Biology:Glucosamine-phosphate N-acetyltransferase
| glucosamine 6-phosphate N-acetyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
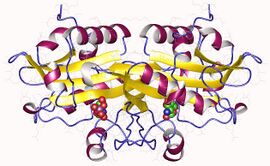 Glucosamine 6-phosphate N-acetyltransferase 1, homodimer, Human | |||||||||
| Identifiers | |||||||||
| EC number | 2.3.1.4 | ||||||||
| CAS number | 9031-91-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
In enzymology, glucosamine-phosphate N-acetyltransferase (GNA) (EC 2.3.1.4) is an enzyme that catalyzes the transfer of an acetyl group from acetyl-CoA to the primary amine in glucosamide-6-phosphate, generating a free CoA and N-acetyl-D-glucosamine-6-phosphate.[1]
This enzyme belongs to the family of transferases, a group of enzymes that transfers a very specific functional group, in this case acetyl, from a donor to a receptor. Specifically, this enzyme can be characterized as part of the acyltransferases family, since it involves the transfer of a general acyl group with a methyl as the substituent.
Nomenclature
The systematic name of this enzyme class is acetyl-CoA:D-glucosamine-6-phosphate N-acetyltransferase. Other names in common use include phosphoglucosamine transacetylase, phosphoglucosamine acetylase, glucosamine-6-phosphate acetylase, D-glucosamine-6-P N-acetyltransferase, aminodeoxyglucosephosphate acetyltransferase, glucosamine 6-phosphate acetylase, glucosamine 6-phosphate N-acetyltransferase, N-acetylglucosamine-6-phosphate synthase, phosphoglucosamine N-acetylase, glucosamine-phosphate N-acetyltransferase, and glucosamine-6-phosphate N-acetyltransferase.
Function
This enzyme is part of the hexosamine biosynthesis pathway[2] (HBP), which is one of the glucose processing pathways in the general metabolism. This pathway shares the initial two steps with glycolysis and diverges only a small portion of glucose flux from this more traditional glycolytic pathway. Therefore, it is favored when there is negative feedback regulation on glycolysis, as in the case of large amounts of free fatty acids. The end product of this pathway is UDP-N-Acetylglucosamine, which is involved in the modification of complex molecules such as glycolipids, proteoglycans[3] and glycoproteins. This end product acts as a carrier of N-Acetylglucosamine, which is the monomeric unit of chitin,[4] a structural polymer that composes the shells of crustaceans and insects, as well as the cell wall of fungi. Furthermore, N-Acetylglucosamine is also a unit of the peptidoglycan polymer that composes the bacteria cell wall[5] along with N-acetylmuramic disaccharide.
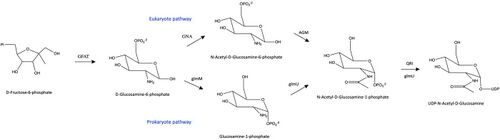
More specifically, the GNA enzyme catalyzes the fourth step of the HBP pathway in eukaryotes, promoting a carbon transfer from Acetyl-CoA to the other substrate, D-Glucosamine-6-phosphate which will finally yield UDP-N-Acetylglucosamine. This is a small, but an important chemical step that is crucial to the properties of the sub-products of this metabolic pathway. The acetylation is carried out until the very end product of the hexamine pathway, and is very characteristic of the polymers formed with N-Acetylglucosamine. For instance, it constitutes one of the main difference in the molecular structure of chitin and cellulose,[7] and explains many of the physical and chemical properties of these polymers. In the case of chitin, for example, computational studies have found that the acylation contributes to the formation of hydrogen bonds that stabilize the crystalline structure of this polymer, providing greater resistance to fracture.[8]
Nevertheless, in prokaryotic metabolism, the hexosamine biosynthesis pathway follows a different reaction step, in which a different enzyme acts upon the same characteristic substrates[6] (Figure 1). In prokaryotes, the phosphate transfer from the 6-carbon to the 1-carbon takes place before the acylation, such that the substrate of carbon-adding reaction is Glucosamine-1-phosphate rather than D-glucosamine-6-phosphate. This time, the enzyme responsible for the acetylation is the bifunctional protein glmU (N-Acetylglucosamine-1-phosphate uridyltransferase),[9] that also catalyzes the addition of UDP to the phosphate group on N-Acetyl-D-Glucosamine-1-Phosphate.
