Biology:Extrachromosomal DNA
Extrachromosomal DNA (abbreviated ecDNA) is any DNA that is found off the chromosomes, either inside or outside the nucleus of a cell. Most DNA in an individual genome is found in chromosomes contained in the nucleus. Multiple forms of extrachromosomal DNA exist, and, while some of these serve important biological functions,[1] they can also play a role in diseases such as cancer.[2][3][4]
In prokaryotes, nonviral extrachromosomal DNA is primarily found in plasmids, whereas, in eukaryotes extrachromosomal DNA is primarily found in organelles.[1] Mitochondrial DNA is a main source of this extrachromosomal DNA in eukaryotes.[5] The fact that this organelle contains its own DNA supports the hypothesis that mitochondria originated as bacterial cells engulfed by ancestral eukaryotic cells.[6] Extrachromosomal DNA is often used in research into replication because it is easy to identify and isolate.[1]
Although extrachromosomal circular DNA (eccDNA) is found in normal eukaryotic cells, extrachromosomal DNA (ecDNA) is a distinct entity that has been identified in the nuclei of cancer cells and has been shown to carry many copies of driver oncogenes.[7][8][3] ecDNA is considered to be a primary mechanism of gene amplification, resulting in many copies of driver oncogenes and very aggressive cancers.
Extrachromosomal DNA in the cytoplasm has been found to be structurally different from nuclear DNA. Cytoplasmic DNA is less methylated than DNA found within the nucleus. It was also confirmed that the sequences of cytoplasmic DNA were different from nuclear DNA in the same organism, showing that cytoplasmic DNAs are not simply fragments of nuclear DNA.[9] In cancer cells, ecDNA have been shown to be primarily isolated to the nucleus (reviewed in [2]).
In addition to DNA found outside the nucleus in cells, infection by viral genomes also provides an example of extrachromosomal DNA.
Prokaryotic

Although prokaryotic organisms do not possess a membrane-bound nucleus like eukaryotes, they do contain a nucleoid region in which the main chromosome is found. Extrachromosomal DNA exists in prokaryotes outside the nucleoid region as circular or linear plasmids. Bacterial plasmids are typically short sequences, consisting of 1 to a few hundred kilobase (kb) segments, and contain an origin of replication which allows the plasmid to replicate independently of the bacterial chromosome.[10] The total number of a particular plasmid within a cell is referred to as the copy number and can range from as few as two copies per cell to as many as several hundred copies per cell.[11] Circular bacterial plasmids are classified according to the special functions that the genes encoded on the plasmid provide. Fertility plasmids, or f plasmids, allow for conjugation to occur whereas resistance plasmids, or r plasmids, contain genes that convey resistance to a variety of different antibiotics such as ampicillin and tetracycline. Virulence plasmids contain the genetic elements necessary for bacteria to become pathogenic. Degradative plasmids that contain genes that allow bacteria to degrade a variety of substances such as aromatic compounds and xenobiotics.[12] Bacterial plasmids can also function in pigment production, nitrogen fixation and the resistance to heavy metals.[13]
Naturally occurring circular plasmids can be modified to contain multiple resistance genes and several unique restriction sites, making them valuable tools as cloning vectors in biotechnology.[10] Circular bacterial plasmids are also the basis for the production of DNA vaccines. Plasmid DNA vaccines are genetically engineered to contain a gene which encodes for an antigen or a protein produced by a pathogenic virus, bacterium or other parasites.[14] Once delivered into the host, the products of the plasmid genes will then stimulate both the innate immune response and the adaptive immune response of the host. The plasmids are often coated with some type of adjuvant prior to delivery to enhance the immune response from the host.[15]
