Biology:Enteropeptidase
| enteropeptidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
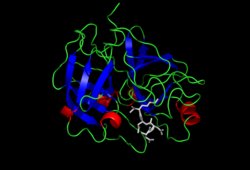 Crystal structure of Enteropeptidase with an
inhibitor | |||||||||
| Identifiers | |||||||||
| EC number | 3.4.21.9 | ||||||||
| CAS number | 9014-74-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| protease, serine, 7 (enteropeptidase) | |
|---|---|
| Identifiers | |
| Symbol | TMPRSS15 |
| NCBI gene | 5651 |
| HGNC | 9490 |
| OMIM | 606635 |
| RefSeq | NM_002772 |
| UniProt | P98073 |
| Other data | |
| Locus | Chr. 21 q21 |
Enteropeptidase (also called enterokinase) is an enzyme produced by cells of the duodenum and is involved in digestion in humans and other animals. Enteropeptidase converts trypsinogen (a zymogen) into its active form trypsin, resulting in the subsequent activation of pancreatic digestive enzymes.[1][2] Absence of enteropeptidase results in intestinal digestion impairment.[3]
Enteropeptidase is a serine protease (EC 3.4.21.9) consisting of a disulfide-linked heavy-chain of 82-140 kDa that anchors enterokinase in the intestinal brush border membrane and a light-chain of 35–62 kDa that contains the catalytic subunit.[4] Enteropeptidase is a part of the chymotrypsin-clan of serine proteases, and is structurally similar to these proteins.[5]
Historical significance
Enteropeptidase was discovered by Ivan Pavlov, who was awarded the 1904 Nobel Prize in Physiology or Medicine for his studies of gastrointestinal physiology. It is the first known enzyme to activate other enzymes, and it remains a remarkable example of how serine proteases have been crafted to regulate metabolic pathways.[6] The inert function of digestive enzymes within the pancreas was known, as compared to their potent activity within the intestine, but the basis of this difference was unknown. In 1899, Pavlov's student, N. P. Schepowalnikov, demonstrated that canine duodenal secretions dramatically stimulated the digestive activity of pancreatic enzymes, especially trypsinogen. The active principle was recognized as a special enzyme in the intestine that could activate other enzymes. Pavlov named it enterokinase. The debate of whether enterokinase was a cofactor or enzyme was resolved by Kunitz, who showed that the activation of trypsinogen by enterokinase was catalytic. In the 1950s, cattle trypsinogen was shown to be activated autocatalytically by cleavage of an N-terminal hexapeptide.[7] The more precise IUBMB name enteropeptidase has been in existence since 1970. However, the original name ‘enterokinase’ has a long history and remains in common use.[8]
Enzyme structure
Enteropeptidase is a type II transmembrane serine protease (TTSP) localized to the brush border of the duodenal and jejunal mucosa and synthesized as a zymogen, proenteropeptidase, which requires activation by duodenase or trypsin.[9] TTSPs are synthesized as single chain zymogens with N-terminal propeptide sequences of different lengths. These enzymes are activated by cleavage at the carboxyl side of lysine or arginine residues present in a highly conserved activation motif. Once activated, TTSPs are predicted to remain membrane-bound through a conserved disulfide bond linking the pro- and catalytic domains.[10]
In the case of cattle enteropeptidase the primary translation product comprises 1035 residues with an expected mass of 114.9 kDa.[11] The detected apparent mass of about 160 kDa is consistent with the specified carbohydrate content of 30 - 40%, with equal amounts of neutral and amino sugars.[12][13] The activation cleavage site after Lys800 splits the heavy and light chains of mature cattle enteropeptidase. There are 17 potential N-linked glycosylation sites in the heavy chain and three in the light chain; most of these are conserved in other species. The heavy chain has a hydrophobic section near the N-terminus that supports the transmembrane anchor.[14][15] The heavy chain influences the specificity of enteropeptidase. Native enteropeptidase is resistant to soybean trypsin inhibitor. However, the isolated light chain is subtle whether prepared by limited reduction of the natural protein[16] or by mutagenesis and expression in COS cells.[17] Native enteropeptidase and the isolated light chain have similar activity toward Gly-(Asp)4-Lys-NHNap, but the secluded light chain has distinctly decreased activity toward trypsinogen . An analogous selective defect in the recognition of trypsinogen can be produced in two-chain enteropeptidase by heating or by acetylation.[18] This behavior implies that the catalytic center and one or more secondary substrate-binding sites are essential for optimal recognition of trypsinogen.
