Biology:Inorganic pyrophosphatase
| inorganic pyrophosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
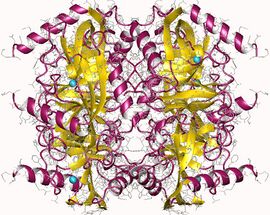 Pyrophosphatase (inorganic) hexamer, E.Coli | |||||||||
| Identifiers | |||||||||
| EC number | 3.6.1.1 | ||||||||
| CAS number | 9024-82-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Soluble inorganic pyrophosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
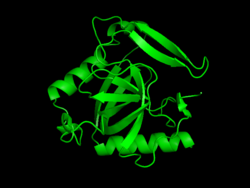 Structure of soluble inorganic pyrophosphatase, isolated from Thermococcus litoralis (PDB: 2PRD). | |||||||||
| Identifiers | |||||||||
| Symbol | Pyrophosphatase | ||||||||
| Pfam | PF00719 | ||||||||
| InterPro | IPR008162 | ||||||||
| PROSITE | PS00387 | ||||||||
| CATH | 2prd | ||||||||
| SCOP2 | 2prd / SCOPe / SUPFAM | ||||||||
| CDD | cd00412 | ||||||||
| |||||||||
| pyrophosphatase (inorganic) 1 | |
|---|---|
| Identifiers | |
| Symbol | PPA1 |
| Alt. symbols | PP |
| NCBI gene | 5464 |
| HGNC | 9226 |
| OMIM | 179030 |
| RefSeq | NM_021129 |
| UniProt | Q15181 |
| Other data | |
| Locus | Chr. 10 q11.1-q24 |
| pyrophosphatase (inorganic) 2 | |
|---|---|
| Identifiers | |
| Symbol | PPA2 |
| NCBI gene | 27068 |
| HGNC | 28883 |
| OMIM | 609988 |
| RefSeq | NM_176869 |
| UniProt | Q9H2U2 |
| Other data | |
| Locus | Chr. 4 q25 |
Inorganic pyrophosphatase (or inorganic diphosphatase, PPase) is an enzyme (EC 3.6.1.1) that catalyzes the conversion of one ion of pyrophosphate to two phosphate ions.[1] This is a highly exergonic reaction, and therefore can be coupled to unfavorable biochemical transformations in order to drive these transformations to completion.[2] The functionality of this enzyme plays a critical role in lipid metabolism (including lipid synthesis and degradation), calcium absorption and bone formation,[3][4] and DNA synthesis,[5] as well as other biochemical transformations.[6][7]
Two types of inorganic diphosphatase, very different in terms of both amino acid sequence and structure, have been characterised to date: soluble and transmembrane proton-pumping pyrophosphatases (sPPases and H(+)-PPases, respectively). sPPases are ubiquitous proteins that hydrolyse pyrophosphate to release heat, whereas H+-PPases, so far unidentified in animal and fungal cells, couple the energy of PPi hydrolysis to proton movement across biological membranes.[8][9]
Structure
Thermostable soluble pyrophosphatase had been isolated from the extremophile Thermococcus litoralis. The 3-dimensional structure was determined using x-ray crystallography, and was found to consist of two alpha-helices, as well as an antiparallel closed beta-sheet. The form of inorganic pyrophosphatase isolated from Thermococcus litoralis was found to contain a total of 174 amino acid residues and have a hexameric oligomeric organization (Image 1).[10]
Humans possess two genes encoding pyrophosphatase, PPA1 and PPA2.[11] PPA1 has been assigned to a gene locus on human chromosome 10,[12] and PPA2 to chromosome 4.[13]
Mechanism
Though the precise mechanism of catalysis via inorganic pyrophosphatase in most organisms remains uncertain, site-directed mutagenesis studies in Escherichia coli have allowed for analysis of the enzyme active site and identification of key amino acids. In particular, this analysis has revealed 17 residues of that may be of functional importance in catalysis.[14]
Further research suggests that the protonation state of Asp67 is responsible for modulating the reversibility of the reaction in Escherichia coli. The carboxylate functional group of this residue has been shown to perform a nucleophilic attack on the pyrophosphate substrate when four magnesium ions are present. Direct coordination with these four magnesium ions and hydrogen bonding interactions with Arg43, Lys29, and Lys142 (all positively charged residues) have been shown to anchor the substrate to the active site. The four magnesium ions are also suggested to be involved in the stabilization of the trigonal bipyramid transition state, which lowers the energetic barrier for the aforementioned nucleophilic attack.[14]
