Biology:Bifidobacteriaceae
The Bifidobacteriaceae are the only family of bacteria in the order Bifidobacteriales.[1]
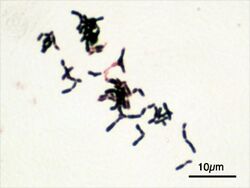
The family Bifidobacteriaceae stain [Gram-positive], range from obligate to faculatively anaerobic, are non-motile, non-filamentous and non-spore forming.[2] Their morphology is varied and ranges from Y- or V-shaped (from which the bifidobacteria derived their name) to ones with enlarged or flattened ends (club- or spatula-shaped).[3] The branching nature of Bifidobacteria can change with different starins and media.[4] These rods appear as a solitary bacilli or as aggregates in chains or in clumps.[2]
These chemoorganotrophic microorganisms are saccharolytic acid producers and do not produce gas.[2]
The Bifidobacteriaceae family is divided into ten genera ( Bifidobacterium, Aeriscardovia, Alloscardovia, Bombiscardovia, Galliscardovia, Gardnerella, Neoscardovia, Parascardovia, Pseudoscardovia, and Scardovia, [5] with three candidate genera Candidatus Ancillula, Candidatus Opitulatrix, and Candidatus Servula.[6]
Genomics
All Bifidobacteraceae contain a metabolic pathway to catabolise six-carbon sugars (hexoses) which involves the key enzyme fructose-6-phosphate phosphoketolase. This pathway is known as the fructose-6-phosphate pathway or the 'bifid shunt'.[2][7][8]
Comparative analysis of aligned protein sequences has led to the discovery of two conserved signature indels which are specific for the order Bifidobacteriales. The first indel, a one amino acid deletion in ribosomal protein L13, is found in all Bifidobacteriales species and no other Actinomycetota, providing a potential molecular marker for the entire Bifidobacteriales order. The second indel that has been identified is a 1 amino acid insertion in glucose-6-phosphate dehydrogenase found in all Bifidobacterium species and G. vaginalis, but not in any other Actinomycetota. This indel is thus characteristic of the clade consisting of Bifidobacterium species and G. vaginalis and can be used to distinguish these genera from the rest of the order Bifidobacteriales. 16 conserved signature proteins have also been identified which are unique to the order Bifidobacteriales and can be used as molecular markers for this order. Additionally, 6 conserved signature proteins which are unique to Bifidobacterium and Gardnerella have been identified, providing further evidence that species from these two genera are closely related and providing molecular markers for the clade consisting of these genera.[9]
| Bifidobacteriaceae | |
|---|---|
| |
| Bifidobacterium adolescentis | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Actinomycetota |
| Class: | Actinomycetia |
| Order: | Bifidobacteriales Stackebrandt et al. 1997[10] |
| Family: | Bifidobacteriaceae Stackebrandt et al. 1997[10] |
| Type genus | |
| Bifidobacterium Orla-Jensen 1924 (Approved Lists 1980)
| |
| Genera[11] | |
| |
The Bifidobacteriaceae are the only family of bacteria in the order Bifidobacteriales.[12]
The Bifidobacteriaceae family is divided into ten genera ( Bifidobacterium, Aeriscardovia, Alloscardovia, Bombiscardovia, Galliscardovia,Gardnerella, Neoscardovia, Parascardovia, Pseudoscardovia and Scardovia [13] with three candidate genera Candidatus Ancillula, Candidatus Opitulatrix, and Candidatus Servula.[14]
Genomics
All Bifidobacteraceae contain a peculiar metabolic pathway to catabolise six-carbon sugars (hexoses) involving the key enzyme Fructose-6-phosphate phosphoketolase (EC 4.1.2.2), known as the fructose-6-phosphate pathway or the 'bifid shunt'.[2]
Comparative analysis of aligned protein sequences has led to the discovery of two conserved signature indels which are specific for the order Bifidobacteriales. The first indel, a 1 amino acid deletion in ribosomal protein L13, is found in all Bifidobacteriales species and no other Actinomycetota, providing a potential molecular marker for the entire Bifidobacteriales order. The second indel that has been identified is a 1 amino acid insertion in glucose-6-phosphate dehydrogenase found in all Bifidobacterium species and G. vaginalis, but not in any other Actinomycetota. This indel is thus characteristic of the clade consisting of Bifidobacterium species and G. vaginalis and can be used to distinguish these genera from the rest of the order Bifidobacteriales. 16 conserved signature proteins have also been identified which are unique to the order Bifidobacteriales and can be used as molecular markers for this order. Additionally, 6 conserved signature proteins which are unique to Bifidobacterium and Gardnerella have been identified, providing further evidence that species from these two genera are closely related and providing molecular markers for the clade consisting of these genera.[15]
Phylogeny
The currently accepted taxonomy is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN).[11] The phylogeny is based on whole-genome analysis.[16]
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| outgroup | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Notes
References
- ↑ "Association between Bifidobacteriaceae and the clinical severity of root caries lesions". Oral Microbiol. Immunol. 24 (1): 32–7. February 2009. PMID 19121067.