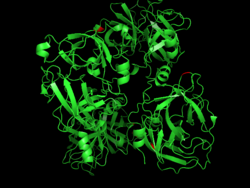
Activity
Despite its alternative name (enterokinase), enteropeptidase is a serine protease that catalyses the hydrolysis of peptide bonds in proteins and, unlike other kinases, does not catalyze transfer of phosphate groups. Enteropeptidase exhibits trypsin-like activity, cleaving proteins following a lysine at a specific cleavage site (Asp-Asp-Asp-Asp-Lys).[19] This cleavage results in trypsindependent activation of other pancreatic zymogens, such as chymotrypsinogen, proelastase, procarboxypeptidase and prolipase in the lumen of the gut.[20] As the pro-region of trypsinogen contains this sequence, enteropeptidase catalyses its activation in vivo:
trypsinogen → trypsin + pro-region (Val-Asp-Asp-Asp-Asp-Lys)
Genetics and disease relevance
In humans, enteropeptidase is encoded by the TMPRSS15 gene (also known as ENTK, and previously as PRSS7) on chromosome 21q21. Some nonsense and frameshift mutations in this gene lead to a rare recessive disorder characterised by severe failure to thrive in affected infants, due to enteropeptidase deficiency.[21] Enteropeptidase mRNA expression is limited to the proximal small intestine, and the protein is found in enterocytes of duodenum and proximal jejunum. Upon secretion from the pancreas into the duodenum, trypsinogen encounters enteropeptidase and is activated. Trypsin then cleaves and activates other pancreatic serine protease zymogens (chymotrypsinogen and proelastases), metalloprotease zymogens (procarboxypeptidases) and prolipases. By means of this simple two-step cascade, the destructive activity of these digestive hydrolases is confined to the lumen of the intestine. The physiological importance of this pathway is demonstrated by the severe intestinal malabsorption caused by congenital deficiency of enteropeptidase.[22][23] This condition can be life-threatening, but responds to oral supplementation with pancreatic extract.
Applications
Enteropeptidase's specificity makes it an ideal tool in biochemical applications; a fusion protein containing a C-terminal affinity tag (such as poly-His) linked by this sequence can be cleaved by enteropeptidase to obtain the target protein following protein purification.[19] On the converse, the N-terminal pro-sequence of proteases that must be cleaved prior to activation can be mutated to enable activation with enteropeptidase.[24]
References
- ↑ Kunitz M (March 1939). "Formation of trypsin from crystalline trypsinogen by means of enterokinase". J. Gen. Physiol. 22 (4): 429–46. doi:10.1085/jgp.22.4.429. PMID 19873112.
- ↑ Kiel B (1971). "Trypsin". in Boyer PS. The Enzymes, 3: Hydrolysis - Peptide Bonds. Amsterdam: Elsevier. pp. 249–75. ISBN 978-0-12-122703-6.
- ↑ "Enterokinase (enteropeptidase) : comparative aspects". Trends Biochem. Sci. 14 (3): 110–2. March 14, 1989. doi:10.1016/0968-0004(89)90133-3. PMID 2658218.
- ↑ "Functional expression and purification of bovine enterokinase light chain in recombinant Escherichia coli". Prep. Biochem. Biotechnol. 37 (3): 205–17. 2007. doi:10.1080/10826060701386695. PMID 17516250.
- ↑ "Evolutionary families of peptidases". Biochem. J. 290 (1): 205–18. February 1993. doi:10.1042/bj2900205. PMID 8439290.
- ↑ "Crystal structure of enteropeptidase light chain complex with an analog of the trypsinogen activation peptide". J Mol Biol 292 (2): 361–73. Sep 17, 1999. doi:10.1006/jmbi.1999.3089. PMID 10493881.
- ↑ Yamashina I. (May 1956). "The action of enterokinase on trypsinogen". Biochim Biophys Acta 20 (2): 433–4. doi:10.1016/0006-3002(56)90329-8. PMID 13328891. http://actachemscand.dk/pdf/acta_vol_10_p0739-0743.pdf.
