Medicine:Congenital stationary night blindness
| Congenital stationary night blindness | |
|---|---|
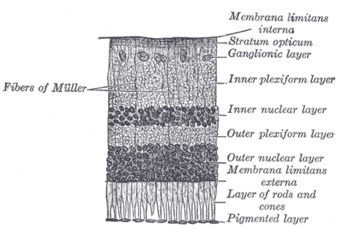 | |
| Malfunction in transmission from the photoreceptors in the outer nuclear layer to bipolar cells in the inner nuclear layer underlies CSNB. |
Congenital stationary night blindness (CSNB) is a rare non-progressive retinal disorder. People with CSNB often have difficulty adapting to low light situations due to impaired photoreceptor transmission. These patients may also have reduced visual acuity, myopia, nystagmus, and strabismus. CSNB has two forms -- complete, also known as type-1 (CSNB1), and incomplete, also known as type-2 (CSNB2), which are distinguished by the involvement of different retinal pathways. In CSNB1, downstream neurons called bipolar cells are unable to detect neurotransmission from photoreceptor cells. CSNB1 can be caused by mutations in various genes involved in neurotransmitter detection, including NYX. In CSNB2, the photoreceptors themselves have impaired neurotransmission function; this is caused primarily by mutations in the gene CACNA1F, which encodes a voltage-gated calcium channel important for neurotransmitter release. CSNB has been identified in horses and dogs as the result of mutations in TRPM1 (Horse, "LP")[1], GRM6 (Horse, "CSNB2")[2], and LRIT3 (Dog, CSNB)[3].
Congenital stationary night blindness (CSNB) can be inherited in an X-linked, autosomal dominant, or autosomal recessive pattern, depending on the genes involved.
Two forms of CSNB can also affect horses, one linked to the leopard complex of equine coat colors and the other found in certain horse breeds. Both are autosomal recessives.[4][5]
Symptoms and signs
The X-linked varieties of congenital stationary night blindness (CSNB) can be differentiated from the autosomal forms by the presence of myopia, which is typically absent in the autosomal forms. Patients with CSNB often have impaired night vision, myopia, reduced visual acuity, strabismus and nystagmus. Individuals with the complete form of CSNB (CSNB1) have highly impaired rod sensitivity (reduced ~300x) as well as cone dysfunction. Patients with the incomplete form can present with either myopia or hyperopia.[6]
Cause
CSNB is caused by malfunctions in neurotransmission from rod and cone photoreceptors to bipolar cells in the retina.[7] At this first synapse, information from photoreceptors is divided into two channels: ON and OFF. The ON pathway detects light onset, while the OFF pathway detects light offset.[8] The malfunctions in CSNB1 specifically affect the ON pathway, by hindering the ability of ON-type bipolar cells to detect neurotransmitter released from photoreceptors.[7] Rods, which are responsible for low-light vision, make contacts with ON-type bipolar cells only, while, cones, which are responsible for bright-light vision, make contacts with bipolar cells of both ON an OFF subtypes.[9] Because the low-light sensing rods feed only into the ON pathway, individuals with CSNB1 typically have problems with night vision, while vision in well-lit conditions is spared.[7] In CSNB2, release of neurotransmitter from photoreceptors is impaired, leading to involvement of both ON and OFF pathways.
The electroretinogram (ERG) is an important tool for diagnosing CSNB. The ERG a-wave, which reflects the function of the phototransduction cascade in response to a light flashes, is typically normal in CSNB patients, although in some cases phototransduction is also affected, leading to a reduced a-wave. The ERG b-wave, which primarily reflects the function of ON-bipolar cells, is greatly reduced in CSNB2 cases, and completely absent in CSNB1 cases.[7][10]
Genetics
Only three rhodopsin mutations have been found associated with congenital stationary night blindness (CSNB).[11] Two of these mutations are found in the second transmembrane helix of rhodopsin at Gly-90 and Thr-94. Specifically, these mutations are the Gly90Asp [12] and the Thr94Ile, which has been the most recent one reported.[13] The third mutation is Ala292Glu, and it is located in the seventh transmembrane helix, in proximity to the site of retinal attachment at Lys-296.[14] Mutations associated with CSNB affect amino acid residues near the protonated Schiff base (PSB) linkage. They are associated with changes in conformational stability and the protonated status of the PSB nitrogen.[15]
Pathophysiology
CSNB1
The complete form of X-linked congenital stationary night blindness, also known as nyctalopia, is caused by mutations in the NYX gene (Nyctalopin on X-chromosome), which encodes a small leucine-rich repeat (LRR) family protein of unknown function.[16][17] This protein consists of an N-terminal signal peptide and 11 LRRs (LRR1-11) flanked by cysteine-rich LRRs (LRRNT and LRRCT). At the C-terminus of the protein there is a putative GPI anchor site. Although the function of NYX is yet to be fully understood, it is believed to be located extracellularly. A naturally occurring deletion of 85 bases in NYX in some mice leads to the "nob" (no b-wave) phenotype, which is highly similar to that seen in CSNB1 patients.[18] NYX is expressed primarily in the rod and cone cells of the retina. There are currently almost 40 known mutations in NYX associated with CSNB1, Table 1., located throughout the protein. As the function of the nyctalopin protein is unknown, these mutations have not been further characterized. However, many of them are predicted to lead to truncated proteins that, presumably, are non-functional.
