Biology:NAD(P)(+)—protein-arginine ADP-ribosyltransferase
| NAD(P)+-protein-arginine ADP-ribosyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 2.4.2.31 | ||||||||
| CAS number | 81457-93-4 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| ART | |||||||||
|---|---|---|---|---|---|---|---|---|---|
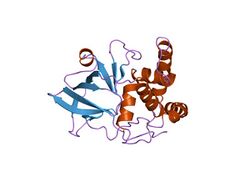 crystal structure of the eucaryotic mono-adp-ribosyltransferase art2.2; crystal form c (p3121) | |||||||||
| Identifiers | |||||||||
| Symbol | ART | ||||||||
| Pfam | PF01129 | ||||||||
| Pfam clan | CL0084 | ||||||||
| InterPro | IPR000768 | ||||||||
| SMART | START | ||||||||
| PROSITE | PDOC00993 | ||||||||
| SCOP2 | 1gy0 / SCOPe / SUPFAM | ||||||||
| |||||||||
In enzymology, a NAD(P)+-protein-arginine ADP-ribosyltransferase (EC 2.4.2.31) is an enzyme that catalyzes the chemical reaction using nicotinamide adenine dinucleotide
- NAD+ + protein L-arginine [math]\displaystyle{ \rightleftharpoons }[/math] nicotinamide + Nomega-(ADP-D-ribosyl)-protein-L-arginine NADP+ + protein L-arginine [math]\displaystyle{ \rightleftharpoons }[/math] nicotinamide + Nomega-[(2'-phospho-ADP)-D-ribosyl]-protein-L-arginine
as well as the corresponding reaction using nicotinamide adenine dinucleotide phosphate
- NADP+ + protein L-arginine [math]\displaystyle{ \rightleftharpoons }[/math] nicotinamide + Nomega-(ADP-D-ribosyl)-protein-L-arginine NADP+ + protein L-arginine [math]\displaystyle{ \rightleftharpoons }[/math] nicotinamide + Nomega-[(2'-phospho-ADP)-D-ribosyl]-protein-L-arginine
Thus, the two substrates of this enzyme are NAD+ (or NADP+) and protein L-arginine, whereas its two products are nicotinamide and Nomega-(ADP-D-ribosyl)-protein-L-arginine (or Nomega-[(2'-phospho-ADP)-D-ribosyl]-protein-L-arginine, respectively).
This enzyme belongs to the family of glycosyltransferases, specifically the pentosyltransferases. The systematic name of this enzyme class is NAD(P)+:protein-L-arginine ADP-D-ribosyltransferase. Other names in common use include ADP-ribosyltransferase, mono(ADP-ribosyl)transferase, NAD+:L-arginine ADP-D-ribosyltransferase, NAD(P)+-arginine ADP-ribosyltransferase, and NAD(P)+:L-arginine ADP-D-ribosyltransferase.
At least five forms of the enzyme have been characterised to date, some of which are attached to the membrane via glycosylphosphatidylinositol (GPI) anchors, while others appear to be secreted. The enzymes contain ~250-300 residues, which encode putative signal sequences and carbohydrate attachment sites. In addition, the N- and C-termini are predominantly hydrophobic, a characteristic of GPI-anchored proteins.[1]
Structural studies
As of late 2007, 6 structures have been solved for this class of enzymes, with PDB accession codes 1GXY, 1GXZ, 1GY0, 1OG1, 1OG3, and 1OG4.
References
- ↑ "Immunological and structural conservation of mammalian skeletal muscle glycosylphosphatidylinositol-linked ADP-ribosyltransferases". Biochemistry 33 (43): 12828–36. November 1994. doi:10.1021/bi00209a014. PMID 7947688.
Further reading
- "Substrate specificity and partial purification of a stereospecific NAD- and guanidine-dependent ADP-ribosyltransferase from avian erythrocytes". J. Biol. Chem. 254 (18): 8891–4. 1979. PMID 225315.
- "Isolation and properties of an NAD- and guanidine-dependent ADP-ribosyltransferase from turkey erythrocytes". J. Biol. Chem. 255 (12): 5838–40. 1980. PMID 6247348.
- "ADP-ribosylation". Annu. Rev. Biochem. 54 (1): 73–100. 1985. doi:10.1146/annurev.bi.54.070185.000445. PMID 3927821.
 |

