Biology:Leptospira
| Leptospira | |
|---|---|
| |
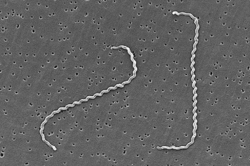
| Scanning electron micrograph of Leptospira interrogans | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Spirochaetota |
| Class: | Spirochaetia |
| Order: | Leptospirales |
| Family: | Leptospiraceae |
| Genus: | Leptospira Noguchi 1917 non Swainson 1840 non Boucot, Johnson & Staton 1964 |
| Type species | |
| Leptospira interrogans (Stimson 1907) Wenyon 1926
| |
| Species | |
|
See text | |
Leptospira (Ancient Greek:, 'fine, thin' and Latin: spira, 'coil')[1] is a genus of spirochaete bacteria, including a small number of pathogenic and saprophytic species.[2] Leptospira was first observed in 1907 in kidney tissue slices of a leptospirosis victim who was described as having died of "yellow fever".[3]
Taxonomy
Leptospira, together with the genera Leptonema and Turneria, is a member of the family Leptospiraceae. The genus Leptospira is divided into 20 species based on DNA hybridization studies.[4][5]
Pathogenic Leptospira
- Leptospira alstonii Smythe et al. 2013 ["Leptospira alstoni" Haake et al. 1993]
- Leptospira interrogans (Stimson 1907) Wenyon 1926 emend. Faine and Stallman 1982 ["Spirochaeta interrogans" Stimson 1907; "Spirochaeta nodosa" Hubener & Reiter 1916; "Spirochaeta icterohaemorrhagiae" Inada et al. 1916; "Spirochaeta icterogenes" Uhlenhuth & Fromme 1916; "Leptospira icteroides" Noguchi 1919]
- Leptospira kirschneri Ramadass et al. 1992
- Leptospira noguchii Yasuda et al. 1987
- Leptospira alexanderi Brenner et al. 1999
- Leptospira weilii Yasuda et al. 1987
- Leptospira borgpetersenii Yasuda et al. 1987
- Leptospira santarosai Yasuda et al. 1987
- Leptospira kmetyi Slack et al. 2009[6]
- Leptospira mayottensis Bourhy et al. 2014
Intermediates or opportunistic Leptospira
- Leptospira inadai Yasuda et al. 1987
- Leptospira fainei Perolat et al. 1998
- Leptospira broomii Levett et al. 2006[7]
- Leptospira licerasiae Matthias et al. 2009[8]
- Leptospira wolffii Slack et al. 2008[9]
Non-pathogenic Leptospira
- Leptospira biflexa (Wolbach and Binger 1914) Noguchi 1918 emend. Faine and Stallman 1982 ["Spirochaeta biflexa" Wolbach & Binger 1914]
- Leptospira idonii Saito et al. 2013
- Leptospira meyeri Yasuda et al. 1987
- Leptospira wolbachii Yasuda et al. 1987
- Leptospira vanthielii Smythe et al. 2013
- Leptospira terpstrae Smythe et al. 2013
- Leptospira yanagawae Smythe et al. 2013
Members of Leptospira are also grouped into serovars according to their antigenic relatedness. There are currently over 200 recognized serovars. A few serovars are found in more than one species of Leptospira.
At its 2002 meeting, the Committee on the Taxonomy of Leptospira of the International Union of Microbiological Societies approved the following nomenclature for serovars of Leptospira. Genus and species names are italicized as usual, with the serovar name not italicized and with an upper case first letter.