| Mutation | Position | References | |
|---|---|---|---|
| Nucleotide | Amino acid | ||
| c.?-1_?-61del | 1_20del | Signal sequence | [17] |
| Splicing | Intron 1 | [19] | |
| c.?-63_1443-?del | 21_481del | [17] | |
| c.48_64del | L18RfsX108 | Signal sequence | [19] |
| c.85_108del | R29_A36del | N-terminal LRR | [16] |
| c.G91C | C31S | LRRNT | [17] |
| c.C105A | C35X | LRRNT | [17] |
| c.C169A | P57T | LRRNT | |
| c.C191A | A64E | LRR1 | [20] |
| c.G281C | R94P | LRR2 | [21] |
| c.301_303del | I101del | LRR2 | [17] |
| c.T302C | I101T | LRR2 | [21] |
| c.340_351del | E114_A118del | LRR3 | [17][19] |
| c.G427C | A143P | LRR4 | [17] |
| c.C452T | P151L | LRR4 | [16] |
| c.464_465insAGCGTGCCCGAGCGCCTCCTG | S149_V150dup+P151_L155dup | LRR4 | [16] |
| c.C524G | P175R | LRR5 | [17] |
| c.T551C | L184P | LRR6 | [16] |
| c.556_618delins | H186?fsX260 | LRR6 | [16] |
| c.559_560delinsAA | A187K | LRR6 | [17] |
| c.613_621dup | 205_207dup | LRR7 | [16][17] |
| c.628_629ins | R209_S210insCLR | LRR7 | [16] |
| c.T638A | L213Q | LRR7 | [16] |
| c.A647G | N216S | LRR7 | [16][19] |
| c.T695C | L232P | LRR8 | [16] |
| c.727_738del | 243_246del | LRR8 | [17] |
| c.C792G | N264K | LRR9 | [16] |
| c.T854C | L285P | LRR10 | [16] |
| c.T893C | F298S | LRR10 | [16] |
| c.C895T | Q299X | LRR10 | [19] |
| c.T920C | L307P | LRR11 | [17] |
| c.A935G | N312S | LRR11 | [17] |
| c.T1040C | L347P | LRRCT | [17] |
| c.G1049A | W350X | LRRCT | [16] |
| c.G1109T | G370V | LRRCT | [17] |
| c.1122_1457del | S374RfsX383 | LRRCT | [17][19] |
| c.1306del | L437WfsX559 | C-terminus | [19] |
| LRR: leucine-rich repeat, LRRNT and LRRCT: N- and C-terminal cysteine-rich LRRs. | |||
CSNB2
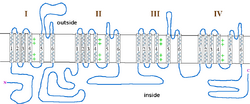
The incomplete form of X-linked congenital stationary night blindness (CSNB2) is caused by mutations in the CACNA1F gene, which encodes the voltage-gated calcium channel CaV1.4 expressed heavily in retina.[22][23] One of the important properties of this channel is that it inactivates at an extremely low rate. This allows it to produce sustained Ca2+ entry upon depolarization. As photoreceptors depolarize in the absence of light, CaV1.4 channels operate to provide sustained neurotransmitter release upon depolarization.[24] This has been demonstrated in CACNA1F mutant mice that have markedly reduced photoreceptor calcium signals.[25] There are currently 55 mutations in CACNA1F located throughout the channel, Table 2 and Figure 1. While most of these mutations result in truncated and, likely, non-functional channels, it is expected that they prevent the ability of light to hyperpolarize photoreceptors. Of the mutations with known functional consequences, 4 produce channels that are either completely non-functional, and two that result in channels which open at far more hyperpolarized potentials than wild-type. This will result in photoreceptors that continue to release neurotransmitter even after light-induced hyperpolarization.
