Biology:EF-G
| Protein-synthesizing GTPase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC number | 3.6.5.3 | ||||||||
| Alt. names | Elongation factor G, EF-G | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
| Translation elongation factor EFG/EF2 | |
|---|---|
| Identifiers | |
| Symbol | Transl_elong_EFG/EF2 |
| InterPro | IPR004540 |
| SCOP2 | 1n0u / SCOPe / SUPFAM |
| EFG/EF2, domain IV | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | EFG_IV | ||||||||
| Pfam | PF03764 | ||||||||
| Pfam clan | CL0329 | ||||||||
| SMART | SM00889 | ||||||||
| CDD | cd01434 | ||||||||
| |||||||||
EF-G (elongation factor G, historically known as translocase) is a prokaryotic elongation factor involved in protein translation. As a GTPase, EF-G catalyzes the movement (translocation) of transfer RNA (tRNA) and messenger RNA (mRNA) through the ribosome.[1]
Structure
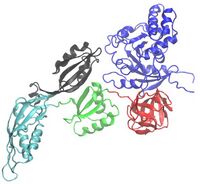
Encoded by the fusA gene on the str operon,[2] EF-G is made up of 704 amino acids that form 5 domains, labeled Domain I through Domain V. Domain I may be referred to as the G-domain or as Domain I(G), since it binds to and hydrolyzes guanosine triphosphate (GTP). Domain I also helps EF-G bind to the ribosome, and contains the N-terminal of the polypeptide chain.[3][4] Domain IV is important for translocation, as it undergoes a significant conformational change and enters the A site on the 30S ribosomal subunit, pushing the mRNA and tRNA molecules from the A site to the P site.[5]
The five domains may be also separated into two super-domains. Super-domain I consists of Domains I and II, and super-domain II consists of Domains III - IV. Throughout translocation, super-domain I will remain relatively unchanged, as it is responsible for binding tightly to the ribosome. However, super-domain II will undergo a large rotational motion from the pre-translocational (PRE) state to the post-translocational (POST) state. Super-domain I is similar to the corresponding sections of EF-Tu.[6][7][8] Super-domain II in the POST state mimics the tRNA molecule of the EF-Tu • GTP • aa-tRNA ternary complex.[9]

EF-G on the ribosome
Binding to L7/L12
L7/L12 is only a multicopy protein on the large ribosomal subunit of the bacterial ribosome that binds to certain GTPases, like Initiation Factor 2, Elongation factor-Tu, Release Factor 3, and EF-G.[10] Specifically, the C-terminal of L7/L12 will bind to EF-G and is necessary for GTP hydrolysis.[4]
Interaction with the GTPase Associated Center
The GTPase Associated Center (GAC) is a region on the large ribosomal subunit that consists of two smaller regions of 23S ribosomal RNA called the L11 stalk and the sarcin-ricin loop (SRL).[11] As a highly conserved rRNA loop in evolution, the SRL is critical in helping GTPases bind to the ribosome, but is not essential for GTP hydrolysis. There is some evidence to support that a phosphate oxygen in the A2662 residue of the SRL may help hydrolyze GTP.[12]
Function in protein elongation
EF-G catalyzes the translocation of the tRNA and mRNA down the ribosome at the end of each round of polypeptide elongation.[1] In this process, the peptidyl transferase center (PTC) has catalyzed the formation of a peptide bond between amino acids, moving the polypeptide chain from the P site tRNA to the A site tRNA. The 50S and 30S ribosomal subunits are now allowed to rotate relative to each other by approximately 7°.[13][14] The subunit rotation is coupled with the movement of the 3' ends of both tRNA molecules on the large subunit from the A and P sites to the P and E sites, respectively, while the anticodon loops remain unshifted. This rotated ribosomal intermediate, in which the first tRNA occupies a hybrid A/P position and the second tRNA occupies a hybrid P/E position is a substrate for EF-G-GTP.[1][13]
As a GTPase, EF-G binds to the rotated ribosome near the A site in its GTP-bound state, and hydrolyzes GTP, releasing GDP and inorganic phosphate:
The hydrolysis of GTP allows for a large conformational change within EF-G, forcing the A/P tRNA to fully occupy the P site, the P/E tRNA to fully occupy the E site (and exit the ribosome complex), and the mRNA to shift three nucleotides down relative to the ribosome. The GDP-bound EF-G molecule then dissociates from the complex, leaving another free A-site where the elongation cycle can start again.[1][15]

