Biology:Arenavirus
| Arenaviridae | |
|---|---|
| |
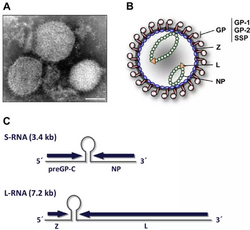
| (A) Electron micrograph of Lassa virus, bar = 100nm (B) diagram and (C) genome of arenavirus | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Negarnaviricota |
| Class: | Ellioviricetes |
| Order: | Bunyavirales |
| Family: | Arenaviridae |
| Genera | |
| |
An arenavirus is a bi- or trisegmented ambisense RNA virus that is a member of the family Arenaviridae.[1][2] These viruses infect rodents and occasionally humans. A class of novel, highly divergent arenaviruses, properly known as reptarenaviruses, have also been discovered which infect snakes to produce inclusion body disease. At least eight arenaviruses are known to cause human disease. The diseases derived from arenaviruses range in severity. Aseptic meningitis, a severe human disease that causes inflammation covering the brain and spinal cord, can arise from the lymphocytic choriomeningitis virus. Hemorrhagic fever syndromes, including Lassa fever, are derived from infections such as Guanarito virus, Junin virus, Lassa virus, Lujo virus,[3] Machupo virus, Sabia virus, or Whitewater Arroyo virus.[4] Because of the epidemiological association with rodents, some arenaviruses and bunyaviruses are designated as roboviruses.
Structure
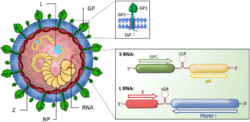
Viewed in cross-section, arenaviruses contain grainy particles that are ribosomes acquired from their host cells. It is from this characteristic that they acquired the name arena, from the Latin root meaning sand.[5] The ribosomal structures are not believed to be essential for virus replication. Virus particles, or virions, are pleomorphic (variable in shape) but are often spherical, with a diameter of 60–300 nm, and are covered with surface glycoprotein spikes.[6]
The virus contains a beaded nucleocapsid with two single-stranded RNA segments. The nucleocapsid consists of a core of nucleic acid enclosed in a protein coat. Although they are categorized as negative-sense viruses,[7] arenaviruses are ambisense. While sections of their genome encode genes in the negative sense (reverse polarity), other sections encode genes in the opposite (forward/positive sense) direction. This complex gene expression structure is theorized to be a primitive regulatory system, allowing the virus to control what proteins are synthesized at what point in the life cycle. The life cycle of the arenavirus is restricted to the cell cytoplasm.[citation needed]
Genome
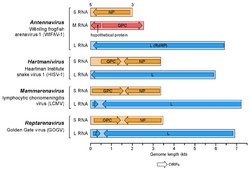
Arenaviruses have a segmented RNA genome that consists of two single-stranded ambisense RNAs.[8] As with all negative-sense RNA viruses, the genomic RNA alone is not infectious and the viral replication machinery is required to initiate infection within a host cell.[9] Genomic sense RNA packaged into the arenavirus virion is designated negative-sense RNA, and must first be copied into a positive-sense mRNA in order to produce viral protein.[10] The RNA segments are denoted Small (S), Medium (M; if present), and Large (L),[8][11] and code for four viral proteins in a unique ambisense coding strategy.[12][13] For mammarenaviruses and reptarenaviruses, each RNA segment codes for two viral proteins in opposite orientation such that the negative-sense RNA genome serves as the template for transcription of a single mRNA and the positive-sense copy of the RNA genome templates a second mRNA.[10] The separate coding sequences of the two viral proteins are divided by an intergenic region RNA sequence that is predicted to fold into a stable hairpin structure.[14]