| Mutation | Position | Effect | References | |
|---|---|---|---|---|
| Nucleotide | Amino Acid | |||
| c.C148T | R50X | N-terminus | ||
| c.151_155delAGAAA | R51PfsX115 | N-terminus | [26] | |
| c.T220C | C74R | N-terminus | [26] | |
| c.C244T | R82X | N-terminus | [27][26] | |
| c.466_469delinsGTAGGGGTGCT CCACCCCGTAGGGGTGCTCCACC |
S156VdelPinsGVKHOVGVLH | D1S2-3 | [27][28][29] | |
| Splicing | Intron 4 | [27] | ||
| c.T685C | S229P | D1S4-5 | [26] | |
| c.G781A | G261R | D1-pore | [26] | |
| c.G832T | E278X | D1-pore | ||
| c.904insG | R302AfsX314 | D1-pore | [28] | |
| c.951_953delCTT | F318del | D1-pore | [27] | |
| c.G1106A | G369D | D1S6 | Activates ~20mV more negative than wild-type, increases time to peak current and decreases inactivation, increased Ca2+ permeability. | [22][24][27][26][30] |
| c.1218delC | W407GfsX443 | D1-2 | [23][27][31] | |
| c.C1315T | Q439X | D1-2 | [26] | |
| c.G1556A | R519Q | D1-2 | Decreased expression | [22][32] |
| c.C1873T | R625X | D2S4 | [27][26] | |
| c.G2021A | G674D | D2S5 | [24][27][28] | |
| c.C2071T | R691X | D2-pore | [20] | |
| c.T2258G | F753C | D2S6 | [26] | |
| c.T2267C | I756T | D2S6 | Activates ~35mV more negative than wild-type, inactivates more slowly | |
| Splicing | Intron 19 | [26] | ||
| c.T2579C | L860P | D2-3 | [26] | |
| c.C2683T | R895X | D3S1-2 | [19][20][23][27] | |
| Splicing | Intron 22 | [26][28] | ||
| Splicing | Intron 22 | [26] | ||
| c.C2783A | A928D | D3S2-3 | [24][27] | |
| c.C2905T | R969X | D3S4 | [22][26] | |
| c.C2914T | R972X | D3S4 | [31] | |
| Splicing | Intron24 | [27] | ||
| c.C2932T | R978X | D3S4 | [28] | |
| c.3006_3008delCAT | I1003del | D3S4-5 | [27] | |
| c.G3052A | G1018R | D3S5 | [26] | |
| c.3125delG | G1042AfsX1076 | D3-pore | [27] | |
| c.3166insC | L1056PfsX1066 | D3-pore | [22][23][27][26] | |
| c.C3178T | R1060W | D3-pore | [22][26] | |
| c.T3236C | L1079P | D3-pore | Does not open without BayK, activates ~5mV more negative than wild-type | [26][30] |
| c.3672delC | L1225SfsX1266 | D4S2 | [23][27] | |
| c.3691_3702del | G1231_T1234del | D4S2 | [22][26] | |
| c.G3794T | S1265I | D4S3 | [20] | |
| c.C3886A | R1296S | D4S4 | [20] | |
| c.C3895T | R1299X | D4S4 | [23][27][26] | |
| Splicing | Intron 32 | [26] | ||
| c.C4075T | Q1359X | D4-pore | [22][26] | |
| c.T4124A | L1375H | D4-pore | Decreased expression | [22][26][32] |
| Splicing | Intron 35 | [26] | ||
| c.G4353A | W1451X | C-terminus | Non-functional | [23][24][27][30] |
| c.T4495C | C1499R | C-terminus | [26] | |
| c.C4499G | P1500R | C-terminus | [26] | |
| c.T4523C | L1508P | C-terminus | [26] | |
| Splicing | intron 40 | [27] | ||
| c.4581delC | F1528LfsX1535 | C-terminus | ||
| c.A4804T | K1602X | C-terminus | [22][26] | |
| c.C5479T | R1827X | C-terminus | [26] | |
| c.5663delG | S1888TfsX1931 | C-terminus | [27] | |
| c.G5789A | R1930H | C-terminus | [20] | |
Diagnosis
Footnotes
- ↑ "Evidence for a retroviral insertion in TRPM1 as the cause of congenital stationary night blindness and leopard complex spotting in the horse". PLOS ONE 8 (10): e78280. 2013-10-22. doi:10.1371/journal.pone.0078280. PMID 24167615. Bibcode: 2013PLoSO...878280B.