- Genus species serovar Serovar_name
For example:
- Leptospira interrogans serovar Australis
- Leptospira biflexa serovar Patoc
Phylogeny
The currently accepted taxonomy is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN)[10] and National Center for Biotechnology Information (NCBI).[11]
| 16S rRNA based LTP_08_2023[12][13][14] | 120 marker proteins based GTDB 08-RS214[15][16][17] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
Species incertae sedis:
- "L. andamana" Tripathy, Evans & Hanson 1971 non Collier 1948
- "L. brihuegai" Grune Loffler et al. 2015
- "L. hardjobovis" Srinivas, Walker & Rippke 2013
- "L. macculloughii" Thibeaux et al. 2018
- L. sanjuanensis Fernandes et al. 2022
Morphology
Although over 200 serotypes oLeptospira have been described, all members of the genus have similar morphology. Leptospira are spiral-shaped bacteria that are 6-20 μm long and 0.1 μm in diameter with a wavelength of about 0.5 μm.[18] One or both ends of the spirochete are usually hooked. Because they are so thin, live Leptospira are best observed by darkfield microscopy.
The bacteria have a number of degrees of freedom; when ready to proliferate via binary fission, the bacterium noticeably bends in the place of the future split.
Cellular structure
Leptospira have a Gram-negative-like cell envelope consisting of a cytoplasmic and outer membrane. However, the peptidoglycan layer is associated with the cytoplasmic rather than the outer membrane, an arrangement that is unique to spirochetes. The two flagella of Leptospira extend from the cytoplasmic membrane at the ends of the bacterium into the periplasmic space and are necessary for the motility of Leptospira.[19]
The outer membrane contains a variety of lipoproteins and transmembrane outer membrane proteins.[20] As expected, the protein composition of the outer membrane differs when comparing Leptospira growing in artificial medium with Leptospira present in an infected animal.[21][22][23] Several leptospiral outer membrane proteins have been shown to attach to the host extracellular matrix and to factor H. These proteins may be important for adhesion of Leptospira to host tissues and in resisting complement, respectively.[24][25][26]
The outer membrane of Leptospira, like those of most other Gram-negative bacteria, contains lipopolysaccharide (LPS). Differences in the highly immunogenic LPS structure account for the numerous serovars of Leptospira.[18] Consequently, immunity is serovar specific; current leptospiral vaccines, which consist of one or several serovars of Leptospira endemic in the population to be immunized, protect only against the serovars contained in the vaccine preparation. Leptospiral LPS has low endotoxin activity.[18] An unusual feature of leptospiral LPS is that it activates host cells via TLR2 rather than TLR4.[27] The unique structure of the lipid A portion of the LPS molecule may account for this observation.[28] Finally, the LPS O antigen content of L. interrogans differs in an acutely infected versus a chronically infected animal.[29] The role of O antigen changes in the establishment or maintenance of acute or chronic infection, if any, is unknown.
Habitat
Leptospira, both pathogenic and saprophytic, can occupy diverse environments, habitats, and life cycles; these bacteria are found throughout the world, except in Antarctica. High humidity and neutral (6.9–7.4) pH are necessary for their survival in the environment, with stagnant water reservoirs—bogs, shallow lakes, ponds, puddles, etc.—being the natural habitat for the bacteria.
Nutrition
Leptospira are cultivated at 30 °C in Ellinghausen-McCullough-Johnson-Harris (EMJH) medium, which can be supplemented with 0.21% rabbit serum to enhance growth of fastidious strains.[30] Growth of pathogenic Leptospira in an artificial nutrient environment such as EMJH becomes noticeable in 4–7 days; growth of saprophytic strains occur within 2–3 days. The minimal growth temperature of pathogenic species is 13–15 °C. Because the minimal growth temperature of the saprophytes is 5–10 °C, the ability of Leptospira to grow at 13 °C can be used to distinguish saprophytic from pathogenic Leptospira species.[30] The optimal pH for growth of Leptospira is 7.2–7.6.
Leptospira are aerobes whose major carbon and energy source during in vitro growth is long-chain fatty acids, which are metabolized by beta-oxidation.[31][32] Fatty acids are provided in EMJH in the form of Tween.[30] Fatty acid molecules are bound by albumin in EMJH and are released slowly into the medium to prevent its toxic accumulation.