The extreme termini of each RNA segment contains a 19 nucleotide highly conserved sequence that is critical for recruitment of the viral replication machinery and initiation of viral mRNA transcription and genomic replication.[15][16][17][18][19] The conserved 5' and 3' RNA termini sequences are complementary and allows each RNA segment to adopt a double-stranded RNA panhandle structure[20] that maintains the termini in close proximity and results in a circular appearance to purified arenavirus genomic templates visualized by electron microscopy.[21][22] The double-stranded RNA panhandle structure is critical for efficient viral RNA synthesis,[18][23] but potential interterminal double-stranded RNA interactions must be transiently relieved in order to recruit the viral polymerase.[19]
The S-segment RNA is approximately 3.5 kb, and encodes the viral nucleocapsid protein (NP) and glycoprotein (GPC).[24] The L-segment RNA is approximately 7.2 kb, and encodes the viral RNA-dependent RNA-polymerase (L) and a small RING-domain containing protein (Z).[25][26][27]
The Z protein forms homo oligomers and a structural component of the virions.[28] The formation of these oligomers is an essential step for particle assembly and budding. Binding between Z and the viral envelope glycoprotein complex is required for virion infectivity. Z also interacts with the L and NP proteins. Polymerase activity appears to be modulated by the association between the L and Z proteins. Interaction between the Z and NP proteins is critical for genome packaging.[citation needed]
Microbiology
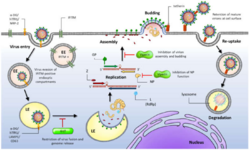
The glycoprotein (GP) is synthesised as a precursor molecule.[29] It is cleaved into three parts - GP1, GP2 and a stable signal peptide (SSP). These reactions are catalysed by cellular signal peptidases and the cellular enzyme Subtilisin Kexin Isozyme-1 (SKI-1)/Site-1 Protease (S1P). These processes are essential for fusion competence and incorporation of mature GP into nascent budding virion particles.[citation needed]
Taxonomy
Within the family Arenaviridae, arenaviruses were formerly all placed in the genus Arenavirus, but in 2015 were divided into the genera Mammarenavirus for those with mammalian hosts and Reptarenavirus for those infecting snakes.[30][31] Reptarenaviruses and mammarenavirus are separated by an impenetrable species barrier. Infected rodents cannot pass disease onto snakes, and IBD in captive snakes is not transmissible to humans.[citation needed]
A third genus, Hartmanivirus (not to be confused with genus Haartmanvirus of vibrio phages in family Demerecviridae, order Caudovirales), has also been established,[32] including other species that infect snakes. The organisation of the genome of this genus is typical of arenaviruses but their glycoproteins resemble those of filoviruses. Species in this genus lack the matrix protein.[33]
A fourth genus, Antennavirus, has also been established[34] to accommodate two arenaviruses found in striated frogfish (Antennarius striatus).[35] A third antennavirus has been detected in Chinook salmon and sockeye salmon.[36]
Mammarenaviruses can be divided into two serogroups, which differ genetically and by geographical distribution:[37] When the virus is classified "Old World" this means it was found in the Eastern Hemisphere in places such as Europe, Asia, and Africa. When it is found in the Western Hemisphere, in places such as Argentina, Bolivia, Venezuela, Brazil, and the United States, it is classified "New World". Lymphocytic choriomeningitis (LCM) virus is the only mammarenavirus found worldwide because of its ubiquitous Old World host, the house mouse. Old and New World area viruses appear to have diverged ~45,000 years ago.[38] The Old World Mammarenaviruses originated ~23.1-1.88 thousand years ago, most likely in Southern Africa, while the New World Mammarenaviruses evolved in the Latin America-Caribbean region ~41.4-3.3 thousand years ago.
