Biology:Cellulose synthase (UDP-forming)
| Cellulose synthase (CesA/BcsA) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Cellulose_synth | ||||||||
| Pfam | PF03552 | ||||||||
| InterPro | IPR005150 | ||||||||
| TCDB | 4.D.3 | ||||||||
| CAZy | GT2 | ||||||||
| |||||||||
| 4p02 chain A; CAZy and TCDB also includes other proteins | |||||||||
| Bacterial cellulose synthase di-GMP-binding regulatory subunit | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | BcsB | ||||||||
| Pfam | PF03170 | ||||||||
| InterPro | IPR018513 | ||||||||
| CATH | 4p02 | ||||||||
| OPM superfamily | 302 | ||||||||
| OPM protein | 4p02 | ||||||||
| Membranome | 539 | ||||||||
| |||||||||
| 4p02 chain B | |||||||||
The UDP-forming form of cellulose synthase (EC 2.4.1.12) is the main enzyme that produces cellulose. Systematically, it is known as UDP-glucose:(1→4)-β-D-glucan 4-β-D-glucosyltransferase in enzymology. It catalyzes the chemical reaction:
- UDP-glucose + [(1→4)-β-D-glucosyl]n = UDP + [(1→4)-β-D-glucosyl]n+1
A similar enzyme utilizes GDP-glucose, cellulose synthase (GDP-forming) (EC 2.4.1.29).
This family of enzymes is found in bacteria and plants alike. Plant members are usually known as CesA (cellulose synthase) or the tentative CslA (cellulose synthase-like), while bacterial members may additionally be known as BcsA (bacterial cellulose synthase) or CelA (simply "cellulose").[1] Plants acquired CesA from the endosymbiosis event that produced the chloroplast.[2] This family belongs to glucosyltransferase family 2 (GT2).[1] Glycosyltransferases are involved in the biosynthesis and hydrolysis of the bulk of earth's biomass.[3] There are known to be about seven subfamilies in the plant CesA superfamily,[4] or ten in the combined plant-algal superfamily.[5] Urochordates are the only group of animals possessing this enzyme, having acquired them by horizontal gene transfer more than 530 million years ago.[6]
Cellulose
Cellulose is an aggregation of unbranched polymer chains made of β-(1→4)-linked glucose residues that makes up a large portion of primary and secondary cell walls.[7][8][9][10] Although important for plants, it is also synthesized by most algae, some bacteria, and some animals.[6][5][11][12] Worldwide, 2 × 1011 tons of cellulose microfibrils are produced,[13] which serves as a critical source of renewable biofuels and other biological-based products, such as lumber, fuel, fodder, paper and cotton.[8][14]
Purpose of cellulose
Cellulose microfibrils are made on the surface of cell membranes to reinforce cells walls, which has been researched extensively by plant biochemists and cell biologist because 1) they regulate cellular morphogenesis and 2) they serve alongside many other constituents (i.e. lignin, hemicellulose, pectin) in the cell wall as a strong structural support and cell shape.[14] Without these support structures, cell growth would cause a cell to swell and spread in all directions, thus losing its shape viability [15]
Structure
This section is missing information about sequence motifs, mentioned in most structural articles. (August 2019) |
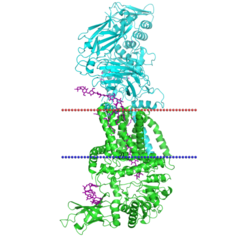
Several structures of the bacterial cellulose synthase BcsAB has been resolved. The bacterial enzyme consists of two different subunits, the catalytic BcsA on the cytoplasmic side, and the regulatory BcsB on the periplasmic side. They are coupled by a series of transmembrane helices, termed by the CATH database as 4p02A01 and 4p02B05. (Divisions for other models, such as 4hg6, follow similarly.) The enzyme is stimulated by cyclic di-GMP. In vivo but not in vitro, a third subunit called BcsC made up of a 18-strand beta barrel is required. Some bacteria contain extra non-essential periplasmic subunits.[16]
BcsA follows a layout of cytoplasmic domains sandwiched between the N- and C-terminal transmembrane domain. It has a typical family 2 GT domain (4p02A02) with a GT-A fold structure. At the C-terminal end is a PilZ domain conserved in bacteria,[16] which forms part of the cyclic di-GMP binding surface together with BcsB and the beta-barrel (4p02A03) domain.[17] Besides the C-terminal TM domain, BcsB is made up of two repeats, each consisting of a carbohydrate-binding module 27 (CATH 2.60.120.260) and an alpha-beta sandwith (CATH 3.30.379.20).[16]
