Biology:Clostridioides difficile
| Clostridioides difficile | |
|---|---|
| |
| C. difficile colonies on a blood agar plate | |

| |
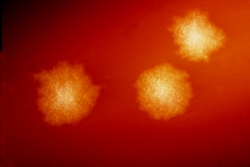
| Electron micrograph of the bacterium | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Bacillota |
| Class: | Clostridia |
| Order: | Eubacteriales |
| Family: | Peptostreptococcaceae |
| Genus: | Clostridioides |
| Species: | C. difficile
|
| Binomial name | |
| Clostridioides difficile (Hall & O'Toole, 1935) Lawson & Rainey, 2016
| |
| Synonyms | |
Clostridioides difficile (syn. Clostridium difficile) is a bacterium known for causing serious diarrheal infections, and may also cause colon cancer.[4][5] It is known also as C. difficile, or C. diff (/siː dɪf/), and is a Gram-positive species of spore-forming bacteria.[6] Clostridioides spp. are anaerobic, motile bacteria, ubiquitous in nature and especially prevalent in soil. Its vegetative cells are rod-shaped, pleomorphic, and occur in pairs or short chains. Under the microscope, they appear as long, irregular (often drumstick- or spindle-shaped) cells with a bulge at their terminal ends (forms subterminal spores). C. difficile cells show optimum growth on blood agar at human body temperatures in the absence of oxygen. C. difficile is catalase- and superoxide dismutase-negative, and produces up to three types of toxins: enterotoxin A, cytotoxin B and Clostridioides difficile transferase.[7] Under stress conditions, the bacteria produce spores that tolerate extreme conditions that the active bacteria cannot tolerate.[8]
Clostridioides difficile is an important human pathogen; according to the CDC, in 2017 there were 223,900 cases in hospitalized patients and 12,800 deaths in the United States.[9] Although C. difficile is known as a hospital- and antibiotic-associated pathogen, at most one third of infections can be traced to transmission from an infected person in hospitals,[10] and only a small number of antibiotics are directly associated with an elevated risk of developing a C. difficile infection (CDI), namely vancomycin, clindamycin, fluoroquinolones and cephalosporins.[11][12][13] Most infections are acquired outside of hospitals, and most antibiotics have similar elevated risk of infection on par with many non-antibiotic risk factors, such as using stool softeners and receiving an enema.[14]
Clostridioides difficile can become established in the human colon without causing disease.[15] Although early estimates indicated that C. difficile was present in 2–5% of the adult population,[8] later research indicated that colonization is closely associated with a history of unrelated diarrheal illnesses, such as food poisoning or laxative abuse.[16] Individuals with no history of gastrointestinal disturbances appear unlikely to become asymptomatic carriers. These carriers are thought to be a major infection reservoir.[17]
Taxonomy
The species was transferred from the genus Clostridium to Clostridioides in 2016, thus giving it the binomial Clostridioides difficile.[18][19][20] This new name reflects the taxonomic differences between this species and members of the genus Clostridium, while maintaining the common name as C. difficile .[19] As of 2018[update], the only other species in this new genus is Clostridioides mangenotii (formerly known as Clostridium mangenotii).[21]
Human pathogen
Pathogenic C. difficile strains produce multiple toxins.[22] The best-characterized are enterotoxin (C. difficile toxin A) and cytotoxin (C. difficile toxin B), both of which may produce diarrhea and inflammation in infected patients (C. difficile colitis), although their relative contributions have been debated. The diarrhea may range from a few days of intestinal fluid loss to life-threatening pseudomembranous colitis, which is associated with intense colon inflammation and pseudomembrane formation on the intestinal mucosal surface.[8] This may progress to toxic megacolon, a severe form of colonic distention that can put a patient at risk for colon perforation, sepsis and shock. Toxins A and B are glucosyltransferases that target and inactivate the Rho family of GTPases. Toxin B (cytotoxin) induces actin depolymerization by a mechanism correlated with a decrease in the ADP-ribosylation of the low molecular mass GTP-binding Rho proteins.[23] A binary toxin (AB toxin), but its role in disease is not fully understood.[24]
Additional virulence factors include an adhesion factor that mediates the binding to human colonic cells and a hyaluronidase.[25] The bacterium also produces the chemical para-cresol, which inhibits the growth of other microbes in its vicinity and allows it to outcompete normal human gut flora.[26]
Antibiotic resistance
Antibiotic treatment of C. difficile infections may be difficult, due both to antibiotic resistance and physiological factors of the bacterium (spore formation, protective effects of the pseudomembrane).[8] A highly toxic strain of C. difficile, resistant to fluoroquinolone antibiotics, such as ciprofloxacin and levofloxacin, said to be causing geographically dispersed outbreaks in North America, was reported in 2005.[27] In 2005 the US Centers for Disease Control in Atlanta warned of an epidemic strain with increased virulence, antibiotic resistance, or both.[28] In 2018, resistance to other antibiotics such as metronidazole, the first choice of antimicrobial drug when treating CDI, was observed in up to 12% of clinical isolates. While treatment with various antibiotics continues, more diverse and stronger resistance will continue in C. difficile populations.[29]
Clostridioides difficile spores resist many disinfectants,[30] including high concentrations of bleach.[31]
A recent study published by researchers in Nature Communications found that a chain mail-like armor, called "The S-layer" may be responsible for C. difficile's apparent immunity to antibiotic treatments, human saliva, as well as an enzyme human host cells normally use to fight viruses.
