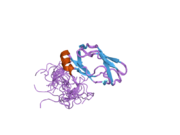Biology:Dihydrolipoyl transacetylase
 Generic protein structure example |
Dihydrolipoyl transacetylase (or dihydrolipoamide acetyltransferase) is an enzyme component of the multienzyme pyruvate dehydrogenase complex. The pyruvate dehydrogenase complex is responsible for the pyruvate decarboxylation step that links glycolysis to the citric acid cycle. This involves the transformation of pyruvate from glycolysis into acetyl-CoA which is then used in the citric acid cycle to carry out cellular respiration.
There are three different enzyme components in the pyruvate dehydrogenase complex. Pyruvate dehydrogenase (EC 1.2.4.1) is responsible for the oxidation of pyruvate, dihydrolipoyl transacetylase (this enzyme; EC 2.3.1.12) transfers the acetyl group to coenzyme A (CoA), and dihydrolipoyl dehydrogenase (EC 1.8.1.4) regenerates the lipoamide. Because dihydrolipoyl transacetylase is the second of the three enzyme components participating in the reaction mechanism for conversion of pyruvate into acetyl CoA, it is sometimes referred to as E2.
In humans, dihydrolipoyl transacetylase enzymatic activity resides in the pyruvate dehydrogenase complex component E2 (PDCE2) that is encoded by the DLAT (dihydrolipoamide S-acetyltransferase) gene.[1]
Nomenclature
The systematic name of this enzyme class is acetyl-CoA:enzyme N6-(dihydrolipoyl)lysine S-acetyltransferase.
Other names in common use include:
- acetyl-CoA:dihydrolipoamide S-acetyltransferase,
- acetyl-CoA:enzyme 6-N-(dihydrolipoyl)lysine S-acetyltransferase.
- dihydrolipoamide S-acetyltransferase,
- dihydrolipoate acetyltransferase,
- dihydrolipoic transacetylase,
- dihydrolipoyl acetyltransferase,
- enzyme-dihydrolipoyllysine:acetyl-CoA S-acetyltransferase,
- lipoate acetyltransferase,
- lipoate transacetylase,
- lipoic acetyltransferase,
- lipoic acid acetyltransferase,
- lipoic transacetylase,
- lipoylacetyltransferase,
- thioltransacetylase A, and
- transacetylase X.
Structure
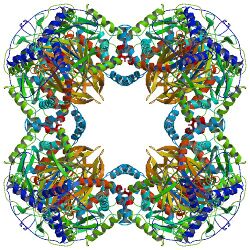
All dihydrolipoyl transacetylases have a unique multidomain structure consisting of (from N to C): 3 lipoyl domains, an interaction domain, and the catalytic domain (see the domain architecture at Pfam). All the domains are connected by disordered, low complexity linker regions.
Depending on the species, multiple subunits of dihydrolipoyl transacetylase enzymes can arrange together into either a cubic or dodecahedral shape. These structure then form the catalytic core of the pyruvate dehydrogenase complex which not only catalyzes the reaction that transfers an acetyl group to CoA, but also performs a crucial structural role in creating the architecture of the overall complex.[3]
Cube
The cubic core structure, found in species such as Azotobacter vinelandii, is made up of 24 subunits total.[4][5] The catalytic domains are assembled into trimers with the active site located at the subunit interface. The topology of this trimer active site is identical to that of chloramphenicol acetyltransferase. Eight of these trimers are then arranged into a hollow truncated cube. The two main substrates, CoA and the lipoamide (Lip(SH)2), are found at two opposite entrances of a 30 Å long channel which runs between the subunits and forms the catalytic center. CoA enters from the inside of the cube, and the lipoamide enters from the outside.[6]
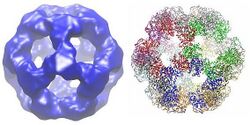
Dodecahedron
In many species, including bacteria such as Geobacillus stearothermophilus and Enterococcus faecalis [3] as well as mammals such as humans[7] and cows,[8] the dodecahedral core structure is made up of 60 subunits total. The subunits are arranged in sets of three, similar to the trimers in the cubic core shape, with each set making up one of the 20 dodecahedral vertices.
