Biology:SUMO enzymes
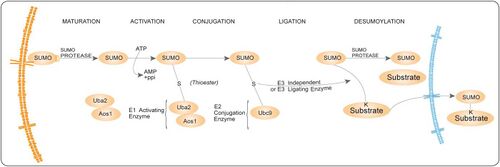
SUMO enzymatic cascade catalyzes the dynamic posttranslational modification process of sumoylation (i.e. transfer of SUMO protein to other proteins). The Small Ubiquitin-related Modifier, SUMO-1,[1][2] is a ubiquitin-like family member that is conjugated to its substrates through three discrete enzymatic steps (see the figure on the right): activation, involving the E1 enzyme (SAE1/SAE2);[3] conjugation, involving the E2 enzyme (UBE2I);[4][5] substrate modification, through the cooperation of the E2 and E3[6] protein ligases.[7]
SUMO pathway modifies hundreds of proteins that participate in diverse cellular processes.[8] SUMO pathway is the most studied ubiquitin-like pathway that regulates a wide range of cellular events,[9] evidenced by a large number of sumoylated proteins identified in more than ten large-scale studies.[10][11][12][13][14][15][16][17][18][19][20][excessive citations]
See also
References
- ↑ Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004)
- ↑ Melchior, F. SUMO–nonclassical ubiquitin. Annu. Rev. Cell Dev. Biol. 16, 591–626 (2000)
- ↑ Boggio R et al., A mechanism for inhibiting the SUMO pathway. Mol Cell. 2004 Nov 19;16(4):549-61
- ↑ Bernier-Villamor, V. , Sampson, D.A. , Matunis, M.J. & Lima, C.D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356
- ↑ Lin, D. et al. Identification of a substrate recognition site on Ubc9. J. Biol. Chem. 277, 21740–21748 (2002)
- ↑ Reverter, D. & Lima, C.D. Insights into E3 ligase activity revealed by a SUMO-RanGAP1-Ubc9-Nup358 complex. Nature 435, 687–692
- ↑ Melchior, F. , Schergaut, M. & Pichler, A. SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem. Sci. 28, 612–618
- ↑ Johnson ES. Protein modification by SUMO. Annu Rev Biochem 2004;73:355–82
- ↑ O. Kerscher, R. Felberbaum and M. Hochstrasser, Modification of proteins by ubiquitin and ubiquitin-like proteins, Annu. Rev. Cell Dev. Biol. (2006) Jun 5, Electronic publication ahead of print
- ↑ W. Zhou, J.J. Ryan and H. Zhou, Global analyses of sumoylated proteins in Saccharomyces cerevisiae. Induction of protein sumoylation by cellular stresses, J. Biol. Chem. 279 (2004), pp. 32262–32268
- ↑ T. Li, E. Evdokimov, R.F. Shen, C.C. Chao, E. Tekle, T. Wang, E.R. Stadtman, D.C. Yang and P.B. Chock, Sumoylation of heterogeneous nuclear ribonucleoproteins, zinc finger proteins, and nuclear pore complex proteins: a proteomic analysis, Proc. Natl. Acad. Sci. U. S. A. 101 (2004), pp. 8551–8556
- ↑ J.A. Wohlschlegel, E.S. Johnson, S.I. Reed and J.R. Yates, 3rd, Global analysis of protein sumoylation in Saccharomyces cerevisiae, J. Biol. Chem. (2004)
- ↑ V.G. Panse, U. Hardeland, T. Werner, B. Kuster and E. Hurt, A proteome-wide approach identifies sumoylated substrate proteins in yeast, J. Biol. Chem. (2004)
- ↑ A.C. Vertegaal, S.C. Ogg, E. Jaffray, M.S. Rodriguez, R.T. Hay, J.S. Andersen, M. Mann and A.I. Lamond, A proteomic study of SUMO-2 target proteins, J. Biol. Chem. 279 (2004), pp. 33791–33798
- ↑ [72] C. Denison, A.D. Rudner, S.A. Gerber, C.E. Bakalarski, D. Moazed and S.P. Gygi, A proteomic strategy for gaining insights into protein sumoylation in yeast, Mol. Cell. Proteomics 4 (2005), pp. 246–254
- ↑ J.T. Hannich, A. Lewis, M.B. Kroetz, S.J. Li, H. Heide, A. Emili and M. Hochstrasser, Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae, J. Biol. Chem. 280 (2005), pp. 4102–4110
- ↑ Y. Zhao, S.W. Kwon, A. Anselmo, K. Kaur and M.A. White, Broad spectrum identification of cellular small ubiquitin-related modifier (SUMO) substrate proteins, J. Biol. Chem. 279 (2004), pp. 20999–21002
- ↑ L.L. Manza, S.G. Codreanu, S.L. Stamer, D.L. Smith, K.S. Wells, R.L. Roberts and D.C. Liebler, Global shifts in protein sumoylation in response to electrophile and oxidative stress, Chem. Res. Toxicol. 17 (2004), pp. 1706–1715
- ↑ T. Li, R. Santockyte, R.F. Shen, E. Tekle, G. Wang, D.C. Yang and P.B. Chock, A general approach for investigating enzymatic pathways and substrates for ubiquitin-like modifiers, Arch. Biochem. Biophys. 453 (2006), pp. 70–74
- ↑ G. Rosas-Acosta, W.K. Russell, A. Deyrieux, D.H. Russell and V.G. Wilson, A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers, Mol. Cell. Proteomics 4 (2005), pp. 56–72
 |