The study also found that removing a region of the S-layer called D2 made C. diff cells susceptible to lysozyme, an enzyme typically found in saliva that tears open microbes' exteriors. Without the protection layer, the bacteria would also be vulnerable to the means of attacking the virus mentioned above.[32]
Persistence in the gut
Beyond spore formation and toxin secretion, an important virulence factor of Clostridioides difficile is its set of cell-surface proteins.[33] These proteins are located on or beyond the bacterial peptidoglycan layer and include the surface (S-) layer, flagella, pili, and approximately 28 accessory cell-wall proteins (CWPs).[33] Cell-surface proteins in C. difficile have been associated with several functions, including adhesion to host cells, biofilm formation, surface interactions, and motility.[33][34][35]
The S-layer is the most abundant and functionally significant cell-surface component.[33][34] Unlike in many other prokaryotes, the C. difficile S-layer is heterodimeric, consisting of low- and high-molecular-weight S-layer proteins.[34] The S-layer contributes to maintaining cell-envelope stability and has been implicated in several virulence-related processes, including spore and toxin production, as well as resistance to antibiotics and environmental stress.[35]
Research
Deletion of the slpA gene, which encodes the S-layer protein in Clostridioides difficile, results in marked changes and impairments in both cell and colony morphology.[35] The mutant strain also exhibits reduced ability to withstand stress during the late stationary phase and shows approximately double the level of biofilm formation compared to the wild type.[35] However, loss of the S-layer protein increases bacterial susceptibility to the antibiotic vancomycin and to environmental stress induced by Triton X-100.[35] The absence of the S-layer further decreases sporulation efficiency, toxin release, and adherence capacity.[35]
Transmission

Clostridioides difficile is transmitted from person or animal to person by the fecal-oral route, shed in faeces. The organism forms heat-resistant aero-tolerant spores that are resistant to alcohol-based hand cleansers or routine surface cleaning. Spores survive in clinical environments for long periods.[36] Any surface, device, or material (e.g., toilets, bathing tubs, or rectal thermometers) that becomes contaminated with feces may serve as a reservoir for C. difficile spores, which can live for long periods of time on surfaces.[37] Because of this, the bacterium may be cultured from almost any surface. Once spores are ingested, their acid resistance allows them to pass through the stomach unscathed. They germinate and multiply into vegetative cells in the colon upon exposure to bile acids. Consequently, the World Health Organization advocates the use of soap in addition to alcohol solutions to limit the spread of the spores.[38] Sporulation was shown to be significantly reduced after inactivation of C. difficile's DNA methyltransferase CamA,[39] raising the prospect of developing a drug that may inhibit this bacterium in a specific manner.
Host range
Clostridioides difficile infects pigs, calves, and humans, and inhabits a natural reservoir of soil, faeces of domestic animals and humans, sewage, the human intestinal tract, and retail meat.[40]
A 2015 CDC study estimated that C. difficile afflicted almost half a million Americans and caused 29,000 deaths in 2011. The study estimated that 40% of cases began in nursing homes or community health-care settings, while 24% occurred in hospitals.[41]
Clostridioides difficile is common in the human digestive system. However, it is often outcompeted for nutrients by other bacteria, limiting its spread. If the sudden introduction of an antibiotic disrupts the microbiome, C. difficile may be able to grow as many of its competitors die. The incubation period is 5–10 days, with a range of 1 day to weeks following antibiotic treatment. Additionally, carriage of C. difficile with high levels of toxins is common in young children, while disease is rare. The production of one or even both toxins is not always sufficient for producing symptoms.[42]
Signs and symptoms
Symptoms of C. difficile infection include diarrhea (at least three loose bowel movements a day), dehydration, abdominal pain that can be severe, loss of appetite, and nausea.[43]
Pathophysiology
Clostridioides difficile is transmitted through the oral-fecal route, and many reproduce through spores. Spore germination depends on the ability to sense primary bile acids in the liver, like taurocholate, which are sensed by the germinant receptor CspC. Secondary bile acids can inhibit these processes in the colon. Spores can grow and colonize the intestine by antibiotic-induced shifts in the host microbiota. C. difficile secretes mucolytic enzymes like CWp84 to degrade the colonic mucosa. These spores can adhere to colon cells. Additionally, C. difficile is motile and can switch between motile and sessile phases, a process regulated by cyclic-di-GMP. C. difficile is also capable of forming biofilms and cell-to-cell signaling.[44] C. difficile is often transferred via the hands of healthcare workers or the overall hospital environment and acquired by ingesting the pathogen. The spores resist the stomach acidity and germinate into their vegetative form in the small intestine. C. difficile can carry a broad spectrum of clinical manifestations, from asymptomatic to severe colitis and death.[45] C. difficile is the most prevalent US healthcare infection, posing serious health risks and substantial care costs.[46]
Immune response
The C. difficile secreted toxins A (TcdA) and B (TcdB) contain immunogenic antigens that are recognised by antibodies and T cells. However, the levels of anti-TcdA and -TcdB IgG antibodies have not been able to discriminate healthy individuals from patients with C. difficile infection, meaning they have limited clinical use.[47][48] Recent work has shown these toxins are also recognised by helper CD4+ T cells, predominantly by the Th17 helper cells, which are important in maintaining a healthy gut environment, although in patients with severe infection these cells are impaired.[49] Interestingly, individuals with severe C. difficile infection had significantly more toxin-specific T cells compared to those with mild infection, indicating T cells are playing a key role in fighting this infection. This immune response can further dysregulate microRNA expression.[50] This is further evidenced by the recovery of the toxin-specific Th17 cells and microRNA expression following fecal microbiota transplant of patients with severe disease.[50][51] New findings show that the loss of the interleukin-10 corresponds to higher levels of interleukin-22, which has been found to be important in a host's response to a C. difficile infection. Thus, IL-10 deficiency can increase a host's defense against the pathogen. This could be of particular interest in future research for treatments.[52]