BcsA and BcsB together form a channel through with the synthesized cellulose exits the cell, and mutations to residues lining the channel are known to reduce the activity of this enzyme.[16] A gating loop in BcsA closes over the channel; it opens when cyclic di-GMP is bound to the enzyme.[17]
Plants
In plants, cellulose is synthesized by large cellulose synthase complexes (CSCs), which consist of synthase protein isoforms (CesA) that are arranged into a unique hexagonal structure known as a “particle rosette” 50 nm wide and 30–35 nm tall.[5][18][19] There are more than 20 of these full-length integral membrane proteins, each of which is around 1000 amino acids long.[8][9] These rosettes, formerly known as granules, were first discovered in 1972 by electron microscopy in green algae species Cladophora and Chaetomorpha[20] (Robinson et al. 1972). Solution x-ray scattering have shown that CesAs are at the surface of a plant cell and are elongated monomers with a two catalytic domains that fuse together into dimers. The center of the dimers is the main point of catalytic activity,[5] and the lobes are presumed to contain the plant specific PC-R and CS-R.[8] Since cellulose is made in all cell walls, CesA proteins are present in all tissues and cell types of plants. Nonetheless, there are different types of CesA, some tissue types may have varying concentrations of one over another. For example, the AtCesA1 (RSW1) protein is involved in the biosynthesis of primary cell walls throughout the whole plant while the AtCesA7 (IRX3) protein is only expressed in the stem for secondary cell wall production.[9]
Compared to the bacterial enzyme, plant versions of the synthase are much harder to crystallize, and as of August 2019 no experimental atomic structures of the plant cellulose synthase catalytic domain is known. However, at least two high-confidence structures have been predicted for these enzymes.[8][11] The broader of the two structures (Sethaphong 2013), which includes the entire middle cytoplasmic domain (again sandwiched between TM helices), gives a useful view of the enzyme: two plant-specific insertions called the PC-R (plant-conserved region, similar in all plants) on the N-terminal end and CS-R (class-specific region, determines the subclass number after CesA) on the C-terminal end punctuate the usual GT catalytic core, probably providing the unique rosette-forming function of plant CesA.[11] (Some CesA proteins possess an additional insertion.)[21] The structure seems to explain the effects of many known mutations. The positioning of the two insertions, however, do not match the scattering result from Olek 2014.[11] An 2016 experimental model of the PC-R (5JNP) domain helps to fill in this gap, as it greatly improves the fit against Olek's previous result.[22] It also matches the Sethaphong 2015 prediction of an antiparallel coiled-coil well. The two groups continue to further their understandings of the CesA structure, with Olek et al focusing on experimental structures and Sethaphong et al focusing on plant studies and building better computer models.[23]
Other differences from the bacterial BcsA includes a different TM helice count (BcsA has 4 helices on each end; CesA has two on the N-terminal and 6 on the C-terminal), and the presence of a zinc finger (1WEO) at the N-terminus.[8]
Activity
Cellulose biosynthesis is the process during which separate homogeneous β-(1→4)-glucan chains, ranging from 2,000 to 25,000 glucose residues in length, are synthesized and then immediately hydrogen bond with one another to form rigid crystalline arrays, or microfibrils.[8] Microfibrils in the primary cell wall are approximately 36 chains long while those of the secondary cell wall are much larger, containing up to 1200 β-(1→4)-glucan chains.[14][9] Uridine diphosphate-glucose (UDP), which is produced by the enzyme sucrose synthase (SuSy) that produces and transports UDP-glucose to the plasma membrane is the substrate used by cellulose synthase to produce the glucan chains.[8][24] The rate at which glucose residues are synthesized per one glucan chain ranges from 300 to 1000 glucose residues per minute, the higher rate being more prevalent in secondary wall particles, such as in the xylem.[25][26]