Several studies have also identified additional substrates that can act as allosteric effectors. In particular, the binding of pyrophosphate (PPi) to the effector site of inorganic pyrophosphatase increases its rate of hydrolysis at the active site.[15] ATP has also been shown to function as an allosteric activator in Escherichia coli,[16] while fluoride has been shown to inhibit hydrolysis of pyrophosphate in yeast.[17]
Biological function and significance
The hydrolysis of inorganic pyrophosphate (PPi) to two phosphate ions is utilized in many biochemical pathways to render reactions effectively irreversible.[18] This process is highly exergonic (accounting for approximately a −19kJ change in free energy), and therefore greatly increases the energetic favorability of reaction system when coupled with a typically less-favorable reaction.[19]
Inorganic pyrophosphatase catalyzes this hydrolysis reaction in the early steps of lipid degradation, a prominent example of this phenomenon. By promoting the rapid hydrolysis of pyrophosphate (PPi), Inorganic pyrophosphatase provides the driving force for the activation of fatty acids destined for beta oxidation.[19]
Before fatty acids can undergo degradation to fulfill the metabolic needs of an organism, they must first be activated via a thioester linkage to coenzyme A. This process is catalyzed by the enzyme acyl CoA synthetase, and occurs on the outer mitochondrial membrane. This activation is accomplished in two reactive steps: (1) the fatty acid reacts with a molecule of ATP to form an enzyme-bound acyl adenylate and pyrophosphate (PPi), and (2) the sulfhydryl group of CoA attacks the acyl adenylate, forming acyl CoA and a molecule of AMP. Each of these two steps is reversible under biological conditions, save for the additional hydrolysis of PPi by inorganic pyrophosphatase.[19] This coupled hydrolysis provides the driving force for the overall forward activation reaction, and serves as a source of inorganic phosphate used in other biological processes.
Evolution
Examination of prokaryotic and eukaryotic forms of soluble inorganic pyrophosphatase (sPPase, Pfam PF00719) has shown that they differ significantly in both amino acid sequence, number of residues, and oligomeric organization. Despite differing structural components, recent work has suggested a large degree of evolutionary conservation of active site structure as well as reaction mechanism, based on kinetic data.[20] Analysis of approximately one million genetic sequences taken from organisms in the Sargasso Sea identified a 57 residue sequence within the regions coding for proton-pumping inorganic pyrophosphatase (H+-PPase) that appears to be highly conserved; this region primarily consisted of the four early amino acid residues Gly, Ala, Val and Asp, suggesting an evolutionarily ancient origin for the protein.[21]
References
- ↑ "Inorganic polyphosphates in biology: structure, metabolism, and function". Bacteriological Reviews 30 (4): 772–94. December 1966. doi:10.1128/MMBR.30.4.772-794.1966. PMID 5342521.
- ↑ "Inorganic pyrophosphate generation and disposition in pathophysiology". American Journal of Physiology. Cell Physiology 281 (1): C1–C11. July 2001. doi:10.1152/ajpcell.2001.281.1.C1. PMID 11401820.
- ↑ "Role of inorganic pyrophosphatase in the mechanism of action of parathyroid hormone and calcitonin". Endocrinology 89 (3): 852–8. September 1971. doi:10.1210/endo-89-3-852. PMID 4327778.
- ↑ "Parathyroid hormone - a bone anabolic and catabolic agent". Current Opinion in Pharmacology 5 (6): 612–7. December 2005. doi:10.1016/j.coph.2005.07.004. PMID 16181808.
- ↑ Nelson, David L.; Cox, Michael M. (2000). Lehninger Principles of Biochemistry, 3rd ed.. New York: Worth Publishers. pp. 937. ISBN 1-57259-153-6. https://archive.org/details/lehningerprincip01lehn/page/937.
- ↑ "PYP-1, inorganic pyrophosphatase, is required for larval development and intestinal function in C. elegans". FEBS Letters 581 (28): 5445–53. November 2007. doi:10.1016/j.febslet.2007.10.047. PMID 17981157.
- ↑ "Inorganic polyphosphate induces osteoblastic differentiation". Journal of Dental Research 89 (5): 504–9. May 2010. doi:10.1177/0022034510363096. PMID 20332330.