Possible role in colon cancer
Studies indicate that C. difficile infection may contribute to colorectal cancer development through mechanisms such as chronic inflammation and epithelial barrier disruption.[32] Toxigenic C. difficile strains isolated from human colon tumors have been shown to drive colonic tumorigenesis in mice, with tumor growth requiring the TcdB and accompanied by activation of Wnt/β-catenin signaling and inflammatory immune responses.[53] C. difficile toxins can also induce cellular senescence; TcdB-triggered senescent cells may promote neoplastic transformation via the senescence-associated secretory phenotype.[54] In addition, C. difficile releases membrane vesicles that directly stimulate epithelial–mesenchymal transition in colonic epithelial cells, as evidenced by upregulation of β-catenin and mesenchymal markers with loss of E-cadherin in both cell cultures and mouse models, thus creating a pro-tumorigenic microenvironment in the colon.[55]
Diagnosis
An infection with C. difficile is often indicated by foul-smelling diarrhea, an active or recent treatment with antibiotics can also point to Clostridium difficile associated diarrhea (CDAD). Clinical diagnosis however requires either the presence of the main toxins in the stool or direct cultivation of C. difficile . To confirm a CDI, a cytotoxin assay detects the cell's toxin B (ToxB) cytotoxicity in the fecal eluate. The presence of C. difficile toxin is confirmed by the anti-toxin antibodies' neutralization of the cytotoxic effect. C. difficile strains can also be cultured before conducting a cytotoxin assay. These cultures detect the C. difficile strain that can produce toxins.[56] However, these enzyme immunoassays are more widely used due to their rapid turnaround, low cost, and simplicity. Additionally, they show lower sensitivity than toxigenic stool cultures. PCR assays have a shorter turnaround time and a higher sensitivity range than the toxigenic stool culture. Using a PCR-based assay helps avoid detection of asymptomatic patients.[45]
Treatment
Patients being treated with antibiotics when symptoms begin should stop taking them, if possible. This break in antibiotic therapy can sometimes lead to spontaneous resolution of symptoms. Patients who do not respond to the cessation of broad-spectrum antibiotics will need to be treated with antibiotics capable of killing C. difficile spores. Primary infections are typically treated with vancomycin, with a usual dosage of 125 mg every 6 hours.[57] The vancomycin regimen has replaced the traditional use of metronidazole due to its greater efficacy, safety profile, and lower recurrence rates. In patients who cannot tolerate vancomycin, fidaxomicin is an acceptable option with similar efficacy and even lower recurrence rates than vancomycin.[58] In cases of fulminant CDI, adjuvant therapy with parenteral metronidazole plus oral vancomycin or fidaxomicin is suggested.[59]
Approximately 15-30%[60] of patients who successfully complete therapy of primary infection with metronidazole or vancomycin will experience a relapse. About 40% of these patients will continue to have recurrent C. difficile infection. The first relapse of C. difficile is usually treated with the same antibiotic used to treat the primary infection. Any subsequent infections should not be treated with metronidazole. Occasionally, a standard 10-day course of oral vancomycin will not work. In these cases, a vancomycin taper is the preferred treatment. Patients take decreasing doses of vancomycin over a period of up to 3 months, depending on the severity of the infection.[43]
Each subsequent relapse of C. difficile tends to be more severe than previous infections. Long-term treatment with a vancomycin taper supplemented with probiotics, especially Saccharomyces boulardii, is associated with a higher rate of success.[61]
After three relapses, patients may be treated with oral fidaxomicin, a narrow-spectrum antibiotic. The usual dosage is 200 mg twice a day orally for 10 days. Fidaxomicin is considered to be superior to vancomycin for severe CDI.[62] The major downside of treatment with fidaxomicin is the cost of medication. A 10-day course may cost up to US$3500. When a patient is deteriorating or progressing to severe-complicated disease the addition of intravenous tigecycline merits considerations.[63][64] Patients with high risk of relapse may also benefit from the addition of the monoclonal antibody bezlotoxumab to the standard of care.[65]
Patients who do not respond to traditional antibiotic therapy may be eligible for a fecal microbiota transplant (FMT). Healthcare providers can transfer stool from a healthy person to the colon of a patient with recurrent CDI. This process is the most successful treatment for severe CDI with a cure rate around 93%. Fecal matter transplants have also been found to be an effective and safe treatment option for children and young adults.[66] Recurrence rates of CDI in patients treated with a FMT are generally low, around 19%, which makes it very effective at treating chronic CDI cases. However, in some cases, flares of inflammatory bowel disease are a possible side effect of the treatment.[67] The state of the host immune system is important when considering the success of microbiota-based treatments in clearing infection.[68] Long-term effects of FMT are unknown, as the procedure has only been FDA-approved for recurrent CDI since 2013 and relatively few procedures have been performed. If transplantation is not an option, removal of the infected part of the colon can cure CDI.[62][43]
In April 2023, the FDA approved the first oral microbiome therapeutic, VOWST for treatment of recurrent CDI.