Mammarenavirus
Old World complex
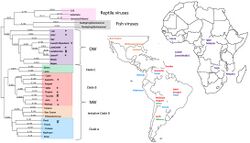
- Alxa virus (ALXV)
- Bitu virus (BITV)
- Dandenong virus (DANV)
- Dhati Welel virus (DHWV)
- Gairo virus (GAIV)
- Gbagroube virus
- Ippy virus (IPPYV)
- Kitale virus (KTLV)
- Kodoko virus (KODV)
- Kwanza virus (KWAV)
- Lassa virus (LASV)
- Lìjiāng virus (LIJV)
- Loei River virus (LORV)
- Lujo virus(LUJV)
- Luna virus (LUAV)
- Lunk virus (LNKV)
- Lymphocytic choriomeningitis virus (LCMV)
- Mafiga virus (MAFV)
- Mariental virus (MRTV)
- Merino Walk virus (MRWV)
- Menekre virus
- Minu virus
- Mobala virus (MOBV)
- Mopeia virus (MOPV)
- Morogoro virus (MORV)
- Okahandja virus (OKAV)
- Ryukyu virus (RYKV)
- Solwezi virus (SOLV)
- Souris virus (SOUV)
- Wenzhou virus (WENV)
New World complex
- Clade A
- Allpahuayo virus (AALV)
- Flexal virus (FLEV)
- Paraná virus (PRAV)
- Pichindé virus (PICHV)
- Pirital virus (PIRV)
- Clade B
- Amaparí virus (AMAV)
- Aporé virus (APOV)
- Chapare virus (CHAPV)
- Cupixi virus (CUPXV)
- Guanarito virus (GTOV)
- Junín virus (JUNV)
- Machupo virus (MACV)
- Ocozocoautla de Espinosa virus
- Real de Catorce virus (RCTV)
- Tacaribe virus (TCRV)
- Xapuri virus (XAPV)
- Sabiá virus (SBAV)
- Clade C
- Latino virus (LATV)
- Oliveros virus (OLVV)
- Clade D
- Bear Canyon virus (BCNV)
- Catarina virus (CTNV)
- Skinner Tank virus (SKTV)
- Tamiami virus (TMMV)
- Whitewater Arroyo virus (WWAV)
- Catarina virus (CTNV)
- Big Brushy Tank virus (BBTV)
- Skinner Tank virus (SKTV)
- Tonto Creek virus (TTCV)
- Others
- Patawa virus
Reptarenavirus
- California reptarenavirus
- Giessen reptarenavirus
- Golden reptarenavirus
- Ordinary reptarenavirus
- Rotterdam reptarenavirus
Hartmanivirus
- Haartman hartmanivirus
- Heimat hartmanivirus
- Muikkunen hartmanivirus
- Schoolhouse hartmanivirus
- Setpatvet hartmanivirus
- Zurich hartmanivirus
Antennavirus
- Hairy antennavirus
- Salmon antennavirus
- Striated antennavirus
Evolution
The evolution of the Mammarenavirus genus has been studied.[38] The New World and Old World species diverged less than 45,000 years ago. The New World species evolved between 41,400 and 3,300 years ago in the Latin America-Caribbean region. The Old World species evolved between 23,100 and 1,880 years ago, most likely in southern Africa.[citation needed]
Reservoirs
Some arenaviruses are zoonotic pathogens and are generally associated with rodent—transmitted disease in humans. Each virus usually is associated with a particular rodent host species in which it is maintained. Arenaviruses persist in nature by infecting rodents first and then transmitted into humans. Humans can be infected through mucosal exposure to aerosols, or by direct contact of abraded skin with the infectious material, derived from infected rodents.[6] Aerosols are fine mists or sprays of rodent dried excreta, especially urine that is dropped in the environment. Most of the Arenaviruses caught by humans are within their own homes when these rodents seek shelter. The virus can be caught in factories, from food that has been contaminated, or within agricultural work areas. Humans' risk of contracting the Arenavirus infection is related to age, race, or sex within the degree of contact with the dried rodent excreta.[citation needed]
Epidemiology
Hosts
| Virus | Disease | Host | Distribution |
|---|---|---|---|
| Dandenong[39] | Dandenong hemorrhagic fever | Unknown | old world ( Australian cases from Serbia ) |
| Lymphocytic choriomeningitis virus | Lymphocytic choriomeningitis | House mouse (Mus musculus) | Worldwide |
| Lassa virus | Lassa fever | Natal Multimammate Mouse (Mastomys natalensis) | West Africa |
| Junin virus | Argentine hemorrhagic fever | Drylands Vesper Mouse (Calomys musculinus) | Argentina |
| Machupo virus | Bolivian hemorrhagic fever | Large Vesper Mouse (Calomys callosus) | Bolivia |
| Guanarito virus | Venezuelan hemorrhagic fever | Short-tailed Cane Mouse (Zygodontomys brevicauda) | Venezuela |
| Sabiá virus | Brazilian hemorrhagic fever | Unknown | Brazil |
| Tacaribe virus | Bat (Artibeus) | Trinidad | |
| Flexal virus | Influenza-like illness | Rice rat (Oryzomys) | Brazil |
| Whitewater Arroyo virus | Hemorrhagic fever | Woodrat (Neotoma) | Southwestern United States |
Clinical diseases
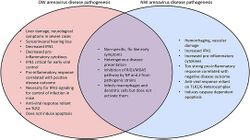
- Lymphocytic choriomeningitis (LCM) viruses cause influenza-like febrile illness, but occasionally they may cause meningitis, characteristically accompanied by large numbers of lymphocytes in the cerebrospinal fluid (as the name LCM suggests).