Function in protein termination
Protein elongation continues until a stop codon appears on the mRNA. A Class I release factor (RF1 or RF2) binds to the stop codon, which induces hydrolysis of the tRNA-peptide bond in the P site, allowing the newly-formed protein to exit the ribosome. The nascent peptide continues to fold and leaves the 70S ribosome, the mRNA, the deacylated tRNA (P site), and the Class I release factor (A site).[16][17]
In a GTP-dependent manner, the subsequent recycling is catalyzed by a Class II release factor named RF3/prfC, Ribosome recycling factor (RRF), Initiation Factor 3 (IF3) and EF-G. The protein RF3 releases the Class I release factor so that it may occupy the ribosomal A site. EF-G hydrolyzes GTP and undergoes a large conformational change to push RF3 down the ribosome, which occurs alongside tRNA dissociation and promotes the ribosomal subunit rotation. This motion actively splits the B2a/B2b bridge, which connects the 30S and the 50S subunits, so that the ribosome can split.[16] IF3 then isolates the 30S subunit to prevent re-association of the large and small subunits.[18]
Clinical significance
EF-G in pathogenic bacteria can be inhibited by antibiotics that prevent EF-G from binding to the ribosome,[19] carrying out translocation[20] or dissociating from the ribosome.[21]
For example, the antibiotic thiostrepton prevents EF-G from binding stably to the ribosome,[19] while the antibiotics dityromycin and GE82832 inhibit the activity of EF-G by preventing the translocation of the A site tRNA. Dityromycin and GE82832 do not affect the binding of EF-G to the ribosome, however.[20]
The antibiotic fusidic acid is known to inhibit Staphylococcus aureus and other bacteria by binding to EF-G after one translocation event on the ribosome, preventing EF-G from dissociating.[21][22] However, some bacterial strains have developed resistance to fusidic acid due to point mutations in the fusA gene, which prevents fusidic acid from binding to EF-G.[23][24]
Evolution
EF-G has a complex evolutionary history, with numerous paralogous versions of the factor present in bacteria, suggesting subfunctionalization of different EF-G variants.[25]
Elongation factors exist in all three domains of life with similar function on the ribosome. The eukaryotic and archeal homologs of EF-G are eEF2 and aEF2, respectively. In bacteria (and some archaea), the fusA gene that encodes EF-G is found within the conserved str gene with the sequence 5′ - rpsL - rpsG - fusA - tufA - 3′.[2] However, two other major forms of EF-G exist in some species of Spirochaetota, Planctomycetota, and δ-Proteobacteria (which has since been split and renamed Bdellovibrionota, Myxococcota, and Thermodesulfobacteriota), which form the spd group of bacteria that have elongation factors spdEFG1 and spdEFG2.[25][26]
From spdEFG1 and spdEFG2 evolved the mitochondrial elongation factors mtEFG1 (GFM1) and mtEFG2 (GFM2), respectively.[25][26] The two roles of EF-G in elongation and termination of protein translation are split amongst the mitochondrial elongation factors, with mtEFG1 responsible for translocation and mtEFG2 responsible for termination and ribosomal recycling with mitochondrial RRF.
See also
- Prokaryotic elongation factors
- EF-Ts (elongation factor thermo stable)
- EF-Tu (elongation factor thermo unstable)
- EF-P (elongation factor P)
- eEF2 (eukaryotic elongation factor 2)
- Protein translation
- GTPase
References
- ↑ 1.0 1.1 1.2 1.3 Shoji, S; Walker, SE; Fredrick, K (2009). "Ribosomal translocation: one step closer to the molecular mechanism". ACS Chem Biol 4 (2): 93–107. doi:10.1021/cb8002946. PMID 19173642.
- ↑ 2.0 2.1 Post, L. E.; Nomura, M. (1980-05-25). "DNA sequences from the str operon of Escherichia coli". The Journal of Biological Chemistry 255 (10): 4660–4666. doi:10.1016/S0021-9258(19)85545-X. ISSN 0021-9258. PMID 6989816.
- ↑ Liu, Kaixian; Rehfus, Joseph E.; Mattson, Elliot; Kaiser, Christian M. (2017-07-01). "The ribosome destabilizes native and non-native structures in a nascent multidomain protein" (in en). Protein Science 26 (7): 1439–1451. doi:10.1002/pro.3189. ISSN 1469-896X. PMID 28474852.
