Biology:Glycoside hydrolase family 56
| Hyaluronidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
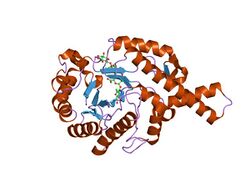 crystal structure of bee venom hyaluronidase in complex with hyaluronic acid tetramer | |||||||||
| Identifiers | |||||||||
| Symbol | Glyco_hydro_56 | ||||||||
| Pfam | PF01630 | ||||||||
| Pfam clan | CL0058 | ||||||||
| InterPro | IPR018155 | ||||||||
| SCOP2 | 1fcv / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 117 | ||||||||
| OPM protein | 1fcq | ||||||||
| CAZy | GH56 | ||||||||
| Membranome | 1078 | ||||||||
| |||||||||
In molecular biology, glycoside hydrolase family 56 is a family of glycoside hydrolases.
Glycoside hydrolases EC 3.2.1. are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycoside hydrolases, based on sequence similarity, has led to the definition of >100 different families.[1][2][3] This classification is available on the CAZy web site,[4][5] and also discussed at CAZypedia, an online encyclopedia of carbohydrate active enzymes.[6][7]
Glycoside hydrolase family 56 CAZY GH_56 includes enzymes with hyaluronidase EC 3.2.1.35 activity. The venom of Apis mellifera (Honeybee) contains several biologically-active peptides and two enzymes, one of which is a hyaluronidase.[8] The amino acid sequence of bee venom hyaluronidase contains 349 amino acids, and includes four cysteines and a number of potential glycosylation sites.[8] The sequence shows a high degree of similarity to PH-20, a membrane protein of mammalian sperm involved in sperm-egg adhesion, supporting the view that hyaluronidases play a role in fertilisation.[8]
PH-20 is required for sperm adhesion to the egg zona pellucida; it is located on both the sperm plasma membrane and acrosomal membrane.[9] The amino acid sequence of the mature protein contains 468 amino acids, and includes six potential N-linked glycosylation sites and twelve cysteines, eight of which are tightly clustered near the C-terminus.[9]
References
- ↑ "Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases". Proceedings of the National Academy of Sciences of the United States of America 92 (15): 7090–4. July 1995. doi:10.1073/pnas.92.15.7090. PMID 7624375. Bibcode: 1995PNAS...92.7090H.
- ↑ "Structures and mechanisms of glycosyl hydrolases". Structure 3 (9): 853–9. September 1995. doi:10.1016/S0969-2126(01)00220-9. PMID 8535779.
- ↑ "Updating the sequence-based classification of glycosyl hydrolases". The Biochemical Journal 316 (Pt 2): 695–6. June 1996. doi:10.1042/bj3160695. PMID 8687420.
- ↑ "Home" (in en). http://www.cazy.org/.
- ↑ "The carbohydrate-active enzymes database (CAZy) in 2013". Nucleic Acids Research 42 (Database issue): D490-5. January 2014. doi:10.1093/nar/gkt1178. PMID 24270786.
- ↑ "Glycoside Hydrolase Family 56" (in en). http://www.cazypedia.org/index.php/Glycoside_Hydrolase_Family_56.
- ↑ CAZypedia Consortium (December 2018). "Ten years of CAZypedia: a living encyclopedia of carbohydrate-active enzymes". Glycobiology 28 (1): 3–8. doi:10.1093/glycob/cwx089. PMID 29040563. https://hal.archives-ouvertes.fr/hal-01886461/file/Hehemann_2018_01.pdf.
- ↑ 8.0 8.1 8.2 "Bee venom hyaluronidase is homologous to a membrane protein of mammalian sperm". Proc. Natl. Acad. Sci. U.S.A. 90 (8): 3569–3573. 1993. doi:10.1073/pnas.90.8.3569. PMID 7682712. Bibcode: 1993PNAS...90.3569G.
- ↑ 9.0 9.1 "cDNA cloning reveals the molecular structure of a sperm surface protein, PH-20, involved in sperm-egg adhesion and the wide distribution of its gene among mammals". J. Cell Biol. 111 (6): 2939–2949. 1990. doi:10.1083/jcb.111.6.2939. PMID 2269661.
 |