Like most bacteria, Leptospira require iron for growth.[33] L. interrogans and L. biflexa have the ability to acquire iron in different forms.[34] A TonB-dependent receptor required for utilization of the ferrous form of the iron has been identified in L. biflexa, and an ortholog of the receptor is encoded in the genome of L. interrogans. L. interrogans can also obtain iron from heme, which is bound to most of the iron in the human body. The HbpA hemin-binding protein, which may be involved in the uptake of hemin, has been identified on the surface of L. interrogans[35] Although other pathogenic species of Leptospira and L. biflexa lack HbpA, yet another hemin-binding protein, LipL41, may account for their ability to use hemin as a source of iron.[35] Although they do not secrete siderophores, L. biflexa and L. interrogans may be capable of obtaining iron from siderophores secreted by other microorganisms.[34]
Genome
The genome of pathogenic Leptospira consists of two chromosomes. The size of the genomes of L. interrogans serovars Copenhageni and Lai is approximately 4.6 Mb.[36][37] However, the genome of L. borgpetersenii serovar Hardjo is only 3.9 Mb in size with a large number of pseudogenes, gene fragments, and insertion sequences relative to the genomes of L. interrogans.[38] L. interrogans and L. borgpetersenii share 2708 genes from which 656 are pathogenic specific genes. The guanine plus cytosine (GC) content is between 35% and 41%.[39] L. borgpetersenii serovar Hardjo is usually transmitted by direct exposure to infected tissues, whereas L. interrogans is often acquired from water or soil contaminated by the urine of carrier animals harboring Leptospira in their kidneys. The high number of defective genes and insertion sequences in L. borgpetersenii Hardjo together with the poor survival outside of the host and difference in transmission patterns compared to L. interrogans suggest that L. borgpetersenii is undergoing insertion-sequence mediated genomic decay, with ongoing loss of genes necessary for survival outside of the host animal.[38]
Genotyping
Genome sequence determination several strains of Leptospira lead to the development of multilocus VNTR (Variable Number of Tandem Repeats) typing and multilocus sequence typing (MLST) for species level identification of pathogenic Leptospira species.[40] Both methods hold the potential to replace the highly ambiguous serotyping method currently in vogue for leptospiral strain identification.[40]
See also
References
- ↑ "leptospirosis". American Heritage Dictionary of the English Language: Fourth Edition. Bartleby.com. 2000. http://www.bartleby.com/61/66/L0126650.html.
- ↑ Ryan KJ, ed (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. ISBN 978-0-8385-8529-0.
- ↑ Stimson AM (1907). "Note on an organism found in yellow-fever tissue". Public Health Reports 22 (18): 541. doi:10.2307/4559008.
- ↑ "Further determination of DNA relatedness between serogroups and serovars in the family Leptospiraceae with a proposal for Leptospira alexanderi sp. nov. and four new Leptospira genomospecies". Int. J. Syst. Bacteriol. 49 (2): 839–58. 1999. doi:10.1099/00207713-49-2-839. PMID 10319510.
- ↑ "Leptospirosis: a zoonotic disease of global importance". The Lancet Infectious Diseases 3 (12): 757–71. 2003. doi:10.1016/S1473-3099(03)00830-2. PMID 14652202.
- ↑ "Leptospira kmetyi sp. nov., isolated from an environmental source in Malaysia". Int. J. Syst. Evol. Microbiol. 59 (Pt 4): 705–8. April 2009. doi:10.1099/ijs.0.002766-0. PMID 19329592.
- ↑ "Leptospira broomii sp. nov., isolated from humans with leptospirosis". Int. J. Syst. Evol. Microbiol. 56 (Pt 3): 671–3. 2006. doi:10.1099/ijs.0.63783-0. PMID 16514048.
- ↑ Picardeau, Mathieu, ed (2008). "Human Leptospirosis Caused by a New, Antigenically Unique Leptospira Associated with a Rattus Species Reservoir in the Peruvian Amazon". PLOS Negl Trop Dis 2 (4): e213. doi:10.1371/journal.pntd.0000213. PMID 18382606.