Linear bacterial plasmids have been identified in several species of spirochete bacteria, including members of the genus Borrelia (to which the pathogen responsible for Lyme disease belongs), several species of the gram positive soil bacteria of the genus Streptomyces, and in the gram negative species Thiobacillus versutus, a bacterium that oxidizes sulfur. Linear plasmids of prokaryotes are found either containing a hairpin loop or a covalently bonded protein attached to the telomeric ends of the DNA molecule. The adenine-thymine rich hairpin loops of the Borrelia bacteria range in size from 5 kilobase pairs (kb) to over 200 kb[16] and contain the genes responsible for producing a group of major surface proteins, or antigens, on the bacteria that allow it to evade the immune response of its infected host.[17] The linear plasmids which contain a protein that has been covalently attached to the 5' end of the DNA strands are known as invertrons and can range in size from 9 kb to over 600 kb consisting of inverted terminal repeats.[16] The linear plasmids with a covalently attached protein may assist with bacterial conjugation and integration of the plasmids into the genome. These types of linear plasmids represent the largest class of extrachromosomal DNA as they are not only present in certain bacterial cells, but all linear extrachromosomal DNA molecules found in eukaryotic cells also take on this invertron structure with a protein attached to the 5' end.[16][17]
The long, linear "borgs" that co-occur with a species of archaeon – which may host them and shares many of their genes – could be an unknown form of extrachromosomal DNA structures.[18][19][20]
Eukaryotic
Mitochondrial
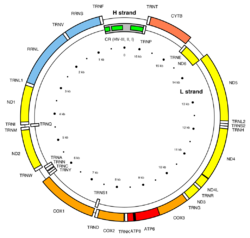
Mitochondria present in eukaryotic cells contain multiple copies of mitochondrial DNA (mtDNA) in the mitochondrial matrix.[21] In multicellular animals, including humans, the circular mtDNA chromosome contains 13 genes that encode proteins that are part of the electron transport chain and 24 genes for mitochondrial RNAs; these genes are broken down into 2 rRNA genes and 22 tRNA genes.[22] The size of an animal mtDNA plasmid is roughly 16.6 kb and, although it contains genes for tRNA and mRNA synthesis, proteins coded for by nuclear genes are still required for the mtDNA to replicate or for mitochondrial proteins to be translated.[23] There is only one region of the mitochondrial chromosome that does not contain a coding sequence, the 1 kb region known as the D-loop to which nuclear regulatory proteins bind.[22] The number of mtDNA molecules per mitochondrion varies from species to species, as well as between cells with different energy demands. For example, muscle and liver cells contain more copies of mtDNA per mitochondrion than blood and skin cells do.[23] Due to the proximity of the electron transport chain within the mitochondrial inner membrane and the production of reactive oxygen species (ROS), and due to the fact that the mtDNA molecule is not bound by or protected by histones, the mtDNA is more susceptible to DNA damage than nuclear DNA.[24] In cases where mtDNA damage does occur, the DNA can either be repaired via base excision repair pathways, or the damaged mtDNA molecule is destroyed (without causing damage to the mitochondrion since there are multiple copies of mtDNA per mitochondrion).[25]
The standard genetic code by which nuclear genes are translated is universal, meaning that each 3-base sequence of DNA codes for the same amino acid regardless of what species from which the DNA comes. However, this code is quite universal and is slightly different in mitochondrial DNA of fungi, animals, protists and plants.[21] While most of the 3-base sequences (codons) in the mtDNA of these organisms do code for the same amino acids as those of the nuclear genetic code, a few are different.
| Genetic code | Translation table | DNA codon involved | RNA codon involved | Translation with this code | Comparison with the universal code |
|---|---|---|---|---|---|
| Vertebrate mitochondrial | 2 | AGA
|
AGA
|
Ter (*)
|
Arg (R)
|
AGG
|
AGG
|
Ter (*)
|
Arg (R)
| ||
ATA
|
AUA
|
Met (M)
|
Ile (I)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
| Yeast mitochondrial | 3 | ATA
|
AUA
|
Met (M)
|
Ile (I)
|
CTT
|
CUU
|
Thr (T)
|
Leu (L)
| ||
CTC
|
CUC
|
Thr (T)
|
Leu (L)
| ||
CTA
|
CUA
|
Thr (T)
|
Leu (L)
| ||
CTG
|
CUG
|
Thr (T)
|
Leu (L)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
CGA
|
CGA
|
absent
|
Arg (R)
| ||
CGC
|
CGC
|
absent
|
Arg (R)
| ||
| Mold, protozoan, and coelenterate mitochondrial | 4 and 7 | TGA
|
UGA
|
Trp (W)
|
Ter (*)
|
| Invertebrate mitochondrial | 5 | AGA
|
AGA
|
Ser (S)
|
Arg (R)
|
AGG
|
AGG
|
Ser (S)
|
Arg (R)
| ||
ATA
|
AUA
|
Met (M)
|
Ile (I)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
| Echinoderm and flatworm mitochondrial | 9 | AAA
|
AAA
|
Asn (N)
|
Lys (K)
|
AGA
|
AGA
|
Ser (S)
|
Arg (R)
| ||
AGG
|
AGG
|
Ser (S)
|
Arg (R)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
