Biology:Glycoside hydrolase family 10
| Glycoside hydrolase, family 10 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
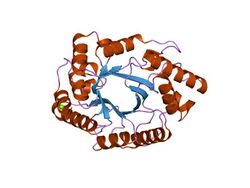 crystal structure of the catalytic domain of xylanase a from streptomyces halstedii jm8 | |||||||||
| Identifiers | |||||||||
| Symbol | Glyco_hydro_10 | ||||||||
| Pfam | PF00331 | ||||||||
| Pfam clan | CL0058 | ||||||||
| InterPro | IPR001000 | ||||||||
| PROSITE | PDOC00510 | ||||||||
| SCOP2 | 2exo / SCOPe / SUPFAM | ||||||||
| CAZy | GH10 | ||||||||
| |||||||||
In molecular biology, Glycoside hydrolase family 10 is a family of glycoside hydrolases.
Glycoside hydrolases EC 3.2.1. are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycoside hydrolases, based on sequence similarity, has led to the definition of >100 different families.[1][2][3] This classification is available on the CAZy web site,[4][5] and also discussed at CAZypedia, an online encyclopedia of carbohydrate active enzymes.[6][7]
Glycoside hydrolase family 10 CAZY GH_10 comprises enzymes with a number of known activities; xylanase (EC 3.2.1.8); endo-1,3-beta-xylanase (EC 3.2.1.32); cellobiohydrolase (EC 3.2.1.91). These enzymes were formerly known as cellulase family F.
The microbial degradation of cellulose and xylans requires several types of enzymes such as endoglucanases (EC 3.2.1.4), cellobiohydrolases (EC 3.2.1.91) (exoglucanases), or xylanases (EC 3.2.1.8).[8][9] Fungi and bacteria produces a spectrum of cellulolytic enzymes (cellulases) and xylanases which, on the basis of sequence similarities, can be classified into families. One of these families is known as the cellulase family F[10] or as the glycosyl hydrolases family 10.[11]
References
- ↑ "Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases". Proceedings of the National Academy of Sciences of the United States of America 92 (15): 7090–4. July 1995. doi:10.1073/pnas.92.15.7090. PMID 7624375. Bibcode: 1995PNAS...92.7090H.
- ↑ "Structures and mechanisms of glycosyl hydrolases". Structure 3 (9): 853–9. September 1995. doi:10.1016/S0969-2126(01)00220-9. PMID 8535779.
- ↑ "Updating the sequence-based classification of glycosyl hydrolases". The Biochemical Journal 316 (Pt 2): 695–6. June 1996. doi:10.1042/bj3160695. PMID 8687420.
- ↑ "Home" (in en). http://www.cazy.org/.
- ↑ "The carbohydrate-active enzymes database (CAZy) in 2013". Nucleic Acids Research 42 (Database issue): D490-5. January 2014. doi:10.1093/nar/gkt1178. PMID 24270786.
- ↑ "Glycoside Hydrolase Family 10" (in en). http://www.cazypedia.org/index.php/Glycoside_Hydrolase_Family_10.
- ↑ CAZypedia Consortium (December 2018). "Ten years of CAZypedia: a living encyclopedia of carbohydrate-active enzymes". Glycobiology 28 (1): 3–8. doi:10.1093/glycob/cwx089. PMID 29040563.
- ↑ "Molecular biology of cellulose degradation". Annual Review of Microbiology 44: 219–48. 1990. doi:10.1146/annurev.mi.44.100190.001251. PMID 2252383.
- ↑ "Domains in microbial beta-1, 4-glycanases: sequence conservation, function, and enzyme families". Microbiological Reviews 55 (2): 303–15. June 1991. doi:10.1128/MMBR.55.2.303-315.1991. PMID 1886523.
- ↑ "Cellulase families revealed by hydrophobic cluster analysis". Gene 81 (1): 83–95. September 1989. doi:10.1016/0378-1119(89)90339-9. PMID 2806912.
- ↑ "A classification of glycosyl hydrolases based on amino acid sequence similarities". The Biochemical Journal 280 (2): 309–16. December 1991. doi:10.1042/bj2800309. PMID 1747104.
 |