- Lassa virus causes Lassa fever. Lassa fever is endemic in west Africa. The virus was first isolated from Americans stationed in the village of Lassa, Nigeria. The virus can be transmitted person-to-person.
- Subclinical diseases: Serological studies suggest that inapparent infections particularly among members of hunting tribes are common.
- Clinical infections: Lassa fever is characterised by high fever, severe myalgia, coagulopathy, haemorrhagic skin rash, and occasional visceral haemorrhage as well as necrosis of liver and spleen.
- Other Arenaviruses like Junin virus, Machupo virus cause haemorrhagic fevers.
All of these diseases pose a great threat to public health in the regions where it is taking place. For example, when the Old World Lassa virus turns into Lassa fever, this usually results in a significant amount of mortality. Similarly the New World Junin virus causes Argentine hemorrhagic fever. This fever is a severe illness with hemorrhagic and neurological manifestations and a case fatality of fifteen to thirty percent.[6] The way this virus spreads is through increased traveling to and from endemic regions. This traveling has led to the importation of Lassa fever into non-endemic metropolitan areas all over the world.
Recent outbreaks
A new species of arenavirus named the Lujo virus has been linked to five patients who exhibited symptoms of viral hemorrhagic fever in South Africa.[40] The disease originated near Lusaka, Zambia and spread to Johannesburg, South Africa , after the first patient was transported to a hospital there. The results of genetic sequencing tests conducted by epidemiologists at Columbia University in New York City , USA, and at the Special Pathogens Branch of the Centers for Disease Control in Atlanta, USA, provided evidence that the causative agent of the disease is a virus from the family Arenaviridae, which ultimately resulted in the deaths of four out of the five infected in Zambia and South Africa during the outbreak which began in September 2008.[citation needed]
Arenavirus has also pinpointed as the cause of death of three donor organ recipients in Australia who contracted the virus after receiving kidney and a liver donations from a single infected organ donor in late 2006. All three died in the first week of 2007.[41][42]
WHO and its Global Outbreak Alert and Response Network (GOARN) partners continue to support the Ministries of Health of the two countries in various facets of the outbreak investigation, including laboratory diagnosis, investigations, active case finding and follow-up of contacts.[43]
Treatments
Very few treatment methods are available. The current lack of a licensed vaccine and limited therapeutic options for the arenavirus make it arguably among the most neglected virus groups. The only licensed drug for the treatment of human arenavirus infection is the nucleoside analogue ribavirin.[44] Ribavirin reduces morbidity and mortality in humans infected with certain arenaviruses, such as LASV and JUNV infections, if it is taken in the early stages of the disease. Ribavirin displays mixed success in treating severe arenaviral disease and is associated with significant toxicities.[45]
Experimental approaches
Effective antiviral drugs need to be produced at a low cost, taken orally, and able to withstand tropical climates due to the regions where these infections are occurring. For this reason high throughput screening (HTS) of small molecular libraries could be the answer to finding a better remedy. HTS collects libraries of small synthetic molecules that can be used to identify protein promoting "agonist" molecules or protein inhibiting "antagonist" interactions.[44] With HTS sustainable antiviral drugs can be discovered against possible new human pathogenic viruses.
Immunotherapy is another potential approach. Monoclonal antibodies against Junin virus have been tested in animal models. An immunotherapeutic agent active against all tested mammarenaviruses that use the transferrin receptor 1 as their receptor was under investigation in 2020.[46]
References
- ↑ Radoshitzky, SR; Buchmeier, MJ; Charrel, RN; Clegg, JCS; Gonzalez, JJ; Günther, S; Hepojoki, J; Kuhn, JH et al. (August 2019). "ICTV Virus Taxonomy Profile: Arenaviridae.". The Journal of General Virology 100 (8): 1200–1201. doi:10.1099/jgv.0.001280. PMID 31192784.