| Ascidian mitochondrial | 13 | AGA
|
AGA
|
Gly (G)
|
Arg (R)
|
AGG
|
AGG
|
Gly (G)
|
Arg (R)
| ||
ATA
|
AUA
|
Met (M)
|
Ile (I)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
| Alternative flatworm mitochondrial | 14 | AAA
|
AAA
|
Asn (N)
|
Lys (K)
|
AGA
|
AGA
|
Ser (S)
|
Arg (R)
| ||
AGG
|
AGG
|
Ser (S)
|
Arg (R)
| ||
TAA
|
UAA
|
Tyr (Y)
|
Ter (*)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
| ||
| Chlorophycean mitochondrial | 16 | TAG
|
UAG
|
Leu (L)
|
Ter (*)
|
| Trematode mitochondrial | 21 | TGA
|
UGA
|
Trp (W)
|
Ter (*)
|
ATA
|
AUA
|
Met (M)
|
Ile (I)
| ||
AGA
|
AGA
|
Ser (S)
|
Arg (R)
| ||
AGG
|
AGG
|
Ser (S)
|
Arg (R)
| ||
AAA
|
AAA
|
Asn (N)
|
Lys (K)
| ||
| Scenedesmus obliquus mitochondrial | 22 | TCA
|
UCA
|
Ter (*)
|
Ser (S)
|
TAG
|
UAG
|
Leu (L)
|
Ter (*)
| ||
| Thraustochytrium mitochondrial | 23 | TTA
|
UUA
|
Ter (*)
|
Leu (L)
|
| Pterobranchia mitochondrial | 24 | AGA
|
AGA
|
Ser (S)
|
Arg (R)
|
AGG
|
AGG
|
Lys (K)
|
Arg (R)
| ||
TGA
|
UGA
|
Trp (W)
|
Ter (*)
|
| Amino acids biochemical properties | nonpolar | polar | basic | acidic | Termination: stop codon |
The coding differences are thought to be a result of chemical modifications in the transfer RNAs that interact with the messenger RNAs produced as a result of transcribing the mtDNA sequences.[26]
Chloroplast
Eukaryotic chloroplasts, as well as the other plant plastids, also contain extrachromosomal DNA molecules. Most chloroplasts house all of their genetic material in a single ringed chromosome, however in some species there is evidence of multiple smaller ringed plasmids.[27][28][29] A recent theory that questions the current standard model of ring shaped chloroplast DNA (cpDNA), suggests that cpDNA may more commonly take a linear shape.[30] A single molecule of cpDNA can contain anywhere from 100 to 200 genes[31] and varies in size from species to species. The size of cpDNA in higher plants is around 120–160 kb.[21] The genes found on the cpDNA code for mRNAs that are responsible for producing necessary components of the photosynthetic pathway as well as coding for tRNAs, rRNAs, RNA polymerase subunits, and ribosomal protein subunits.[32] Like mtDNA, cpDNA is not fully autonomous and relies upon nuclear gene products for replication and production of chloroplast proteins. Chloroplasts contain multiple copies of cpDNA and the number can vary not only from species to species or cell type to cell type, but also within a single cell depending upon the age and stage of development of the cell. For example, cpDNA content in the chloroplasts of young cells, during the early stages of development where the chloroplasts are in the form of indistinct proplastids, are much higher than those present when that cell matures and expands, containing fully mature plastids.[33]
Circular
Extrachromosomal circular DNA (eccDNA) are present in all eukaryotic cells, are usually derived from genomic DNA, and consist of repetitive sequences of DNA found in both coding and non-coding regions of chromosomes. EccDNA can vary in size from less than 2000 base pairs to more than 20,000 base pairs.[34] In plants, eccDNA contain repeated sequences similar to those that are found in the centromeric regions of the chromosomes and in repetitive satellite DNA.[35] In animals, eccDNA molecules have been shown to contain repetitive sequences that are seen in satellite DNA, 5S ribosomal DNA and telomere DNA.[34] Certain organisms, such as yeast, rely on chromosomal DNA replication to produce eccDNA[35] whereas eccDNA formation can occur in other organisms, such as mammals, independently of the replication process.[36] The function of eccDNA have not been widely studied, but it has been proposed that the production of eccDNA elements from genomic DNA sequences add to the plasticity of the eukaryotic genome and can influence genome stability, cell aging and the evolution of chromosomes.[37]
A distinct type of extrachromosomal DNA, denoted as ecDNA, is commonly observed in human cancer cells.[2][3][4] ecDNA found in cancer cells contain one or more genes that confer a selective advantage. ecDNA are much larger than eccDNA, and are visible by light microscopy. ecDNA in cancers generally range in size from 1-3 MB and beyond.[2] Large ecDNA molecules have been found in the nuclei of human cancer cells and are shown to carry many copies of driver oncogenes, which are transcribed in tumor cells. Based on this evidence it is thought that ecDNA contributes to cancer growth.