The prediction of C. difficile recurrence has been of great interest, but there has been no consensus on significantly associated risk factors.[69]
Prevention
C. difficile infection is spread through the fecal-oral route through ingestion and acid-resistant spores. Appropriate hand hygiene of healthcare workers is vital to remove spores, which includes thoroughly washing one's hands with soap and warm water. Additionally, isolation of patients with acute diarrhea can prevent the spread of spores within the hospital.[70] Another way to prevent CDIs is wearing personal protective equipment when interacting with C. difficile patients. Furthermore, CDI transmission can be prevented by daily environmental sporicidal disinfection in patient rooms. Also, reducing the length of antibiotic therapy decreases the CDI rates in hospitals.[71]
Strains
In 2005, molecular analysis led to the identification of the C. difficile strain type characterized as group BI by restriction endonuclease analysis, as North American pulse-field-type NAP1 by pulsed-field gel electrophoresis and as ribotype 027; the difficileering terminology reflects the predominant techniques used for epidemiological typing. This strain is referred to as C. difficile BI/NAP1/027.[56]
As of 2016, the NAP1 strain has been replaced by novel strains in some areas of British Columbia. These novel strains include NAP2 and NAP4, and some strains that do not have a NAP designation. The frequency of these novel strains increased from 2008 to 2013 in one studied region, displacing the originally more common and recognizable NAP1 bacteria.[72]
Two strains, ribotypes RT078 and RT027, can live on low concentrations of the sugar trehalose; both strains became more common after trehalose was introduced as a food additive in the early 2000s, thus increasing dietary trehalose intake.[73]
Genome
| NCBI genome ID | 535 |
|---|---|
| Ploidy | haploid |
| Genome size | 4.3 Mb |
| Number of chromosomes | 1 |
| Year of completion | 2005 |
The first complete genome sequence of a C. difficile strain was published in 2005 by the Sanger Institute in the UK.[74] This was of strain 630, a virulent and multiple drug-resistant strain isolated in Switzerland in 1982. By 2010 scientists at the Sanger Institute had sequenced genomes of about 30 C. difficile isolates using next-generation sequencing technologies from 454 Life Sciences and Illumina.[75]
Researchers at McGill University in Montreal sequenced the genome of the highly virulent Quebec strain of C. difficile in 2005 using ultra-high throughput sequencing technology. The tests involved doing 400,000 DNA parallel-sequencing reactions of the bacterium's genome, which had been fragmented for sequencing. These sequences were assembled computationally to form a complete genome sequence.[27][76]
In 2012, scientists at University of Oxford sequenced C. difficile genomes from 486 cases arising over four years in Oxfordshire using next-generation sequencing technologies from Illumina.[77]
Epigenome
Clostridioides difficile has a highly diverse epigenome, with 17 high-quality methylation motifs reported so far, the majority pertaining to the 6mA type. Methylation at one of these motifs - CAAAAA, was shown to impact sporulation, a key step in C. difficile disease transmission, as well as cell length, biofilm formation, and host colonization.[39]
Bacteriophage
At least eight mainly temperate bacteriophages have been isolated from C. difficile, ranging in genome size from about 30 to about 60 kbp.[78] Both environmentally and clinically derived C. difficile strains carry a diverse and prevalent set of prophages.[78]
Etymology and pronunciation
References
- ↑ "Intestinal flora in new-born infants: with a description of a new pathogenic anaerobe, Bacillus difficilis". American Journal of Diseases of Children 49 (2): 390–402. 1935. doi:10.1001/archpedi.1935.01970020105010.
- ↑ "Études de systématique bactérienne. IV. Critique de la conception actuelle du genre Clostridium". Annales de l'Institut Pasteur 61 (1): 84. 1938. https://gallica.bnf.fr/ark:/12148/bpt6k5846341v/f79.item. Retrieved December 15, 2018.
- ↑ Page Species: Clostridioides difficile on "LPSN - List of Prokaryotic names with Standing in Nomenclature". Deutsche Sammlung von Mikroorganismen und Zellkulturen. https://lpsn.dsmz.de/species/clostridioides-difficile.
- ↑ "Human Colon Cancer-Derived Clostridioides difficile Strains Drive Colonic Tumorigenesis in Mice". Cancer Discovery 12 (8): 1873–1885. August 2022. doi:10.1158/2159-8290.CD-21-1273. PMID 35678528.
- ↑ "Johns Hopkins Doctors Discover That a Common Infection May Cause Cancer". August 24, 2022. https://scitechdaily.com/johns-hopkins-doctors-discover-that-a-common-infection-may-cause-cancer/.
- ↑ "Clostridium difficile: a cause of diarrhea in children". JAMA Pediatrics 167 (6): 592. June 2013. doi:10.1001/jamapediatrics.2013.2551. PMID 23733223.
- ↑ "Clostridioides difficile toxins: mechanisms of action and antitoxin therapeutics". Nature Reviews. Microbiology 20 (5): 285–298. May 2022. doi:10.1038/s41579-021-00660-2. PMID 34837014.