Function
| dihydrolipoyllysine-residue acetyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 2.3.1.12 | ||||||||
| CAS number | 9032-29-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Dihydrolipoyl transacetylase participates in the pyruvate decarboxylation reaction that links glycolysis to the citric acid cycle. These metabolic processes are important for cellular respiration—the conversion of biochemical energy from nutrients into adenosine triphosphate (ATP) which can then be used to carry out numerous biological reactions within a cell. The various parts of cellular respiration take place in different parts of the cell. In eukaryotes, glycolysis occurs in the cytoplasm, pyruvate decarboxylation in the mitochondria, the citric acid cycle within the mitochondrial matrix, and oxidative phosphorylation via the electron transport chain on the mitochondrial cristae. Thus pyruvate dehydrogenase complexes (containing the dihydrolipoyl transacetylase enzymes) are found in the mitochondria of eukaryotes (and simply in the cytosol of prokaryotes).
Mechanism
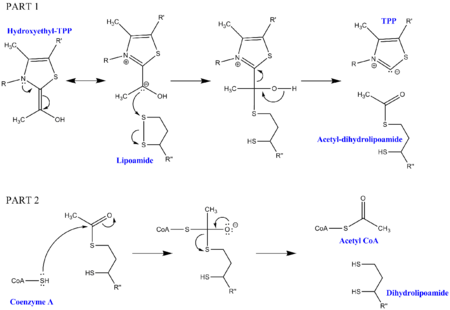
Pyruvate decarboxylation requires a few cofactors in addition to the enzymes that make up the complex. The first is thiamine pyrophosphate (TPP), which is used by pyruvate dehydrogenase to oxidize pyruvate and to form a hydroxyethyl-TPP intermediate. This intermediate is taken up by dihydrolipoyl transacetylase and reacted with a second lipoamide cofactor to generate an acetyl-dihydrolipoyl intermediate, releasing TPP in the process. This second intermediate can then be attacked by the nucleophilic sulfur attached to Coenzyme A, and the dihydrolipoamide is released. This results in the production of acetyl CoA, which is the end goal of pyruvate decarboxylation. The dihydrolipoamide is taken up by dihydrolipoyl dehydrogenase, and with the additional cofactors FAD and NAD+, regenerates the original lipoamide (with NADH as a useful side product).
Interactive pathway map
Clinical significance
Primary biliary cirrhosis
Primary biliary cirrhosis (PBC) is an autoimmune disease characterized by autoantibodies against mitochondrial and nuclear antigens. These are called anti-mitochondrial antibodies (AMA) and anti-nuclear antibodies (ANA), respectively. These antibodies are detectable in the sera of PBC patients and vary greatly with regards to epitope specificity from patient to patient. Of the mitochondrial antigens that can generate autoantibody reactivity in PBC patients, the E2 subunit of the pyruvate dehydrogenase complex, dihydrolipoyl transacetylase, is the most common epitope (other antigens include enzymes of the 2-oxoacid dehydrogenase complexes as well as the other enzymes of the pyruvate dehydrogenase complexes).[9] Recent evidence has suggested that peptides within the catalytic site may present the immunodominant epitopes recognized by the anti-PDC-E2 antibodies in PBC patients.[10] There is also evidence of anti-PDC-E2 antibodies in autoimmune hepatitis (AIH) patients.[11]
Pyruvate dehydrogenase deficiency
Pyruvate dehydrogenase deficiency (PDH) is a genetic disease resulting in lactic acidosis as well as neurological dysfunction in infancy and early childhood. Typically PDH is the result of a mutation in the X-linked gene for the E1 subunit of the pyruvate dehydrogenase complex. However, there have been a few rare cases in which a patient with PDH actually has a mutation in the autosomal gene for the E2 subunit instead. These patients have been reported to have much less severe symptoms, with the most prominent disease manifestation being episodic dystonia, though both hypotonia and ataxia were also present.[12]
References
- ↑ "Chromosome localization and RFLP analysis of PDC-E2: the major autoantigen of primary biliary cirrhosis". Autoimmunity 14 (4): 335–40. 1993. doi:10.3109/08916939309079237. PMID 8102256.