- ↑ "Leptospira wolffii sp. nov., isolated from a human with suspected leptospirosis in Thailand". International Journal of Systematic and Evolutionary Microbiology 58 (Pt 10): 2305–8. October 2008. doi:10.1099/ijs.0.64947-0. PMID 18842846.
- ↑ J.P. Euzéby. "Leptospira". List of Prokaryotic names with Standing in Nomenclature (LPSN). https://lpsn.dsmz.de/genus/leptospira.
- ↑ Sayers. "Leptospira". National Center for Biotechnology Information (NCBI) taxonomy database. https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Tree&id=171&lvl=3&lin=f&keep=1&srchmode=1&unlock.
- ↑ "The LTP". https://imedea.uib-csic.es/mmg/ltp/#LTP.
- ↑ "LTP_all tree in newick format". https://imedea.uib-csic.es/mmg/ltp/wp-content/uploads/ltp/LTP_all_08_2023.ntree.
- ↑ "LTP_08_2023 Release Notes". https://imedea.uib-csic.es/mmg/ltp/wp-content/uploads/ltp/LTP_08_2023_release_notes.pdf.
- ↑ "GTDB release 08-RS214". https://gtdb.ecogenomic.org/about#4%7C.
- ↑ "bac120_r214.sp_label". https://data.gtdb.ecogenomic.org/releases/release214/214.0/auxillary_files/bac120_r214.sp_labels.tree.
- ↑ "Taxon History". https://gtdb.ecogenomic.org/taxon_history/.
- ↑ 18.0 18.1 18.2 Levett PN (2001). "Leptospirosis". Clin. Microbiol. Rev. 14 (2): 296–326. doi:10.1128/CMR.14.2.296-326.2001. PMID 11292640.
- ↑ "First evidence for gene replacement in Leptospira spp. Inactivation of L. biflexa flaB results in non-motile mutants deficient in endoflagella". Mol. Microbiol. 40 (1): 189–99. 2001. doi:10.1046/j.1365-2958.2001.02374.x. PMID 11298286.
- ↑ "Global Analysis of Outer Membrane Proteins from Leptospira interrogans Serovar Lai". Infect. Immun. 70 (5): 2311–8. 2002. doi:10.1128/IAI.70.5.2311-2318.2002. PMID 11953365.
- ↑ "Characterization of Leptospiral Outer Membrane Lipoprotein LipL36: Downregulation Associated with Late-Log-Phase Growth and Mammalian Infection". Infect. Immun. 66 (4): 1579–87. 1998. doi:10.1128/IAI.66.4.1579-1587.1998. PMID 9529084.
- ↑ "Cloning and Molecular Characterization of an Immunogenic LigA Protein of Leptospira interrogans". Infect. Immun. 70 (11): 5924–30. 2002. doi:10.1128/IAI.70.11.5924-5930.2002. PMID 12379666.
- ↑ "Characterization of the Outer Membrane Proteome of Leptospira interrogans Expressed during Acute Lethal Infection". Infect. Immun. 75 (2): 766–73. 2007. doi:10.1128/IAI.00741-06. PMID 17101664.
- ↑ "LfhA, a Novel Factor H-Binding Protein of Leptospira interrogans". Infect. Immun. 74 (5): 2659–66. 2006. doi:10.1128/IAI.74.5.2659-2666.2006. PMID 16622202.
- ↑ "A Newly Identified Leptospiral Adhesin Mediates Attachment to Laminin". Infect. Immun. 74 (11): 6356–64. 2006. doi:10.1128/IAI.00460-06. PMID 16954400.
- ↑ "Physiological Osmotic Induction of Leptospira interrogans Adhesion: LigA and LigB Bind Extracellular Matrix Proteins and Fibrinogen". Infect. Immun. 75 (5): 2441–50. 2007. doi:10.1128/IAI.01635-06. PMID 17296754.