- ↑ "Whole-genome sequencing identifies missense mutation in GRM6 as the likely cause of congenital stationary night blindness in a Tennessee Walking Horse". Equine Veterinary Journal 53 (2): 316–323. March 2021. doi:10.1111/evj.13318. PMID 32654228.
- ↑ "Genome-wide association study and whole-genome sequencing identify a deletion in LRIT3 associated with canine congenital stationary night blindness". Scientific Reports 9 (1): 14166. October 2019. doi:10.1038/s41598-019-50573-7. PMID 31578364. Bibcode: 2019NatSR...914166D.
- ↑ "Appaloosa Panel 2 | Veterinary Genetics Laboratory". https://vgl.ucdavis.edu/panel/appaloosa-panel-2.
- ↑ "Congenital Stationary Night Blindness (CSNB2) in Tennessee Walking Horses | Veterinary Genetics Laboratory" (in en). https://vgl.ucdavis.edu/test/csnb-tennessee-walking-horse.
- ↑ "Evidence for genetic heterogeneity in X-linked congenital stationary night blindness". American Journal of Human Genetics 62 (4): 865–875. April 1998. doi:10.1086/301781. PMID 9529339.
- ↑ 7.0 7.1 7.2 7.3 "Congenital stationary night blindness: an analysis and update of genotype-phenotype correlations and pathogenic mechanisms". Progress in Retinal and Eye Research 45: 58–110. March 2015. doi:10.1016/j.preteyeres.2014.09.001. PMID 25307992.
- ↑ "Retinal bipolar cells: elementary building blocks of vision". Nature Reviews. Neuroscience 15 (8): 507–519. August 2014. doi:10.1038/nrn3783. PMID 25158357.
- ↑ "Wiring patterns in the mouse retina: collecting evidence across the connectome, physiology and light microscopy". The Journal of Physiology 592 (22): 4809–4823. November 2014. doi:10.1113/jphysiol.2014.277228. PMID 25172948.
- ↑ "The negative ERG: clinical phenotypes and disease mechanisms of inner retinal dysfunction". Survey of Ophthalmology 53 (1): 16–40. 2008. doi:10.1016/j.survophthal.2007.10.010. PMID 18191655.
- ↑ "The eye photoreceptor protein rhodopsin. Structural implications for retinal disease". FEBS Letters 528 (1–3): 17–22. September 2002. doi:10.1016/s0014-5793(02)03241-6. PMID 12297272.
- ↑ "Rhodopsin mutation G90D and a molecular mechanism for congenital night blindness". Nature 367 (6464): 639–42. February 1994. doi:10.1038/367639a0. PMID 8107847. Bibcode: 1994Natur.367..639R.
- ↑ N. al-Jandal, G.J. Farrar, A.S. Kiang, M.M. Humphries, N. Bannon, J.B. Findlay, P. Humphries and P.F. Kenna Hum. Mutat. 13 (1999), pp. 75–81.
- ↑ "Heterozygous missense mutation in the rhodopsin gene as a cause of congenital stationary night blindness". Nature Genetics 4 (3): 280–3. July 1993. doi:10.1038/ng0793-280. PMID 8358437.
- ↑ "Dark-light: model for nightblindness from the human rhodopsin Gly-90-->Asp mutation". Proceedings of the National Academy of Sciences of the United States of America 92 (3): 880–4. January 1995. doi:10.1073/pnas.92.3.880. PMID 7846071. Bibcode: 1995PNAS...92..880S.
- ↑ 16.00 16.01 16.02 16.03 16.04 16.05 16.06 16.07 16.08 16.09 16.10 16.11 16.12 16.13 16.14 "Mutations in NYX, encoding the leucine-rich proteoglycan nyctalopin, cause X-linked complete congenital stationary night blindness". Nature Genetics 26 (3): 319–323. November 2000. doi:10.1038/81619. PMID 11062471.
- ↑ 17.00 17.01 17.02 17.03 17.04 17.05 17.06 17.07 17.08 17.09 17.10 17.11 17.12 17.13 17.14 17.15 17.16 "The complete form of X-linked congenital stationary night blindness is caused by mutations in a gene encoding a leucine-rich repeat protein". Nature Genetics 26 (3): 324–327. November 2000. doi:10.1038/81627. PMID 11062472.
