + protein L-glutamate methyl ester
Thus, the two substrates of this enzyme are S-adenosyl methionine and protein L-glutamic acid, whereas its two products are S-adenosylhomocysteine and protein L-glutamate methyl ester.
This enzyme belongs to the family of transferases, specifically those transferring one-carbon group methyltransferases. The systematic name of this enzyme class is S-adenosyl-L-methionine:protein-L-glutamate O-methyltransferase. Other names in common use include methyl-accepting chemotaxis protein O-methyltransferase, S-adenosylmethionine-glutamyl methyltransferase, methyl-accepting chemotaxis protein methyltransferase II, S-adenosylmethionine:protein-carboxyl O-methyltransferase, protein methylase II, MCP methyltransferase I, MCP methyltransferase II, protein O-methyltransferase, protein(aspartate)methyltransferase, protein(carboxyl)methyltransferase, protein carboxyl-methylase, protein carboxyl-O-methyltransferase, protein carboxylmethyltransferase II, protein carboxymethylase, protein carboxymethyltransferase, and protein methyltransferase II. This enzyme participates in bacterial chemotaxis - general and bacterial chemotaxis - organism-specific.
CheR proteins are part of the chemotaxis signaling mechanism which methylates the chemotaxis receptor at specific glutamate residues. Methyl transfer from the ubiquitous S-adenosyl-L-methionine (AdoMet/SAM) to either nitrogen, oxygen or carbon atoms is frequently employed in diverse organisms ranging from bacteria to plants and mammals. The reaction is catalysed by methyltransferases (Mtases) and modifies DNA, RNA, proteins and small molecules, such as catechol for regulatory purposes. The various aspects of the role of DNA methylation in prokaryotic restriction-modification systems and in a number of cellular processes in eukaryotes including gene regulation and differentiation is well documented.
Flagellated bacteria swim towards favourable chemicals and away from deleterious ones. Sensing of chemoeffector gradients involves chemotaxis receptors, transmembrane (TM) proteins that detect stimuli through their periplasmic domains and transduce the signals via their cytoplasmic domains .[1] Signalling outputs from these receptors are influenced both by the binding of the chemoeffector ligand to their periplasmic domains and by methylation of specific glutamate residues on their cytoplasmic domains. Methylation is catalysed by CheR, an S-adenosylmethionine-dependent methyltransferase,[1] which reversibly methylates specific glutamate residues within a coiled coil region, to form gamma-glutamyl methyl ester residues.[1][2] The structure of the Salmonella typhimurium chemotaxis receptor methyltransferase CheR, bound to S-adenosylhomocysteine, has been determined to a resolution of 2.0 Angstrom.[1] The structure reveals CheR to be a two-domain protein, with a smaller N-terminal helical domain linked via a single polypeptide connection to a larger C-terminal alpha/beta domain. The C-terminal domain has the characteristics of a nucleotide-binding fold, with an insertion of a small anti-parallel beta-sheet subdomain. The S-adenosylhomocysteine-binding site is formed mainly by the large domain, with contributions from residues within the N-terminal domain and the linker region.[1]
Structural studies
As of late 2007, two structures have been solved for this class of enzymes, with PDB accession codes 1AF7 and 1BC5.
References
- ↑ 1.0 1.1 1.2 1.3 1.4 "Crystal structure of the chemotaxis receptor methyltransferase CheR suggests a conserved structural motif for binding S-adenosylmethionine". Structure 5 (4): 545–58. April 1997. doi:10.1016/S0969-2126(97)00210-4. PMID 9115443.
- ↑ "Chemotaxis receptor recognition by protein methyltransferase CheR". Nat. Struct. Biol. 5 (6): 446–50. June 1998. doi:10.1038/nsb0698-446. PMID 9628482.
Further reading
- "Purification and characterization of Bacillus subtilis methyl-accepting chemotaxis protein methyltransferase II". J. Biol. Chem. 257 (14): 8412–7. 1982. doi:10.1016/S0021-9258(18)34347-3. PMID 6806296.
- "Isolation of glutamic acid methyl ester from an Escherichia coli membrane protein involved in chemotaxis". J. Biol. Chem. 252 (10): 3214–8. 1977. doi:10.1016/S0021-9258(17)40373-5. PMID 16888.
- "Purification and characterization of the S-adenosylmethionine:glutamyl methyltransferase that modifies membrane chemoreceptor proteins in bacteria". J. Biol. Chem. 262 (18): 8537–43. 1987. doi:10.1016/S0021-9258(18)47447-9. PMID 3298235.
- "Identification of a protein methyltransferase as the cheR gene product in the bacterial sensing system". Proc. Natl. Acad. Sci. U.S.A. 74 (2): 533–7. 1977. doi:10.1073/pnas.74.2.533. PMID 322131. Bibcode: 1977PNAS...74..533S.
 |