- ↑ "Functional complementation of yeast cytosolic pyrophosphatase by bacterial and plant H+-translocating pyrophosphatases". Proc. Natl. Acad. Sci. U.S.A. 99 (25): 15914–9. December 2002. doi:10.1073/pnas.242625399. PMID 12451180. Bibcode: 2002PNAS...9915914P.
- ↑ "H+ -PPases: a tightly membrane-bound family". FEBS Lett. 457 (3): 527–33. September 1999. doi:10.1016/S0014-5793(99)90617-8. PMID 10523139.
- ↑ "Crystal structure of inorganic pyrophosphatase from Thermus thermophilus". Protein Science 3 (7): 1098–107. July 1994. doi:10.1002/pro.5560030713. PMID 7920256.
- ↑ "Cloning and expression profile of human inorganic pyrophosphatase". Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression 1447 (2–3): 133–6. October 1999. doi:10.1016/s0167-4781(99)00175-x. PMID 10542310.
- ↑ "Assignment of the inorganic pyrophosphatase gene locus (PP) to chromosome 10 in man". Cytogenetics and Cell Genetics 16 (1–5): 201–3. 1976. doi:10.1159/000130590. PMID 975879.
- ↑ "PPA2 pyrophosphatase (inorganic) 2 [Homo sapiens (human)"]. https://www.ncbi.nlm.nih.gov/gene/27068.
- ↑ 14.0 14.1 "DFT study on the mechanism of Escherichia coli inorganic pyrophosphatase". The Journal of Physical Chemistry B 113 (18): 6505–10. May 2009. doi:10.1021/jp810003w. PMID 19366250.
- ↑ "Binding of substrate at the effector site of pyrophosphatase increases the rate of its hydrolysis at the active site". Biochemistry. Biokhimiia 72 (1): 68–76. January 2007. doi:10.1134/s0006297907010087. PMID 17309439.
- ↑ "ATP as effector of inorganic pyrophosphatase of Escherichia coli. Identification of the binding site for ATP". Biochemistry. Biokhimiia 72 (1): 93–9. January 2007. doi:10.1134/s0006297907010117. PMID 17309442.
- ↑ "[Two-stage mechanism of the fluoride inhibition of inorganic pyrophosphatase using the fluoride ion]" (in ru). Biokhimiia 48 (10): 1643–53. October 1983. PMID 6139128.
- ↑ "Cellular signaling mediated by calphoglin-induced activation of IPP and PGM". Biochemical and Biophysical Research Communications 325 (1): 203–14. December 2004. doi:10.1016/j.bbrc.2004.10.021. PMID 15522220.
- ↑ 19.0 19.1 19.2 "Roles of phosphatidate phosphatase enzymes in lipid metabolism". Trends in Biochemical Sciences 31 (12): 694–9. December 2006. doi:10.1016/j.tibs.2006.10.003. PMID 17079146.
- ↑ "Evolutionary conservation of the active site of soluble inorganic pyrophosphatase". Trends in Biochemical Sciences 17 (7): 262–6. July 1992. doi:10.1016/0968-0004(92)90406-y. PMID 1323891.
- ↑ "Analysis of ancient sequence motifs in the H-PPase family". The FEBS Journal 273 (22): 5183–93. November 2006. doi:10.1111/j.1742-4658.2006.05514.x. PMID 17054711.
External links
- pyrophosphatases at the US National Library of Medicine Medical Subject Headings (MeSH)
Further reading
- "Purification and properties of yeast pyrophosphatase". The Biochemical Journal 38 (5): 394–8. 1944. doi:10.1042/bj0380394. PMID 16747821.
- "Crystalline inorganic pyrophosphatase isolated from baker's yeast". The Journal of General Physiology 35 (3): 423–50. January 1952. doi:10.1085/jgp.35.3.423. PMID 14898026.
- "Molecular cloning and sequence of cDNA encoding the pyrophosphate-energized vacuolar membrane proton pump of Arabidopsis thaliana". Proceedings of the National Academy of Sciences of the United States of America 89 (5): 1775–9. March 1992. doi:10.1073/pnas.89.5.1775. PMID 1311852. Bibcode: 1992PNAS...89.1775S.
- "A thermostable vacuolar-type membrane pyrophosphatase from the archaeon Pyrobaculum aerophilum: implications for the origins of pyrophosphate-energized pumps". FEBS Letters 460 (3): 505–12. November 1999. doi:10.1016/S0014-5793(99)01404-0. PMID 10556526.
 |