- ↑ Shi, Mang; Lin, Xian-Dan; Chen, Xiao; Tian, Jun-Hua; Chen, Liang-Jun; Li, Kun; Wang, Wen; Eden, John-Sebastian et al. (2018). "The evolutionary history of vertebrate RNA viruses". Nature 561 (7722): 202. doi:10.1038/s41586-018-0310-0. PMID 29946168.
- ↑ "Genetic detection and characterization of Lujo virus, a new hemorrhagic fever-associated arenavirus from southern Africa". PLOS Pathog. 5 (5): e1000455. May 2009. doi:10.1371/journal.ppat.1000455. PMID 19478873.
- ↑ Botten, J; Whitton, JL; Barrowman, P; Sidney, J; Whitmire, JK; Alexander, J; Kotturi, MF; Sette, A et al. (2010). "A Multivalent Vaccination Strategy for the Prevention of Old World Arenavirus Infection in Humans". Journal of Virology 84 (19): 9947–56. doi:10.1128/JVI.00672-10. PMID 20668086.
- ↑ Rowe, Wallace P.; Murphy, Frederick A.; Bergold, Gernot H.; Casals, Jordi; Hotchin, John; Johnson, Karl M.; Lehmann-Grube, Fritz; Mims, Cedric A. et al. (May 1970). "Arenoviruses: Proposed Name for a Newly Defined Virus Group". Journal of Virology 5 (5): 652. doi:10.1128/jvi.5.5.651-652.1970. PMID 4986852.
- ↑ 6.0 6.1 6.2 Emonet, Sebastien E.; Urata, Shuzo; De La Torre, Juan C. (2011). "Arenavirus reverse genetics: New approaches for the investigation of arenavirus biology and development of antiviral strategies". Virology 411 (2): 416–425. doi:10.1016/j.virol.2011.01.013. PMID 21324503.
- ↑ "Arenaviridae - Negative Sense RNA Viruses - Negative Sense RNA Viruses (2011)" (in en). https://talk.ictvonline.org/ictv-reports/ictv_9th_report/negative-sense-rna-viruses-2011/w/negrna_viruses/203/arenaviridae.
- ↑ 8.0 8.1 "RNA composition of Junin virus". J. Virol. 18 (3): 833–8. June 1976. doi:10.1128/JVI.18.3.833-838.1976. PMID 178925.
- ↑ "NP and L proteins of lymphocytic choriomeningitis virus (LCMV) are sufficient for efficient transcription and replication of LCMV genomic RNA analogs". J. Virol. 74 (8): 3470–7. April 2000. doi:10.1128/jvi.74.8.3470-3477.2000. PMID 10729120.
- ↑ 10.0 10.1 "Arenaviruses: Genomic RNAs, Transcription, and Replication". Arenaviruses I. Current Topics in Microbiology and Immunology. 262. 2002. 139–57. doi:10.1007/978-3-642-56029-3_6. ISBN 978-3-540-42244-0.
- ↑ Pedersen IR (March 1973). "Different classes of ribonucleic acid isolated from lymphocytic choriomeningitis virus". J. Virol. 11 (3): 416–23. doi:10.1128/JVI.11.3.416-423.1973. PMID 4734917.
- ↑ "Sequencing studies of pichinde arenavirus S RNA indicate a novel coding strategy, an ambisense viral S RNA". J. Virol. 52 (3): 897–904. December 1984. doi:10.1128/JVI.52.3.897-904.1984. PMID 6492264.
- ↑ "Nucleotide sequence of the Lassa virus (Josiah strain) S genome RNA and amino acid sequence comparison of the N and GPC proteins to other arenaviruses". Virology 168 (2): 421–5. February 1989. doi:10.1016/0042-6822(89)90287-0. PMID 2916333. https://zenodo.org/record/1258276.
- ↑ "The sequences of the N protein gene and intergenic region of the S RNA of pichinde arenavirus". Virology 134 (1): 208–19. April 1984. doi:10.1016/0042-6822(84)90286-1. PMID 6324469.
- ↑ "Analyses of the genomes of prototype pichinde arenavirus and a virulent derivative of Pichinde Munchique: evidence for sequence conservation at the 3' termini of their viral RNA species". Virology 116 (1): 363–7. January 1982. doi:10.1016/0042-6822(82)90429-9. PMID 6278715.