- ↑ 8.0 8.1 8.2 8.3 Sherris Medical Microbiology (4th ed.). McGraw Hill. 2004. pp. 322–4. ISBN 978-0-8385-8529-0. https://archive.org/details/sherrismedicalmi00ryan.
- ↑ "Clostridioides difficile Infection | HAI | CDC" (in en-us). January 2, 2020. https://www.cdc.gov/hai/organisms/cdiff/cdiff_infect.html.
- ↑ "Diverse sources of C. difficile infection identified on whole-genome sequencing". The New England Journal of Medicine 369 (13): 1195–1205. September 2013. doi:10.1056/NEJMoa1216064. PMID 24066741.
- ↑ "Risk Factors for Community-Associated Clostridium difficile Infection in Adults: A Case-Control Study". Open Forum Infectious Diseases 4 (4). October 1, 2017. doi:10.1093/ofid/ofx171. PMID 29732377.
- ↑ "Meta-analysis of antibiotics and the risk of community-associated Clostridium difficile infection". Antimicrobial Agents and Chemotherapy 57 (5): 2326–2332. May 2013. doi:10.1128/AAC.02176-12. PMID 23478961.
- ↑ "Antibiotics Associated With Clostridium difficile Infection". Cureus 15 (5). May 2023. doi:10.7759/cureus.39029. PMID 37323360.
- ↑ "Risk factors for Clostridium difficile carriage and C. difficile-associated diarrhea in a cohort of hospitalized patients". The Journal of Infectious Diseases 162 (3): 678–684. September 1990. doi:10.1093/infdis/162.3.678. PMID 2387993.
- ↑ "Nosocomial acquisition of Clostridium difficile infection". The New England Journal of Medicine 320 (4): 204–210. January 1989. doi:10.1056/NEJM198901263200402. PMID 2911306.
- ↑ "Diarrhoeal events can trigger long-term Clostridium difficile colonization with recurrent blooms". Nature Microbiology 5 (4): 642–650. April 2020. doi:10.1038/s41564-020-0668-2. PMID 32042128.
- ↑ "Asymptomatic Clostridium difficile colonisation and onward transmission". PLOS ONE 8 (11). November 12, 2013. doi:10.1371/journal.pone.0078445. PMID 24265690. Bibcode: 2013PLoSO...878445E.
- ↑ "List of new names and new combinations previously effectively, but not validly, published". International Journal of Systematic and Evolutionary Microbiology 67 (9): 3140–3143. September 2017. doi:10.1099/ijsem.0.002278. PMID 28891789.
- ↑ 19.0 19.1 "Reclassification of Clostridium difficile as Clostridioides difficile (Hall and O'Toole 1935) Prévot 1938". Anaerobe 40: 95–99. August 2016. doi:10.1016/j.anaerobe.2016.06.008. PMID 27370902. Bibcode: 2016Anaer..40...95L.
- ↑ "Clostridioides difficile Biology: Sporulation, Germination, and Corresponding Therapies for C. difficile Infection". Frontiers in Cellular and Infection Microbiology 8. 2018. doi:10.3389/fcimb.2018.00029. PMID 29473021.
- ↑ "Phylogenomic analysis of the family Peptostreptococcaceae (Clostridium cluster XI) and proposal for reclassification of Clostridium litorale (Fendrich et al. 1991) and Eubacterium acidaminophilum (Zindel et al. 1989) as Peptoclostridium litorale gen. nov. comb. nov. and Peptoclostridium acidaminophilum comb. nov". International Journal of Systematic and Evolutionary Microbiology 66 (12): 5506–5513. December 2016. doi:10.1099/ijsem.0.001548. PMID 27902180.
- ↑ "Clostridium difficile Toxins A and B: Insights into Pathogenic Properties and Extraintestinal Effects". Toxins 8 (5): 134. May 2016. doi:10.3390/toxins8050134. PMID 27153087.
- ↑ "The low molecular mass GTP-binding protein Rho is affected by toxin A from Clostridium difficile". The Journal of Clinical Investigation 95 (3): 1026–1031. March 1995. doi:10.1172/JCI117747. PMID 7883950.
- ↑ "Binary bacterial toxins: biochemistry, biology, and applications of common Clostridium and Bacillus proteins". Microbiology and Molecular Biology Reviews 68 (3): 373–402, table of contents. September 2004. doi:10.1128/MMBR.68.3.373-402.2004. PMID 15353562. Bibcode: 2004MMBR...68..373B.
- ↑ [Medical Micriobiology, Fifth Edition, Patrick Murray, Elsevier Mosby, 2005, page 412]
- ↑ "The chemical weapon that helps bacterium wreak havoc in the gut". Nature 561 (7723): 288. September 14, 2018. doi:10.1038/d41586-018-06650-4. Bibcode: 2018Natur.561S.288.. https://www.nature.com/articles/d41586-018-06650-4. Retrieved October 8, 2018.
- ↑ 27.0 27.1 "A predominantly clonal multi-institutional outbreak of Clostridium difficile-associated diarrhea with high morbidity and mortality". The New England Journal of Medicine 353 (23): 2442–2449. December 2005. doi:10.1056/NEJMoa051639. PMID 16322602.