In humans, glucosamine-phosphate N-acetyltransferase is a dimer with two identical subunits,[10] and is encoded in the gene GNPNAT[11] (HGNC Symbol). More specifically, the enzyme is strongly expressed in the liver, stomach and gastrointestinal tract tissues, and within the cell, it is located in endosomes and in the Golgi apparatus (by manual annotation).[11]
Mechanism
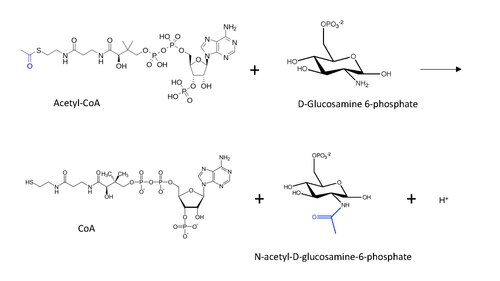
The molecular structure of the reaction catalyzed by GNA is shown below, with the transferred acetyl group in blue.
The general reaction mechanism postulated for protein N-end acetylation (inspired by lysine acetylation mechanism) with Acetyl-CoA involves a nucleophilic attack of the amino group (in this case from D-Glucosamine-6-phosphate) on the terminal carbonyl in the carbon transfer, leading to the formation of a carbon tetrahedral intermediate.[13] The reaction proceeds with the restoration of the carbonyl by removing the CoA as a leaving group, such that now the acetyl group is connected to the amino group in the other substrate.
Specifically for this N-acetyltransferase catalysts, studies with S. cerevisiae GNA enzyme have shown that some specific amino acids contribute to substrate binding, increased nucleophilicity of the amino group and finally catalysis, which supports the postulated mechanism described above.[14] Glu98, Asp99, and Ile100 polarize the carbonyl bond in the Acetyl-CoA, increasing the carbon electrophilicity as well as stabilizing the carbon tetrahedral intermediate. Tyr143 is responsible for stabilizing the thiolate anion, favoring the S-CoA as a leaving group from the tetrahedral carbon. Finally, Asp134 enhances the nucleophilicity of the amino group in D-Glucosamide-6-phosphate by donating electron density to the nitrogen atom. In a different organism, C. albicans, a similar set of amino acids were found to be essential to the catalytic activity,[15] respectively the Glu88-Asp-89-Ile90 system, Asp125 and Tyr133.
Structure
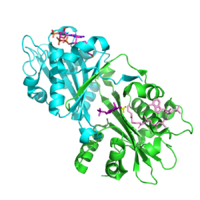
As of late 2019, 13 structures have been solved for this class of enzymes in different species, with PDB access codes 1I12 (Saccharomyces cerevisiae), 1I1D (Saccharomyces cerevisiae), 1I21 (Saccharomyces cerevisiae), 2HUZ (Homo sapiens), 2O28 (Homo sapiens), 4AG7 (Caenorhabditis elegans), among others.
Figure 3 shows the proposed crystal structure of GNA in humans,[17] with each catalytic subunit in a different color. The Acetyl-CoA bounded to the enzyme is shown in light pink, and the product still bound to the catalytic site is shown in purple. The transferred acetyl group in the N-acetyl-D-glucosamine-6-phosphate product in purple is shown in yellow. This proposed 3d structure of the protein shows that the specific parts of the substrates involved in this reaction - the terminal end of the linear portion of Acetyl-CoA and the nitrogen group attached to the glucosamine ring - are in great proximity.
References
- ↑ "Mosquito glucosamine-6-phosphate N-acetyltransferase: cDNA, gene structure and enzyme kinetics". Insect Biochemistry and Molecular Biology 35 (6): 637–46. June 2005. doi:10.1016/j.ibmb.2005.02.005. PMID 15857769.