- ↑ "Crystallographic analysis of substrate binding and catalysis in dihydrolipoyl transacetylase (E2p)". Biochemistry 32 (15): 3887–901. Apr 1993. doi:10.1021/bi00066a007. PMID 8471601.
- ↑ 3.0 3.1 3.2 PDB: 1B5S; "Principles of quasi-equivalence and Euclidean geometry govern the assembly of cubic and dodecahedral cores of pyruvate dehydrogenase complexes". Proc. Natl. Acad. Sci. U.S.A. 96 (4): 1240–5. February 1999. doi:10.1073/pnas.96.4.1240. PMID 9990008. Bibcode: 1999PNAS...96.1240I.
- ↑ "The pyruvate dehydrogenase multi-enzyme complex from Gram-negative bacteria". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology 1385 (2): 353–66. Jun 1998. doi:10.1016/S0167-4838(98)00079-X. PMID 9655933.
- ↑ "The quaternary structure of the dihydrolipoyl transacetylase component of the pyruvate dehydrogenase complex from Azotobacter vinelandii. A reconsideration". European Journal of Biochemistry 179 (2): 287–92. Feb 1989. doi:10.1111/j.1432-1033.1989.tb14553.x. PMID 2917567.
- ↑ "Atomic structure of the cubic core of the pyruvate dehydrogenase multienzyme complex". Science 255 (5051): 1544–50. Mar 1992. doi:10.1126/science.1549782. PMID 1549782. Bibcode: 1992Sci...255.1544M.
- ↑ "Subunit and catalytic component stoichiometries of an in vitro reconstituted human pyruvate dehydrogenase complex". The Journal of Biological Chemistry 284 (19): 13086–98. May 2009. doi:10.1074/jbc.M806563200. PMID 19240034.
- ↑ "The remarkable structural and functional organization of the eukaryotic pyruvate dehydrogenase complexes". Proceedings of the National Academy of Sciences of the United States of America 98 (26): 14802–7. Dec 2001. doi:10.1073/pnas.011597698. PMID 11752427. Bibcode: 2001PNAS...9814802Z.
- ↑ "The peculiar autoimmunity of primary biliary cirrhosis". Immunological Reviews 174: 226–37. Apr 2000. doi:10.1034/j.1600-0528.2002.017410.x. PMID 10807519. http://www3.interscience.wiley.com/resolve/openurl?genre=article&sid=nlm:pubmed&issn=0105-2896&date=2000&volume=174&spage=226.
- ↑ "Catalytic domain of PDC-E2 contains epitopes recognized by antimitochondrial antibodies in primary biliary cirrhosis". World Journal of Gastroenterology 16 (8): 973–81. Feb 2010. doi:10.3748/wjg.v16.i8.973. PMID 20180236.
- ↑ "Long-term follow-up of antimitochondrial antibody-positive autoimmune hepatitis". Hepatology 48 (2): 550–6. Aug 2008. doi:10.1002/hep.22380. PMID 18666262.
- ↑ "Clinical and genetic spectrum of pyruvate dehydrogenase deficiency: dihydrolipoamide acetyltransferase (E2) deficiency". Annals of Neurology 58 (2): 234–41. Aug 2005. doi:10.1002/ana.20550. PMID 16049940.
Further reading
- "Crystallographic analysis of substrate binding and catalysis in dihydrolipoyl transacetylase (E2p)". Biochemistry 32 (15): 3887–901. Apr 1993. doi:10.1021/bi00066a007. PMID 8471601.
- "Enzymatic thioltransacetylation". The Journal of Biological Chemistry 211 (2): 621–9. Dec 1954. doi:10.1016/S0021-9258(18)71152-6. PMID 13221570.
- "Biosynthesis and structure of lipoic acid derivatives". J. Am. Chem. Soc. 78 (8): 1763–1766. 1956. doi:10.1021/ja01589a079.
- "Swinging arms and swinging domains in multifunctional enzymes: catalytic machines for multistep reactions". Annual Review of Biochemistry 69: 961–1004. 2000. doi:10.1146/annurev.biochem.69.1.961. PMID 10966480.