Supporting structures
Microfibril synthesis is guided by cortical microtubules, which lie beneath the plasma membrane of elongating cells, in that they form a platform on which the CSCs can convert glucose molecules into the crystalline chains. The microtubule–microfibril alignment hypothesis proposes that cortical microtubules, which lie beneath the plasma membrane of elongating cells, provide tracks for CSCs that convert glucose molecules into crystalline cellulose microfibrils.[27] The direct hypothesis postulates some types of direct linkage between CESA complexes and microtubules.[24] Additionally, the KORRIGAN (KOR1) protein is thought to be a critical component of cellulose synthesis in that it acts as a cellulase at the plasma membrane-cell wall interface. KOR1 interacts with a two specific CesA proteins, possibly by proof-reading and relieving stress created by glucan chain synthesis, by hydrolyzing disordered amorphous cellulose.[28]
Environmental influences
Cellulose synthesis activity is affected by many environmental stimuli, such as hormones, light, mechanical stimuli, nutrition, and interactions with the cytoskeleton. Interactions with these factors may influence cellulose deposition in that it affects the amount of substrate produced and the concentration and/or activity of CSCs in the plasma membrane.[8][5]
References
- ↑ 1.0 1.1 "BcsA and BcsB form the catalytically active core of bacterial cellulose synthase sufficient for in vitro cellulose synthesis". Proceedings of the National Academy of Sciences of the United States of America 110 (44): 17856–61. October 2013. doi:10.1073/pnas.1314063110. PMID 24127606. Bibcode: 2013PNAS..11017856O.
- ↑ "Evolution and diversity of plant cell walls: from algae to flowering plants". Annual Review of Plant Biology 62: 567–90. 2011. doi:10.1146/annurev-arplant-042110-103809. PMID 21351878.
- ↑ "A classification of nucleotide-diphospho-sugar glycosyltransferases based on amino acid sequence similarities". The Biochemical Journal 329 (Pt 3) (3): 719. February 1998. doi:10.1042/bj3290719. PMID 9445404.
- ↑ "The cellulose synthase superfamily". Plant Physiology 124 (2): 495–8. October 2000. doi:10.1104/pp.124.2.495. PMID 11027699.
- ↑ 5.0 5.1 5.2 5.3 5.4 "The cellulose synthase superfamily in fully sequenced plants and algae". BMC Plant Biology 9: 99. July 2009. doi:10.1186/1471-2229-9-99. PMID 19646250.
- ↑ 6.0 6.1 "The evolutionary origin of animal cellulose synthase". Development Genes and Evolution 214 (2): 81–8. February 2004. doi:10.1007/s00427-003-0379-8. PMID 14740209.
- ↑ "A classification of nucleotide-diphospho-sugar glycosyltransferases based on amino acid sequence similarities". The Biochemical Journal 326 ( Pt 3) (3): 929–39. September 1997. doi:10.1042/bj3260929u. PMID 9334165.
- ↑ 8.0 8.1 8.2 8.3 8.4 8.5 8.6 8.7 8.8 "The structure of the catalytic domain of a plant cellulose synthase and its assembly into dimers". The Plant Cell 26 (7): 2996–3009. July 2014. doi:10.1105/tpc.114.126862. PMID 25012190.
- ↑ 9.0 9.1 9.2 9.3 "Higher plant cellulose synthases". Genome Biology 1 (4): REVIEWS3001. 2000. doi:10.1186/gb-2000-1-4-reviews3001. PMID 11178255.
- ↑ "Cellulose synthase complexes: composition and regulation". Frontiers in Plant Science 3: 75. 2012. doi:10.3389/fpls.2012.00075. PMID 22639663.
- ↑ 11.0 11.1 11.2 11.3 "Tertiary model of a plant cellulose synthase". Proceedings of the National Academy of Sciences of the United States of America 110 (18): 7512–7. April 2013. doi:10.1073/pnas.1301027110. PMID 23592721. Bibcode: 2013PNAS..110.7512S.
- ↑ "Functional analysis of complexes with mixed primary and secondary cellulose synthases". Plant Signaling & Behavior 8 (3): e23179. March 2013. doi:10.4161/psb.23179. PMID 23299322.