- ↑ "Nucleotide sequence conservation at the 3' termini of the virion RNA species of New World and Old World arenaviruses". Virology 121 (1): 200–3. August 1982. doi:10.1016/0042-6822(82)90130-1. PMID 6287720.
- ↑ Mattsson L (1989). "Chronic non-A, non-B hepatitis with special reference to the transfusion-associated form". Scand J Infect Dis Suppl 59: 1–55. doi:10.3109/inf.1988.20.suppl-59.01. PMID 2502835.
- ↑ 18.0 18.1 "Mutational analysis of the lassa virus promoter". J. Virol. 80 (24): 12414–9. December 2006. doi:10.1128/JVI.01374-06. PMID 17005649.
- ↑ 19.0 19.1 "Assembly of a functional Machupo virus polymerase complex". Proc. Natl. Acad. Sci. U.S.A. 107 (46): 20069–74. November 2010. doi:10.1073/pnas.1007152107. PMID 20978208. Bibcode: 2010PNAS..10720069K.
- ↑ "Arenavirus Gene Structure and Organization". Arenaviruses. Current Topics in Microbiology and Immunology. 133. 1987. 5–17. doi:10.1007/978-3-642-71683-6_2. ISBN 978-3-642-71685-0.
- ↑ "Fine structure analysis of Pichinde virus nucleocapsids". J. Gen. Virol. 64 (4): 833–42. April 1983. doi:10.1099/0022-1317-64-4-833. PMID 6682139.
- ↑ "The circular, segmented nucleocapsid of an arenavirus-Tacaribe virus". J. Gen. Virol. 36 (3): 541–5. September 1977. doi:10.1099/0022-1317-36-3-541. PMID 199698.
- ↑ "Characterization of the genomic promoter of the prototypic arenavirus lymphocytic choriomeningitis virus". J. Virol. 77 (2): 1184–94. January 2003. doi:10.1128/jvi.77.2.1184-1194.2003. PMID 12502835.
- ↑ "Molecular organization of Junin virus S RNA: complete nucleotide sequence, relationship with other members of the Arenaviridae and unusual secondary structures". J. Gen. Virol. 72 (9): 2129–41. September 1991. doi:10.1099/0022-1317-72-9-2129. PMID 1654373.
- ↑ "The 5' region of Tacaribe virus L RNA encodes a protein with a potential metal binding domain". Virology 173 (1): 357–61. November 1989. doi:10.1016/0042-6822(89)90257-2. PMID 2510403.
- ↑ "Tacaribe virus L gene encodes a protein of 2210 amino acid residues". Virology 170 (1): 40–7. May 1989. doi:10.1016/0042-6822(89)90349-8. PMID 2718387.
- ↑ "The completed sequence of lymphocytic choriomeningitis virus reveals a unique RNA structure and a gene for a zinc finger protein". Virology 173 (1): 1–10. November 1989. doi:10.1016/0042-6822(89)90216-X. PMID 2510401.
- ↑ "Uncovering viral protein-protein interactions and their role in arenavirus life cycle". Viruses 4 (9): 1651–67. September 2012. doi:10.3390/v4091651. PMID 23170177.
- ↑ "Envelope glycoprotein of arenaviruses". Viruses 4 (10): 2162–81. October 2012. doi:10.3390/v4102162. PMID 23202458.
- ↑ ICTV proposals 2014.011a-dV et al. , Mark D. Stenglein et al.
- ↑ ICTV proposals 2014.012aV et al. , Michael J. Buchmeier et al.
- ↑ "Identification, characterization, and in vitro culture of highly divergent arenaviruses from boa constrictors and annulated tree boas: candidate etiological agents for snake inclusion body disease". mBio 3 (4): e00180–12. 2012. doi:10.1128/mBio.00180-12. PMID 22893382.
- ↑ Hepojoki, Jussi; Hepojoki, Satu; Smura, Teemu; Szirovicza, Leonóra; Dervas, Eva; Prähauser, Barbara; Nufer, Lisbeth; Schraner, Elisabeth M. et al. (2018). "Characterization of Haartman Institute snake virus-1 (HISV-1) and HISV-like viruses—The representatives of genus Hartmanivirus, family Arenaviridae". PLOS Pathogens 14 (11): e1007415. doi:10.1371/journal.ppat.1007415. PMID 30427944.