- ↑ 4.0 4.1 Carlson, Markus A.; Haddad, Bassam G.; Weis, Amanda J.; Blackwood, Colby S.; Shelton, Catherine D.; Wuerth, Michelle E.; Walter, Justin D.; Spiegel, Paul Clint (2017-06-01). "Ribosomal protein L7/L12 is required for GTPase translation factors EF-G, RF3, and IF2 to bind in their GTP state to 70S ribosomes" (in en). The FEBS Journal 284 (11): 1631–1643. doi:10.1111/febs.14067. ISSN 1742-4658. PMID 28342293.
- ↑ Salsi, Enea; Farah, Elie; Dann, Jillian; Ermolenko, Dmitri N. (2014). "Following movement of domain IV of elongation factor G during ribosomal translocation". Proceedings of the National Academy of Sciences 111 (42): 15060–15065. doi:10.1073/pnas.1410873111. PMID 25288752. Bibcode: 2014PNAS..11115060S.
- ↑ Lin, Jinzhong; Gagnon, Matthieu G.; Bulkley, David; Steitz, Thomas A. (2015). "Conformational Changes of Elongation Factor G on the Ribosome during tRNA Translocation". Cell 160 (1–2): 219–227. doi:10.1016/j.cell.2014.11.049. PMID 25594181.
- ↑ Li, Wen; Trabuco, Leonardo G.; Schulten, Klaus; Frank, Joachim (2011-05-01). "Molecular dynamics of EF-G during translocation" (in en). Proteins: Structure, Function, and Bioinformatics 79 (5): 1478–1486. doi:10.1002/prot.22976. ISSN 1097-0134. PMID 21365677.
- ↑ Zhang, Dejiu; Yan, Kaige; Zhang, Yiwei; Liu, Guangqiao; Cao, Xintao; Song, Guangtao; Xie, Qiang; Gao, Ning et al. (2015). "New insights into the enzymatic role of EF-G in ribosome recycling". Nucleic Acids Research 43 (21): 10525–33. doi:10.1093/nar/gkv995. PMID 26432831.
- ↑ Nyborg, J.; Nissen, P.; Kjeldgaard, M.; Thirup, S.; Polekhina, G.; Clark, B. F. (March 1996). "Structure of the ternary complex of EF-Tu: macromolecular mimicry in translation". Trends in Biochemical Sciences 21 (3): 81–82. doi:10.1016/S0968-0004(96)30008-X. ISSN 0968-0004. PMID 8882578.
- ↑ Mandava, C. S.; Peisker, K.; Ederth, J.; Kumar, R.; Ge, X.; Szaflarski, W.; Sanyal, S. (2011-11-18). "Bacterial ribosome requires multiple L12 dimers for efficient initiation and elongation of protein synthesis involving IF2 and EF-G". Nucleic Acids Research 40 (5): 2054–2064. doi:10.1093/nar/gkr1031. ISSN 0305-1048. PMID 22102582.
- ↑ Maklan, E. J. (2012). Genetic and Biochemical Analysis of the GTPase Associated Center of the Ribosome. UC Santa Cruz. Merritt ID: ark:/13030/m5js9t4d. Retrieved from https://escholarship.org/uc/item/7gh9v43h
- ↑ Shi, Xinying; Khade, Prashant K.; Sanbonmatsu, Karissa Y.; Joseph, Simpson (2012). "Functional Role of the Sarcin–Ricin Loop of the 23S rRNA in the Elongation Cycle of Protein Synthesis". Journal of Molecular Biology 419 (3–4): 125–138. doi:10.1016/j.jmb.2012.03.016. PMID 22459262.
- ↑ 13.0 13.1 Choi, Junhong; Puglisi, Joseph D. (2017). "Three tRNAs on the ribosome slow translation elongation". Proceedings of the National Academy of Sciences 114 (52): 13691–13696. doi:10.1073/pnas.1719592115. PMID 29229848.
- ↑ Guo, Z.; Noller, H. F. (2012). "Rotation of the head of the 30S ribosomal subunit during mRNA translocation". Proceedings of the National Academy of Sciences 109 (50): 20391–20394. doi:10.1073/pnas.1218999109. PMID 23188795. Bibcode: 2012PNAS..10920391G.
- ↑ da Cunha, CE; Belardinelli, R; Peske, F; Holtkamp, W; Wintermeyer, W; Rodnina, MV (2013). "Dual use of GTP hydrolysis by elongation factor G on the ribosome". Translation 1 (1): e24315. doi:10.4161/trla.24315. PMID 26824016.