- ↑ "Leptospiral lipopolysaccharide activates cells through a TLR2-dependent mechanism". Nat. Immunol. 2 (4): 346–52. 2001. doi:10.1038/86354. PMID 11276206.
- ↑ "A Methylated Phosphate Group and Four Amide-linked Acyl Chains in Leptospira interrogans Lipid A. The Membrane Anchor of an Unusual Lipopolysaccharide that Activates TLR2". J. Biol. Chem. 279 (24): 25420–9. 2004. doi:10.1074/jbc.M400598200. PMID 15044492.
- ↑ "Changes in Lipopolysaccharide O Antigen Distinguish Acute versus Chronic Leptospira interrogans Infections". Infect. Immun. 73 (6): 3251–60. 2005. doi:10.1128/IAI.73.6.3251-3260.2005. PMID 15908349.
- ↑ 30.0 30.1 30.2 "Differentiation of Pathogenic and Saprophytic Leptospires I. Growth at Low Temperatures". J. Bacteriol. 94 (1): 27–31. 1967. doi:10.1128/jb.94.1.27-31.1967. PMID 6027998.
- ↑ "NUTRITION OF LEPTOSPIRA POMONA II. : Fatty Acid Requirements". J. Bacteriol. 85 (5): 976–82. 1963. doi:10.1128/jb.85.5.976-982.1963. PMID 14044026.
- ↑ "Beta-oxidation of fatty acids by Leptospira". Can. J. Microbiol. 16 (1): 41–5. 1970. doi:10.1139/m70-007. PMID 5415967.
- ↑ Faine S (1959). "Iron as a growth requirement for pathogenic Leptospira". J. Gen. Microbiol. 20 (2): 246–51. doi:10.1099/00221287-20-2-246. PMID 13654718.
- ↑ 34.0 34.1 "Comparative and Functional Genomic Analyses of Iron Transport and Regulation in Leptospira spp". J. Bacteriol. 188 (22): 7893–904. 2006. doi:10.1128/JB.00711-06. PMID 16980464.
- ↑ 35.0 35.1 "Expression and Characterization of an Iron-Regulated Hemin-Binding Protein, HbpA, from Leptospira interrogans Serovar Lai". Infect. Immun. 75 (9): 4582–91. 2007. doi:10.1128/IAI.00324-07. PMID 17576761.
- ↑ "Unique physiological and pathogenic features of Leptospira interrogans revealed by whole-genome sequencing". Nature 422 (6934): 888–93. 2003. doi:10.1038/nature01597. PMID 12712204. Bibcode: 2003Natur.422..888R.
- ↑ "Comparative Genomics of Two Leptospira interrogans Serovars Reveals Novel Insights into Physiology and Pathogenesis". J. Bacteriol. 186 (7): 2164–72. 2004. doi:10.1128/JB.186.7.2164-2172.2004. PMID 15028702.
- ↑ 38.0 38.1 "Genome reduction in Leptospira borgpetersenii reflects limited transmission potential". Proc. Natl. Acad. Sci. U.S.A. 103 (39): 14560–5. 2006. doi:10.1073/pnas.0603979103. PMID 16973745. Bibcode: 2006PNAS..10314560B.
- ↑ "Leptospira: the dawn of the molecular genetics era for an emerging zoonotic pathogen". Nat. Rev. Microbiol. 7 (10): 736–47. October 2009. doi:10.1038/nrmicro2208. PMID 19756012.
- ↑ 40.0 40.1 "A century of Leptospira strain typing". Infect. Genet. Evol. 9 (5): 760–8. September 2009. doi:10.1016/j.meegid.2009.06.009. PMID 19540362.
External links
- Leptospira page at Kenyon College MicrobeWiki.
- "Leptospira". NCBI Taxonomy Browser. https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&id=171.
Wikidata ☰ Q133841 entry
 |