- ↑ "Clostridium difficile: responding to a new threat from an old enemy". Infection Control and Hospital Epidemiology 26 (8): 672–675. August 2005. doi:10.1086/502600. PMID 16156321.
- ↑ "Clostridium difficile Infections: A Global Overview of Drug Sensitivity and Resistance Mechanisms". BioMed Research International 2018. February 21, 2018. doi:10.1155/2018/8414257. PMID 29682562.
- ↑ "Activity of Hospital Disinfectants against Vegetative Cells and Spores of Clostridioides difficile Embedded in Biofilms". Antimicrobial Agents and Chemotherapy 64 (1). December 2019. doi:10.1128/AAC.01031-19. PMID 31611365.
- ↑ "Bleach does not kill common superbug, study finds". BBC News. November 22, 2023. https://www.bbc.com/news/articles/c4n8304nr5ko.
- ↑ 32.0 32.1 Nezhadi, Javad; Lahouty, Masoud; Rezaee, Mohammad Ahangarzadeh; Fadaee, Manouchehr (2025-05-24). "Clostridium difficile as a potent trigger of colorectal carcinogenesis" (in en). Discover Oncology 16 (1). doi:10.1007/s12672-025-02742-6. ISSN 2730-6011. PMID 40411629.
- ↑ 33.0 33.1 33.2 33.3 Chilton, Caroline H.; Viprey, Virginie; Normington, Charmaine; Moura, Ines B.; Buckley, Anthony M.; Freeman, Jane; Davies, Kerrie; Wilcox, Mark H. (2025). "Clostridioides difficile pathogenesis and control". Nature Reviews Microbiology. doi:10.1038/s41579-025-01242-2. PMID 41039149.
- ↑ 34.0 34.1 34.2 Bradshaw, William J.; Roberts, April K.; Shone, Clifford C.; Acharya, K. Ravi (2018). "The structure of the S-layer of Clostridium difficile". Journal of Cell Communication and Signaling 12 (1): 319–331. doi:10.1007/s12079-017-0429-z. PMID 29170885.
- ↑ 35.0 35.1 35.2 35.3 35.4 35.5 Wang, Shaohua; Courreges, Maria C.; Xu, Lingjun; Gurung, Bijay; Berryman, Mark; Gu, Tingyue (2024). "Revealing roles of S-layer protein (SlpA) in Clostridioides difficile pathogenicity by generating the first slpA gene deletion mutant". Microbiology Spectrum 12 (6). doi:10.1128/spectrum.04005-23. PMID 38709045.
- ↑ Di Bella, Stefano; Sanson, Gianfranco; Monticelli, Jacopo; Zerbato, Verena; Principe, Luigi; Giuffrè, Mauro; Pipitone, Giuseppe; Luzzati, Roberto (February 29, 2024). Staley, Christopher. ed. Mayuresh Abhyankar. "Clostridioides difficile infection: history, epidemiology, risk factors, prevention, clinical manifestations, treatment, and future options" (in en). Clinical Microbiology Reviews 37 (2): e0013523. doi:10.1128/cmr.00135-23. ISSN 0893-8512. PMID 38421181.
- ↑ "Clostridium difficile Infection Information for Patients". Health-care Associated Infections (HAI). U.S. Centers for Disease Control and Prevention. https://www.cdc.gov/hai/organisms/cdiff/cdiff-patient.html.
- ↑ "WHO Guidelines on Hand Hygiene in Health Care: a Summary". 2009. p. 31. https://www.who.int/gpsc/5may/tools/who_guidelines-handhygiene_summary.pdf.
- ↑ 39.0 39.1 "Epigenomic characterization of Clostridioides difficile finds a conserved DNA methyltransferase that mediates sporulation and pathogenesis". Nature Microbiology 5 (1): 166–180. January 2020. doi:10.1038/s41564-019-0613-4. PMID 31768029.
- ↑ "Clostridium difficile in food and domestic animals: a new foodborne pathogen?". Clinical Infectious Diseases 51 (5): 577–582. September 2010. doi:10.1086/655692. PMID 20642351.
- ↑ "Death Toll From C. Difficile Is Raised". The New York Times. February 25, 2015. https://www.nytimes.com/2015/02/26/us/death-toll-from-bacteria-is-raised.html?_r=0.
- ↑ Medical Microbiology (Fifth ed.). Elsevier Mosby. 2005. p. 412.
- ↑ 43.0 43.1 43.2 "Could you have deadly diarrhea (C. Diff)?". January 4, 2019. https://www.cdc.gov/hai/organisms/cdiff/cdiff-patient.html.
- ↑ "Clostridium difficile infection". Nature Reviews Disease Primers 2. April 7, 2017. doi:10.1038/nrdp.2016.20. PMID 27158839.
- ↑ 45.0 45.1 "Current Status of Clostridium difficile Infection Epidemiology". Clinical Infectious Diseases 55 (Suppl 2): S65–S70. August 1, 2012. doi:10.1093/cid/cis319. PMID 22752867.
- ↑ "Management of Clostridioides difficile infection: Diagnosis, Treatment, and Future Perspectives". Am J Med 137 (7): 571–576. March 2024. doi:10.1016/j.amjmed.2024.03.024. PMID 38508330.