- ↑ Schwarzer, Michael; Doenst, Torsten (2016). The Scientist's Guide to Cardiac Metabolism. Academic Press. pp. 39–55. ISBN 9780128023945.
- ↑ "Glycolysis and the Hexosamine Biosynthetic Pathway as Novel Targets for Upper and Lower Airway Inflammation". Allergy, Asthma & Immunology Research 10 (1): 6–11. January 2018. doi:10.4168/aair.2018.10.1.6. PMID 29178672.
- ↑ "Chitin synthesis and inhibition: a revisit". Pest Management Science 57 (10): 946–50. October 2001. doi:10.1002/ps.363. PMID 11695188.
- ↑ "Three-dimensional structure of the bacterial cell wall peptidoglycan". Proceedings of the National Academy of Sciences of the United States of America 103 (12): 4404–9. March 2006. doi:10.1073/pnas.0510182103. PMID 16537437. Bibcode: 2006PNAS..103.4404M.
- ↑ 6.0 6.1 "Crystal structure and functional characterization of a glucosamine-6-phosphate N-acetyltransferase from Arabidopsis thaliana". The Biochemical Journal 443 (2): 427–37. April 2012. doi:10.1042/BJ20112071. PMID 22329777.
- ↑ "Fig. 1 Chemical structure of cellulose and chitin" (in en). https://www.researchgate.net/figure/Chemical-structure-of-cellulose-and-chitin_fig1_284921454.
- ↑ "Effect of Acetyl Group on Mechanical Properties of Chitin/Chitosan Nanocrystal: A Molecular Dynamics Study". International Journal of Molecular Sciences 17 (1): 61. January 2016. doi:10.3390/ijms17010061. PMID 26742033.
- ↑ "GlmU (N-acetylglucosamine-1-phosphate uridyltransferase) bound to three magnesium ions and ATP at the active site". Acta Crystallographica Section F 70 (Pt 6): 703–8. June 2014. doi:10.1107/S2053230X14008279. PMID 24915076.
- ↑ "Acceptor substrate binding revealed by crystal structure of human glucosamine-6-phosphate N-acetyltransferase 1". FEBS Letters 582 (20): 2973–8. September 2008. doi:10.1016/j.febslet.2008.07.040. PMID 18675810.
- ↑ 11.0 11.1 "GNPNAT1 - Glucosamine 6-phosphate N-acetyltransferase - Homo sapiens (Human) - GNPNAT1 gene & protein". https://www.uniprot.org/uniprot/Q96EK6.
- ↑ "FlyBase - glucosamine 6-phosphate N-acetyltransferase activity". http://flybase.org/cgi-bin/cvreport.pl?cvterm=GO:0004343&childdepth=2.
- ↑ "Regulation of mitochondrial functions by protein phosphorylation and dephosphorylation". Cell & Bioscience 6: 25. 2016-04-14. doi:10.1186/s13578-016-0089-3. PMID 27087918.
- ↑ "Saccharomyces cerevisiae GNA1, an essential gene encoding a novel acetyltransferase involved in UDP-N-acetylglucosamine synthesis". The Journal of Biological Chemistry 274 (1): 424–9. January 1999. doi:10.1074/jbc.274.1.424. PMID 9867860.
- ↑ "Enzymes of UDP-GlcNAc biosynthesis in yeast". Yeast 23 (1): 1–14. January 2006. doi:10.1002/yea.1337. PMID 16408321.
- ↑ PDB: 2O28; "Crystal Structure of Glucosamine-Phosphate N-Acetyltransferase 1". Worldwide Protein Data Bank. doi:10.2210/pdb2o28/pdb. http://www.rcsb.org/structure/2O28.
- ↑ "The crystal structures of Apo and complexed Saccharomyces cerevisiae GNA1 shed light on the catalytic mechanism of an amino-sugar N-acetyltransferase". The Journal of Biological Chemistry 276 (19): 16328–34. May 2001. doi:10.1074/jbc.M009988200. PMID 11278591.
 |