- "Three-dimensional structure of the major autoantigen in primary biliary cirrhosis". Gastroenterology 115 (1): 139–46. Jul 1998. doi:10.1016/S0016-5085(98)70375-0. PMID 9649469.
- "Comprehensive mapping of HLA-A0201-restricted CD8 T-cell epitopes on PDC-E2 in primary biliary cirrhosis". Hepatology 36 (5): 1125–34. Nov 2002. doi:10.1053/jhep.2002.36161. PMID 12395322.
- "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene 200 (1–2): 149–56. Oct 1997. doi:10.1016/S0378-1119(97)00411-3. PMID 9373149.
- "Binding of pyruvate dehydrogenase to the core of the human pyruvate dehydrogenase complex". FEBS Letters 582 (3): 468–72. Feb 2008. doi:10.1016/j.febslet.2007.12.041. PMID 18206651.
- "Clinical and genetic spectrum of pyruvate dehydrogenase deficiency: dihydrolipoamide acetyltransferase (E2) deficiency". Annals of Neurology 58 (2): 234–41. Aug 2005. doi:10.1002/ana.20550. PMID 16049940.
- "Disease-specific cross-reactivity between mimicking peptides of heat shock protein of Mycobacterium gordonae and dominant epitope of E2 subunit of pyruvate dehydrogenase is common in Spanish but not British patients with primary biliary cirrhosis". Journal of Autoimmunity 22 (4): 353–62. Jun 2004. doi:10.1016/j.jaut.2004.03.002. PMID 15120760.
- "Apotopes and the biliary specificity of primary biliary cirrhosis". Hepatology 49 (3): 871–9. Mar 2009. doi:10.1002/hep.22736. PMID 19185000. PMC 2665925. https://air.unimi.it/bitstream/2434/55031/2/22736_ftp.pdf.
- "Differential epitope mapping of antibodies to PDC-E2 in patients with hematologic malignancies after allogeneic hematopoietic stem cell transplantation and primary biliary cirrhosis". Blood 109 (5): 2001–7. Mar 2007. doi:10.1182/blood-2006-06-030304. PMID 17068145.
- "Facilitated interaction between the pyruvate dehydrogenase kinase isoform 2 and the dihydrolipoyl acetyltransferase". The Journal of Biological Chemistry 278 (36): 33681–93. Sep 2003. doi:10.1074/jbc.M212733200. PMID 12816949.
- "Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappaB signalling". Gut 58 (8): 1078–83. Aug 2009. doi:10.1136/gut.2008.169052. PMID 19240061. https://research.vumc.nl/en/publications/9f0b4115-baf6-4bc6-ab6b-4fd7f23f0f0a.
- "Organization of the cores of the mammalian pyruvate dehydrogenase complex formed by E2 and E2 plus the E3-binding protein and their capacities to bind the E1 and E3 components". The Journal of Biological Chemistry 279 (8): 6921–33. Feb 2004. doi:10.1074/jbc.M308172200. PMID 14638692.
- "The glucose-lactic acid cycle and gluconeogenesis". Current Topics in Cellular Regulation 18: 377–87. 1981. doi:10.1016/B978-0-12-152818-8.50028-1. ISBN 9780121528188. PMID 7273846.
- "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene 138 (1–2): 171–4. Jan 1994. doi:10.1016/0378-1119(94)90802-8. PMID 8125298.
- "Interaction between the individual isoenzymes of pyruvate dehydrogenase kinase and the inner lipoyl-bearing domain of transacetylase component of pyruvate dehydrogenase complex". The Biochemical Journal 366 (Pt 1): 129–36. Aug 2002. doi:10.1042/BJ20020301. PMID 11978179.
External links
- PDB: 1EAA, PDB: 1dpb
- Dihydrolipoyl+transacetylase at the US National Library of Medicine Medical Subject Headings (MeSH)
- Overview of all the structural information available in the PDB for UniProt: P10515 (Dihydrolipoyl transacetylase) at the PDBe-KB.
 |