- ↑ "High prevalence of subclass-specific binding and neutralizing antibodies against Clostridium difficile toxins in adult cystic fibrosis sera: possible mode of immunoprotection against symptomatic C. difficile infection". Clinical and Experimental Gastroenterology 10: 169–175. July 2017. doi:10.2147/CEG.S133939. PMID 28765714.
- ↑ "IgG antibody response to toxins A and B in patients with Clostridium difficile infection". Clinical and Vaccine Immunology 19 (9): 1552–1554. September 2012. doi:10.1128/CVI.00210-12. PMID 22787196.
- ↑ "Recurrent Clostridioides difficile Infection Is Associated With Impaired T Helper Type 17 Immunity to C difficile Toxin B". Gastroenterology 160 (4): 1410–1413.e4. March 2021. doi:10.1053/J.GASTRO.2020.11.043. PMID 33253683.
- ↑ 50.0 50.1 "Overview of current detection methods and microRNA potential in Clostridioides difficile infection screening". World Journal of Gastroenterology 29 (22): 3385–3399. June 2023. doi:10.3748/wjg.v29.i22.3385. PMID 37389232.
- ↑ "Fecal Microbiota Transplantation for Recurrent Clostridioides difficile Infection Enhances Adaptive Immunity to C difficile Toxin B". Gastroenterology 160 (6): 2155–2158.e4. May 2021. doi:10.1053/J.GASTRO.2021.01.009. PMID 33444574.
- ↑ Cribas, Emily S.; Denny, Joshua E.; Maslanka, Jeffrey R.; Abt, Michael C. (April 16, 2021). "Loss of Interleukin-10 (IL-10) Signaling Promotes IL-22-Dependent Host Defenses against Acute Clostridioides difficile Infection". Infection and Immunity 89 (5): e00730–20. doi:10.1128/IAI.00730-20. ISSN 1098-5522. PMID 33649048.
- ↑ Drewes, Julia L.; Chen, Jie; Markham, Nicholas O.; Knippel, Reece J.; Domingue, Jada C.; Tam, Ada J.; Chan, June L.; Kim, Lana et al. (2022-08-05). "Human Colon Cancer–Derived Clostridioides difficile Strains Drive Colonic Tumorigenesis in Mice" (in en). Cancer Discovery 12 (8): 1873–1885. doi:10.1158/2159-8290.CD-21-1273. ISSN 2159-8274. PMID 35678528.
- ↑ Fettucciari, Katia; Fruganti, Alessandro; Stracci, Fabrizio; Spaterna, Andrea; Marconi, Pierfrancesco; Bassotti, Gabrio (2023-05-02). "Clostridioides difficile Toxin B Induced Senescence: A New Pathologic Player for Colorectal Cancer?" (in en). International Journal of Molecular Sciences 24 (9): 8155. doi:10.3390/ijms24098155. ISSN 1422-0067. PMID 37175861.
- ↑ Azimirad, Masoumeh; Noori, Maryam; Mazhari, Sogol; Azizi Raftar, Shahrbanoo Keshavarz; Ghorbaninejad, Mahsa; Meyfour, Anna; Mortazavi, Pejman; Zali, Mohammad Reza et al. (2025). "Membrane vesicles from selected Clostridioides difficile strains induce epithelial-mesenchymal transition in colonic epithelial cells: insights from in vitro and in vivo studies" (in en). Microbial Pathogenesis 208. doi:10.1016/j.micpath.2025.107988. PMID 40816603. https://linkinghub.elsevier.com/retrieve/pii/S0882401025007132.
- ↑ 56.0 56.1 "Clostridium difficile infection: new developments in epidemiology and pathogenesis". Nature Reviews. Microbiology 7 (7): 526–536. July 2009. doi:10.1038/nrmicro2164. PMID 19528959.
- ↑ "Clinical Practice Guidelines for Clostridium difficile Infection in Adults and Children: 2017 Update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA)". Clinical Infectious Diseases 66 (7): e1–e48. March 2018. doi:10.1093/cid/cix1085. PMID 29462280.
- ↑ "Efficacy of fidaxomicin versus vancomycin as therapy for Clostridium difficile infection in individuals taking concomitant antibiotics for other concurrent infections". Clinical Infectious Diseases 53 (5): 440–447. September 2011. doi:10.1093/cid/cir404. PMID 21844027.
- ↑ "2017 Infectious Diseases Society of America Clinical Practice Guidelines for the Diagnosis and Management of Infectious Diarrhea". Clinical Infectious Diseases 65 (12): e45–e80. November 2017. doi:10.1093/cid/cix669. PMID 29053792.
- ↑ "Recurrent Clostridium difficile Infection: Risk Factors, Treatment, and Prevention". Gut and Liver 13 (1): 16–24. January 2019. doi:10.5009/gnl18071. PMID 30400734.
- ↑ "Treatment of Recurrent Clostridium difficile Colitis with Vancomycin and Saccharomyces boulardii". The American Journal of Gastroenterology. http://www.optibacprobiotics.sg/uploads/surawicz_et_al_(1989)_treatment_of_recurrent_clostridium_difficile_colitis_with_vancomycin_and_saccharomyces_boulardii_optibac_probiotics_www.optibacprobiotics.co.uk.pdf. Retrieved April 28, 2018.
- ↑ 62.0 62.1 "Guidelines for diagnosis, treatment, and prevention of Clostridium difficile infections". The American Journal of Gastroenterology 108 (4): 478–98; quiz 499. April 2013. doi:10.1038/ajg.2013.4. PMID 23439232.