- ↑ 16.0 16.1 Das, Debasis; Samanta, Dibyendu; Bhattacharya, Arpita; Basu, Arunima; Das, Anindita; Ghosh, Jaydip; Chakrabarti, Abhijit; Gupta, Chanchal Das (2017-01-18). "A Possible Role of the Full-Length Nascent Protein in Post-Translational Ribosome Recycling" (in en). PLOS ONE 12 (1): e0170333. doi:10.1371/journal.pone.0170333. ISSN 1932-6203. PMID 28099529. Bibcode: 2017PLoSO..1270333D.
- ↑ "Splitting of the posttermination ribosome into subunits by the concerted action of RRF and EF-G". Molecular Cell 18 (6): 675–686. 2005. doi:10.1016/j.molcel.2005.05.016. PMID 15949442.
- ↑ Hirokawa, Go; Nijman, Romana M.; Raj, V. Samuel; Kaji, Hideko; Igarashi, Kazuei; Kaji, Akira (2005-08-01). "The role of ribosome recycling factor in dissociation of 70S ribosomes into subunits" (in en). RNA 11 (8): 1317–1328. doi:10.1261/rna.2520405. ISSN 1355-8382. PMID 16043510.
- ↑ 19.0 19.1 Walter, Justin D.; Hunter, Margaret; Cobb, Melanie; Traeger, Geoff; Spiegel, P. Clint (2012-01-01). "Thiostrepton inhibits stable 70S ribosome binding and ribosome-dependent GTPase activation of elongation factor G and elongation factor 4" (in en). Nucleic Acids Research 40 (1): 360–370. doi:10.1093/nar/gkr623. ISSN 0305-1048. PMID 21908407.
- ↑ 20.0 20.1 Bulkley, David; Brandi, Letizia; Polikanov, Yury S.; Fabbretti, Attilio; O’Connor, Michael; Gualerzi, Claudio O.; Steitz, Thomas A. (2014). "The Antibiotics Dityromycin and GE82832 Bind Protein S12 and Block EF-G-Catalyzed Translocation". Cell Reports 6 (2): 357–365. doi:10.1016/j.celrep.2013.12.024. PMID 24412368.
- ↑ 21.0 21.1 Belardinelli, Riccardo; Rodnina, Marina V. (2017-09-05). "Effect of Fusidic Acid on the Kinetics of Molecular Motions During EF-G-Induced Translocation on the Ribosome" (in En). Scientific Reports 7 (1): 10536. doi:10.1038/s41598-017-10916-8. ISSN 2045-2322. PMID 28874811. Bibcode: 2017NatSR...710536B.
- ↑ Koripella, Ravi Kiran; Chen, Yang; Peisker, Kristin; Koh, Cha San; Selmer, Maria; Sanyal, Suparna (2012). "Mechanism of Elongation Factor-G-mediated Fusidic Acid Resistance and Fitness Compensation inStaphylococcus aureus". Journal of Biological Chemistry 287 (36): 30257–30267. doi:10.1074/jbc.m112.378521. PMID 22767604.
- ↑ "Hyper-susceptibility of a fusidic acid-resistant mutant of Salmonella to different classes of antibiotics". FEMS Microbiology Letters 247 (2): 215–20. June 2005. doi:10.1016/j.femsle.2005.05.007. PMID 15935566.
- ↑ "Fusidic acid-resistant EF-G perturbs the accumulation of ppGpp". Molecular Microbiology 37 (1): 98–107. July 2000. doi:10.1046/j.1365-2958.2000.01967.x. PMID 10931308.
- ↑ 25.0 25.1 25.2 G C Atkinson; S L Baldauf (2011). "Evolution of elongation factor G and the origins of mitochondrial and chloroplast forms". Molecular Biology and Evolution 28 (3): 1281–92. doi:10.1093/molbev/msq316. PMID 21097998.
- ↑ 26.0 26.1 Margus, Tõnu; Remm, Maido; Tenson, Tanel (2011-08-04). "A Computational Study of Elongation Factor G (EFG) Duplicated Genes: Diverged Nature Underlying the Innovation on the Same Structural Template" (in en). PLOS ONE 6 (8): e22789. doi:10.1371/journal.pone.0022789. ISSN 1932-6203. PMID 21829651. Bibcode: 2011PLoSO...622789M.
Further reading
- Carbone, Christine E.; Loveland, Anna B.; Gamper, Howard B.; Hou, Ya-Ming; Demo, Gabriel; Korostelev, Andrei A. (December 2021). "Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP". Nature Communications 12 (1): 7236. doi:10.1038/s41467-021-27415-0.
External links
- Peptide+Elongation+Factor+G at the US National Library of Medicine Medical Subject Headings (MeSH)
 |