- ↑ 2.0 2.1 2.2 2.3 2.4 D.A. Russell, R.P. Ross, G.F. Fitzgerald, C. Stanton (2011). "Metabolic activities and probiotic potential of bifidobacteria". International Journal of Food Microbiology 149 (1): 88-105. doi:10.1016/j.ijfoodmicro.2011.06.003. ISSN 0168-1605.
- ↑ Alessandri G, van Sinderen D, Ventura M. (2021). "The genus bifidobacterium: From genomics to functionality of an important component of the mammalian gut microbiota running title: Bifidobacterial adaptation to and interaction with the host.". Comput Struct Biotechnol J. 9 (19): 1472-1487. doi:10.1016/j.csbj.2021.03.006. PMID 33777340.
- ↑ {{cite journal|author=G.W. Tannock|title= Identification of lactobacilli and bifidobacteria|journal= Current Issues in Molecular Biology|volume= 1|year= 1999|page=53-64
- ↑ Ludwig, W., Euzéby, J.P. and Whitman, W.B. (2015). M.E. Trujillo, S. Dedysh, P. DeVos, B. Hedlund, P. Kämpfer, F.A. Rainey and W.B. Whitman. ed. "Taxonomic outline of the phylum Actinobacteria". Bergey's Manual of Systematics of Archaea and Bacteria.
- ↑ Silva JK, Herve V, Mies US, Platt K, Brune A. (2025). "A Novel Lineage of Endosymbiotic Actinomycetales: Genome Reduction and Acquisition of New Functions in Bifidobacteriaceae Associated With Termite Gut Flagellates.". Environ Microbiol 27.
- ↑ https://pubchem.ncbi.nlm.nih.gov/pathway/BioCyc:META_P124-PWY
- ↑ Turroni F, Milani C, Duranti S, Mahony J, van Sinderen D, Ventura M. (2018). "Glycan Utilization and Cross-Feeding Activities by Bifidobacteria.". Trends Microbiol. 26 (4): 339-350.. doi:10.1016/j.tim.2017.10.001.
- ↑ Gao, B.; Gupta, R. S. (2012). "Phylogenetic Framework and Molecular Signatures for the Main Clades of the Phylum Actinobacteria". Microbiology and Molecular Biology Reviews 76 (1): 66–112. doi:10.1128/MMBR.05011-11. PMID 22390973.
- ↑ 10.0 10.1 "Proposal for a new hierarchic classification system, Actinobacteria classis nov.". Int. J. Syst. Bacteriol. 47 (2): 479–491. 1997. doi:10.1099/00207713-47-2-479.
- ↑ 11.0 11.1 "Bifidobacteriaceae". List of Prokaryotic names with Standing in Nomenclature (LPSN). https://lpsn.dsmz.de/family/bifidobacteriaceae.
- ↑ "Association between Bifidobacteriaceae and the clinical severity of root caries lesions". Oral Microbiol. Immunol. 24 (1): 32–7. February 2009. doi:10.1111/j.1399-302X.2008.00470.x. PMID 19121067.
- ↑ Ludwig, W., Euzéby, J.P. and Whitman, W.B. (2015). M.E. Trujillo, S. Dedysh, P. DeVos, B. Hedlund, P. Kämpfer, F.A. Rainey and W.B. Whitman. ed. "Taxonomic outline of the phylum Actinobacteria". Bergey's Manual of Systematics of Archaea and Bacteria.
- ↑ Silva JK, Herve V, Mies US, Platt K, Brune A. (2025). "A Novel Lineage of Endosymbiotic Actinomycetales: Genome Reduction and Acquisition of New Functions in Bifidobacteriaceae Associated With Termite Gut Flagellates.". Environ Microbiol 27.
- ↑ Gao, B.; Gupta, R. S. (2012). "Phylogenetic Framework and Molecular Signatures for the Main Clades of the Phylum Actinobacteria". Microbiology and Molecular Biology Reviews 76 (1): 66–112. doi:10.1128/MMBR.05011-11. PMID 22390973.
- ↑ "Genome-Based Taxonomic Classification of the Phylum Actinobacteria". Front. Microbiol. 9: 2007. 2018. doi:10.3389/fmicb.2018.02007. PMID 30186281.
External links
| Wikispecies has information related to Bifidobacteriaceae |
Wikidata ☰ Q4904765 entry
Notes
References
External links
| Wikispecies has information related to Bifidobacteriaceae |
Wikidata ☰ Q4904765 entry
 |