- ↑ "European Society of Clinical Microbiology and Infectious Diseases: 2021 update on the treatment guidance document for Clostridioides difficile infection in adults". Clinical Microbiology and Infection 27 (Suppl 2): S1–S21. December 2021. doi:10.1016/j.cmi.2021.09.038. PMID 34678515. https://findresearcher.sdu.dk/ws/files/197926149/Open_Access_Version.pdf.
- ↑ "Is tigecycline a suitable option for Clostridium difficile infection? Evidence from the literature". International Journal of Antimicrobial Agents 46 (1): 8–12. July 2015. doi:10.1016/j.ijantimicag.2015.03.012. PMID 25982915.
- ↑ "Bezlotoxumab for Preventing Recurrent Clostridioides difficile Infection: A Narrative Review from Pathophysiology to Clinical Studies". Infectious Diseases and Therapy 9 (3): 481–494. September 2020. doi:10.1007/s40121-020-00314-5. PMID 32632582.
- ↑ AKAGAWA, Shohei; AKAGAWA, Yuko; YAMANOUCHI, Sohsaku; KIMATA, Takahisa; TSUJI, Shoji; KANEKO, Kazunari (August 25, 2020). "Development of the gut microbiota and dysbiosis in children". Bioscience of Microbiota, Food and Health 40 (1): 12–18. doi:10.12938/bmfh.2020-034. PMID 33520564.
- ↑ "Effect of Faecal Microbiota Transplantation for Treatment of Clostridium difficile Infection in Patients With Inflammatory Bowel Disease: A Systematic Review and Meta-Analysis of Cohort Studies". Journal of Crohn's & Colitis 12 (6): 710–717. May 2018. doi:10.1093/ecco-jcc/jjy031. PMID 29528385.
- ↑ Littmann, Eric R.; Lee, Jung-Jin; Denny, Joshua E.; Alam, Zahidul; Maslanka, Jeffrey R.; Zarin, Isma; Matsuda, Rina; Carter, Rebecca A. et al. (February 2, 2021). "Host immunity modulates the efficacy of microbiota transplantation for treatment of Clostridioides difficile infection". Nature Communications 12 (1): 755. doi:10.1038/s41467-020-20793-x. ISSN 2041-1723. PMID 33531483. Bibcode: 2021NatCo..12..755L.
- ↑ Stewart, David B.; Berg, Arthur; Hegarty, John (January 2013). "Predicting Recurrence of C. difficile Colitis Using Bacterial Virulence Factors: Binary Toxin Is the Key" (in en). Journal of Gastrointestinal Surgery 17 (1): 118–125. doi:10.1007/s11605-012-2056-6. PMID 23086451. https://linkinghub.elsevier.com/retrieve/pii/S1091255X23071019. Retrieved March 20, 2024.
- ↑ "Clostridium difficile Infection: A Worldwide Disease". Gut Liver 8 (1): 1–6. January 29, 2014. doi:10.5009/gnl.2014.8.1.1. PMID 24516694.
- ↑ "Guidance document for prevention of Clostridium difficile infection in acute healthcare settings". Clinical Microbiology and Infection 24 (10): 1051–1054. October 2018. doi:10.1016/j.cmi.2018.02.020. PMID 29505879.
- ↑ "Characterization of Clostridium difficile Strains in British Columbia, Canada: A Shift from NAP1 Majority (2008) to Novel Strain Types (2013) in One Region". The Canadian Journal of Infectious Diseases & Medical Microbiology 2016. March 29, 2016. doi:10.1155/2016/8207418. PMID 27366181.
- ↑ "Dietary trehalose enhances virulence of epidemic Clostridium difficile". Nature 553 (7688): 291–294. January 2018. doi:10.1038/nature25178. PMID 29310122. Bibcode: 2018Natur.553..291C.
- ↑ Sebaihia, Mohammed; Wren, Brendan W; Mullany, Peter et al. (July 2006). "The multidrug-resistant human pathogen Clostridium difficile has a highly mobile, mosaic genome". Nature Genetics 38 (7): 779–786. doi:10.1038/ng1830. PMID 16804543. Bibcode: 2006NaGen..38..779S.
- ↑ "Evolutionary dynamics of Clostridium difficile over short and long time scales". Proceedings of the National Academy of Sciences of the United States of America 107 (16): 7527–7532. April 2010. doi:10.1073/pnas.0914322107. PMID 20368420. Bibcode: 2010PNAS..107.7527H.
- ↑ Scientists map C. difficile strain – Institute of Public Affairs, Montreal
- ↑ "Microevolutionary analysis of Clostridium difficile genomes to investigate transmission". Genome Biology 13 (12). December 2012. doi:10.1186/gb-2012-13-12-r118. PMID 23259504.
- ↑ 78.0 78.1 "Clostridium difficile phages: still difficult?". Frontiers in Microbiology 5: 184. 2014. doi:10.3389/fmicb.2014.00184. PMID 24808893.
External links
- Canada Pathogen Safety Data Sheets: Infectious Substances – Clostridium difficile, Public Health Agency, Canada, September 10, 2014.
- Type strain of Clostridium difficile, BacDive—the Bacterial Diversity Metadatabase
Wikidata ☰ {{{from}}} entry
 |