- ↑ Rawlings, Neil D.; Salvesen, Guy (2013). Handbook of Proteolytic Enzymes (3rd ed.). ISBN 978-0-12-382219-2. http://www.sciencedirect.com/science/book/9780123822192. Retrieved February 20, 2014.
- ↑ "Activation of recombinant proenteropeptidase by duodenase". FEBS Lett. 466 (2–3): 295–9. Jan 28, 2000. doi:10.1016/s0014-5793(00)01092-9. PMID 10682847.
- ↑ "Type II transmembrane serine proteases. Insights into an emerging class of cell surface proteolytic enzymes.". J Biol Chem 276 (2): 857–60. Jan 12, 2001. doi:10.1074/jbc.r000020200. PMID 11060317.
- ↑ "Enterokinase, the initiator of intestinal digestion, is a mosaic protease composed of a distinctive assortment of domains". Proc Natl Acad Sci USA 91 (16): 7588–92. August 2, 1994. doi:10.1073/pnas.91.16.7588. PMID 8052624. Bibcode: 1994PNAS...91.7588K.
- ↑ "Bovine enterokinase. Purification, specificity and some molecular properties". Biochemistry 16 (15): 3354–60. July 26, 1977. doi:10.1021/bi00634a011. PMID 889800.
- ↑ "The preparation and properties of bovine enterokinase". J Biol Chem 254 (5): 1677–83. March 10, 1979. doi:10.1016/S0021-9258(17)37826-2. PMID 762166.
- ↑ "Incorporation of bovine enterokinase in reconstituted soybean phospholipid vesicles". J Biol Chem 258 (5): 3069–74. March 10, 1983. doi:10.1016/S0021-9258(18)32831-X. PMID 6338012.
- ↑ "Bovine proenteroptidase is activated by trypsin, and the specificity of enteropeptidase depends on the heavy chain". J Biol Chem 272 (50): 31293–300. December 12, 1997. doi:10.1074/jbc.272.50.31293. PMID 9395456.
- ↑ "The preparation and properties of the catalytic subunit of bovine enterokinase". J Biol Chem 259 (21): 13195–8. November 10, 1984. doi:10.1016/S0021-9258(18)90676-9. PMID 6386810.
- ↑ "Cloning and functional expression of a cDNA encoding the catalytic subunit of bovine enterokinase". J Biol Chem 268 (31): 23311–7. November 5, 1993. doi:10.1016/S0021-9258(19)49464-7. PMID 8226855.
- ↑ "On the catalytic and binding sites of porcine enteropeptidase". Biochim Biophys Acta 452 (2): 488–96. December 8, 1976. doi:10.1016/0005-2744(76)90199-6. PMID 12810.
- ↑ 19.0 19.1 Terpe K (2003). "Overview of tag protein fusions: from molecular and biochemical fundamentals to commercial systems". Appl Microbiol Biotechnol 60 (5): 523–33. doi:10.1007/s00253-002-1158-6. PMID 12536251. http://wolfson.huji.ac.il/expression/local/tag-protein-fusions.pdf.
- ↑ "Isolation from beef pancreas of crystalline trypsinogen, trypsin, a trypsin inhibitor, and an inhibitor-trypsin compound". J Gen Physiol 19 (6): 991–1007. Jul 20, 1936. doi:10.1085/jgp.19.6.991. PMID 19872978.
- ↑ "Mutations in the proenteropeptidase gene are the molecular cause of congenital enteropeptidase deficiency". Am. J. Hum. Genet. 70 (1): 20–5. January 2002. doi:10.1086/338456. PMID 11719902.
- ↑ "Intestinal enterokinase deficiency". Lancet 1 (7599): 812–3. April 19, 1969. doi:10.1016/s0140-6736(69)92071-6. PMID 4180366.
- ↑ "Malabsorption and growth failure due to intestinal enterokinase deficiency". J. Pediatr. 78 (3): 481–90. March 1971. doi:10.1016/s0022-3476(71)80231-7. PMID 4322674.
- ↑ "Production of active recombinant human chymase from a construct containing the enterokinase cleavage site of trypsinogen in place of the native propeptide sequence". Biol Chem Hoppe-Seyler 376 (11): 681–84. Nov 1995. doi:10.1515/bchm3.1995.376.11.681. PMID 8962677.
External links
- Enteropeptidase at the US National Library of Medicine Medical Subject Headings (MeSH)
 |