- ↑ "Identification of the gene and the mutation responsible for the mouse nob phenotype". Investigative Ophthalmology & Visual Science 44 (1): 378–384. January 2003. doi:10.1167/iovs.02-0501. PMID 12506099.
- ↑ 19.0 19.1 19.2 19.3 19.4 19.5 19.6 19.7 "Mutations in the CACNA1F and NYX genes in British CSNBX families". Human Mutation 21 (2): 169. February 2003. doi:10.1002/humu.9106. PMID 12552565.
- ↑ 20.0 20.1 20.2 20.3 20.4 20.5 Cite error: Invalid
<ref>tag; no text was provided for refs namedZeitz_2005 - ↑ 21.0 21.1 "CSNB1 in Chinese families associated with novel mutations in NYX". Journal of Human Genetics 51 (7): 634–640. 2006. doi:10.1007/s10038-006-0406-5. PMID 16670814.
- ↑ 22.0 22.1 22.2 22.3 22.4 22.5 22.6 22.7 22.8 22.9 "An L-type calcium-channel gene mutated in incomplete X-linked congenital stationary night blindness". Nature Genetics 19 (3): 260–263. July 1998. doi:10.1038/940. PMID 9662399.
- ↑ 23.0 23.1 23.2 23.3 23.4 23.5 23.6 "Loss-of-function mutations in a calcium-channel alpha1-subunit gene in Xp11.23 cause incomplete X-linked congenital stationary night blindness". Nature Genetics 19 (3): 264–267. July 1998. doi:10.1038/947. PMID 9662400.
- ↑ 24.0 24.1 24.2 24.3 24.4 "The CACNA1F gene encodes an L-type calcium channel with unique biophysical properties and tissue distribution". The Journal of Neuroscience 24 (7): 1707–1718. February 2004. doi:10.1523/JNEUROSCI.4846-03.2004. PMID 14973233.
- ↑ "Mutation of the calcium channel gene Cacna1f disrupts calcium signaling, synaptic transmission and cellular organization in mouse retina". Human Molecular Genetics 14 (20): 3035–3046. October 2005. doi:10.1093/hmg/ddi336. PMID 16155113.
- ↑ 26.00 26.01 26.02 26.03 26.04 26.05 26.06 26.07 26.08 26.09 26.10 26.11 26.12 26.13 26.14 26.15 26.16 26.17 26.18 26.19 26.20 26.21 26.22 26.23 26.24 26.25 26.26 26.27 26.28 "Thirty distinct CACNA1F mutations in 33 families with incomplete type of XLCSNB and Cacna1f expression profiling in mouse retina". European Journal of Human Genetics 10 (8): 449–456. August 2002. doi:10.1038/sj.ejhg.5200828. PMID 12111638.
- ↑ 27.00 27.01 27.02 27.03 27.04 27.05 27.06 27.07 27.08 27.09 27.10 27.11 27.12 27.13 27.14 27.15 27.16 27.17 27.18 Cite error: Invalid
<ref>tag; no text was provided for refs namedBoycott_2001 - ↑ 28.0 28.1 28.2 28.3 28.4 "Novel CACNA1F mutations in Japanese patients with incomplete congenital stationary night blindness". Investigative Ophthalmology & Visual Science 42 (7): 1610–1616. June 2001. PMID 11381068.
- ↑ "Retinal and optic disc atrophy associated with a CACNA1F mutation in a Japanese family". Archives of Ophthalmology 121 (7): 1028–1033. July 2003. doi:10.1001/archopht.121.7.1028. PMID 12860808.
- ↑ 30.0 30.1 30.2 "Congenital stationary night blindness type 2 mutations S229P, G369D, L1068P, and W1440X alter channel gating or functional expression of Ca(v)1.4 L-type Ca2+ channels". The Journal of Neuroscience 25 (1): 252–259. January 2005. doi:10.1523/JNEUROSCI.3054-04.2005. PMID 15634789.
- ↑ 31.0 31.1 Cite error: Invalid
<ref>tag; no text was provided for refs namedAllen_2003 - ↑ 32.0 32.1 "Effects of congenital stationary night blindness type 2 mutations R508Q and L1364H on Cav1.4 L-type Ca2+ channel function and expression". Journal of Neurochemistry 96 (6): 1648–1658. March 2006. doi:10.1111/j.1471-4159.2006.03678.x. PMID 16476079.
External links
| Classification | |
|---|---|
| External resources |
 |