- ↑ "Virus Taxonomy: 2019 Release". International Committee on Taxonomy of Viruses. https://ictv.global/taxonomy.
- ↑ Zhang YZ, Wu WC, Shi M, Holmes EC (2018) The diversity, evolution and origins of vertebrate RNA viruses. Curr Opin Virol 31:9-16
- ↑ Mordecai, GJ; Miller, KM; Di Cicco, E; Schulze, AD; Kaukinen, KH; Ming, TJ; Li, S; Tabata, A et al. (2019). "Endangered wild salmon infected by newly discovered viruses". eLife 8: e47615. doi:10.7554/eLife.47615. PMID 31478480.
- ↑ "Chapare virus, a newly discovered arenavirus isolated from a fatal hemorrhagic fever case in Bolivia". PLOS Pathog. 4 (4): e1000047. April 2008. doi:10.1371/journal.ppat.1000047. PMID 18421377.
- ↑ 38.0 38.1 Forni, D; Pontremoli, C; Pozzoli, U; Clerici, M; Cagliani, R; Sironi, M (2018-03-01). "Ancient evolution of Mammarenaviruses: Adaptation via changes in the L protein and no evidence for host-virus codivergence". Genome Biology and Evolution 10 (3): 863–874. doi:10.1093/gbe/evy050. PMID 29608723. PMC 5863214. https://europepmc.org/abstract/pmc/pmc5863214. Retrieved 2019-02-16.
- ↑ Paweska, Janusz T.; Sewlall, Nivesh H.; Ksiazek, Thomas G.; Blumberg, Lucille H.; Hale, Martin J.; Lipkin, W. Ian; Weyer, Jacqueline; Nichol, Stuart T. et al. (October 2009). "Nosocomial Outbreak of Novel Arenavirus Infection, Southern Africa". Emerging Infectious Diseases 15 (10): 1598–1602. doi:10.3201/eid1510.090211. PMID 19861052.
- ↑ Scientists identify new lethal virus in Africa
- ↑ "Three women had organ transplants from one donor. They all died in the same week" (in en-AU). ABC News. 2020-10-10. https://www.abc.net.au/news/health/2020-10-11/nobody-knew-karens-donor-kidney-was-carrying-a-rare-virus/12743548.
- ↑ "Virus identified - nurse ill". News24.com. http://www.news24.com/News24/South_Africa/News/0,,2-7-1442_2408685,00.html.
- ↑ "Virus kills organ recipients". www.theage.com.au. http://www.theage.com.au/victoria/virus-kills-organ-recipients-20100915-15cn8.html.
- ↑ 44.0 44.1 Lee, A. M.; Pasquato, A.; Kunz, S. (2011). "Novel approaches in anti-arenaviral drug development". Virology 411 (2): 163–169. doi:10.1016/j.virol.2010.11.022. PMID 21183197.
- ↑ Mendenhall, M.; Russell, A.; Juelich, T.; Messina, E. L.; Smee, D. F.; Freiberg, A. N.; Holbrook, M. R.; Furuta, Y. et al. (2010). "T-705 (Favipiravir) Inhibition of Arenavirus Replication in Cell Culture". Antimicrobial Agents and Chemotherapy 55 (2): 782–787. doi:10.1128/AAC.01219-10. PMID 21115797.
- ↑ Hadas Cohen-Dvashi; Ron Amon; Krystle N. Agans; Robert W. Cross; Aliza Borenstein-Katz; Mathieu Mateo; Sylvain Baize; Vered Padler-Karavani et al. (2020). "Rational design of universal immunotherapy for TfR1-tropic arenaviruses". Nature Communications 11 (1): 67. doi:10.1038/s41467-019-13924-6. PMID 31900422. Bibcode: 2020NatCo..11...67C.
External links
- ICTV Report: Arenaviridae
- Viralzone: Arenavirus
- Virus Pathogen Database and Analysis Resource (ViPR): Arenaviridae
- Detailed genomic and bioinformatic information about Arenaviridae at NIH-funded database.
- Arenaviridae Genomes database search results from the Viral Bioinformatics Resource Center.
- Google.Org blog info on recent outbreak.
- Arenaviruses
- "Arenavirus". NCBI Taxonomy Browser. https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&id=11618.
Wikidata ☰ Q2368907 entry
de:Arenaviridae
 |