- ↑ Measurement of calorific values. Primary productivity of the biosphere. Ecological Studies. 14. New York: Springer. 1975. pp. 119–129. doi:10.1007/978-3-642-80913-2. ISBN 978-3-642-80915-6.
- ↑ 14.0 14.1 14.2 "Cloning in silico". Current Biology 7 (2): R108-11. February 1997. doi:10.1016/S0960-9822(06)00050-9. PMID 9081659.
- ↑ "Effects of colchicine on cell shape and on microfibril arrangement in the cell wall of Closterium acerosum". Planta 140 (1): 15–8. January 1978. doi:10.1007/BF00389374. PMID 24414355.
- ↑ 16.0 16.1 16.2 16.3 "Crystallographic snapshot of cellulose synthesis and membrane translocation". Nature 493 (7431): 181–6. January 2013. doi:10.1038/nature11744. PMID 23222542. Bibcode: 2013Natur.493..181M.
- ↑ 17.0 17.1 "Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP". Nature Structural & Molecular Biology 21 (5): 489–96. May 2014. doi:10.1038/nsmb.2803. PMID 24704788.
- ↑ "Visualization of particle complexes in the plasma membrane of Micrasterias denticulata associated with the formation of cellulose fibrils in primary and secondary cell walls". The Journal of Cell Biology 84 (2): 327–39. February 1980. doi:10.1083/jcb.84.2.327. PMID 7189756.
- ↑ "The cytoplasmic domain of the cellulose-synthesizing complex in vascular plants". Protoplasma 233 (1–2): 115–27. 2008. doi:10.1007/s00709-008-0302-2. PMID 18709477.
- ↑ "Fine structure of swarmers of Cladophora and Chaetomorpha : III. Wall synthesis and development". Planta 107 (2): 131–44. June 1972. doi:10.1007/BF00387719. PMID 24477398.
- ↑ "Understanding Plant Cellulose Synthases through a Comprehensive Investigation of the Cellulose Synthase Family Sequences". Frontiers in Plant Science 2: 5. 2011. doi:10.3389/fpls.2011.00005. PMID 22629257.
- ↑ "Rice Cellulose SynthaseA8 Plant-Conserved Region Is a Coiled-Coil at the Catalytic Core Entrance". Plant Physiology 173 (1): 482–494. January 2017. doi:10.1104/pp.16.00739. PMID 27879387.
- ↑ "Prediction of the structures of the plant-specific regions of vascular plant cellulose synthases and correlated functional analysis". Cellulose 23 (1): 145–161. 24 October 2015. doi:10.1007/s10570-015-0789-6.
- ↑ 24.0 24.1 "A unified hypothesis for the role of membrane bound enzyme complexes and microtubules in plant cell wall synthesis". Journal of Theoretical Biology 48 (2): 445–9. December 1974. doi:10.1016/S0022-5193(74)80011-1. PMID 4459594. https://www.researchgate.net/publication/18705609.
- ↑ "Visualization of cellulose synthase demonstrates functional association with microtubules". Science 312 (5779): 1491–5. June 2006. doi:10.1126/science.1126551. PMID 16627697. Bibcode: 2006Sci...312.1491P.
- ↑ "The roles of the cytoskeleton during cellulose deposition at the secondary cell wall". The Plant Journal 54 (5): 794–805. June 2008. doi:10.1111/j.1365-313X.2008.03444.x. PMID 18266917.
- ↑ "Mechanism for Plant Cellular Morphogenesis". Science 138 (3548): 1404–5. December 1962. doi:10.1126/science.138.3548.1404. PMID 17753861. Bibcode: 1962Sci...138.1404G.
- ↑ "KORRIGAN1 interacts specifically with integral components of the cellulose synthase machinery". PLOS ONE 9 (11): e112387. 2014. doi:10.1371/journal.pone.0112387. PMID 25383767. Bibcode: 2014PLoSO...9k2387M.
Further reading
- "The synthesis of cellulose in cell-free extracts of Acetobacter xylinum". The Journal of Biological Chemistry 232 (2): 627–36. June 1958. doi:10.1016/S0021-9258(19)77383-9. PMID 13549448.
 |