Specialized tools exist that allow ecDNA to be identified, such as
- software developed by Paul Mischel and Vineet Bafna that allows ecDNA to be identified in microscopic images
- "Circle-Seq, a method for physically isolating ecDNA from cells, removing any remaining linear DNA with enzymes, and sequencing the circular DNA that remains", developed by Birgitte Regenberg and her team at the University of Copenhagen.[38]
Viral
Viral DNA are an example of extrachromosomal DNA. Understanding viral genomes is very important for understanding the evolution and mutation of the virus.[39] Some viruses, such as HIV and oncogenic viruses, incorporate their own DNA into the genome of the host cell.[40] Viral genomes can be made up of single stranded DNA (ssDNA), double stranded DNA (dsDNA) and can be found in both linear and circular form.[41]
One example of infection of a virus constituting as extrachromosomal DNA is the human papillomavirus (HPV). The HPV DNA genome undergoes three distinct stages of replication: establishment, maintenance and amplification. HPV infects epithelial cells in the anogenital tract and oral cavity. Normally, HPV is detected and cleared by the immune system. The recognition of viral DNA is an important part of immune responses. For this virus to persist, the circular genome must be replicated and inherited during cell division.[42]
Recognition by host cell
Cells can recognize foreign cytoplasmic DNA. Understanding the recognition pathways has implications towards prevention and treatment of diseases.[43] Cells have sensors that can specifically recognize viral DNA such as the Toll-like receptor (TLR) pathway.[44]
The Toll Pathway was recognized, first in insects, as a pathway that allows certain cell types to act as sensors capable of detecting a variety of bacterial or viral genomes and PAMPS (pathogen-associated molecular patterns). PAMPs are known to be potent activators of innate immune signaling. There are approximately 10 human Toll-Like Receptors (TLRs). Different TLRs in human detect different PAMPS: lipopolysaccharides by TLR4, viral dsRNA by TLR3, viral ssRNA by TLR7/TLR8, viral or bacterial unmethylated DNA by TLR9. TLR9 has evolved to detect CpG DNA commonly found in bacteria and viruses and to initiate the production of IFN (type I interferons ) and other cytokines.[44]
Inheritance

Inheritance of extrachromosomal DNA differs from the inheritance of nuclear DNA found in chromosomes. Unlike chromosomes, ecDNA does not contain centromeres and therefore exhibits a non-Mendelian inheritance pattern that gives rise to heterogeneous cell populations. In humans, virtually all of the cytoplasm is inherited from the egg of the mother.[45] For this reason, organelle DNA, including mtDNA, is inherited from the mother. Mutations in mtDNA or other cytoplasmic DNA will also be inherited from the mother. This uniparental inheritance is an example of non-Mendelian inheritance. Plants also show uniparental mtDNA inheritance. Most plants inherit mtDNA maternally with one noted exception being the redwood Sequoia sempervirens that inherit mtDNA paternally.[46]
There are two theories why the paternal mtDNA is rarely transmitted to the offspring. One is simply the fact that paternal mtDNA is at such a lower concentration than the maternal mtDNA and thus it is not detectable in the offspring. A second, more complex theory, involves the digestion of the paternal mtDNA to prevent its inheritance. It is theorized that the uniparental inheritance of mtDNA, which has a high mutation rate, might be a mechanism to maintain the homoplasmy of cytoplasmic DNA.[46]
Clinical significance
Sometimes called EEs, extrachromosomal elements, have been associated with genomic instability in eukaryotes. Small polydispersed DNAs (spcDNAs), a type of eccDNA, are commonly found in conjunction with genome instability. SpcDNAs are derived from repetitive sequences such as satellite DNA, retrovirus-like DNA elements, and transposable elements in the genome. They are thought to be the products of gene rearrangements.
Extrachromosomal DNA (ecDNA) found in cancer have historically been referred to as Double minute chromosomes (DMs), which present as paired chromatin bodies under light microscopy. Double minute chromosomes represent ~30% of the cancer-containing spectrum of ecDNA, including single bodies and have been found to contain identical gene content as single bodies.[3] The ecDNA notation encompasses all forms of the large, oncogene-containing, extrachromosomal DNA found in cancer cells. This type of ecDNA is commonly seen in cancer cells of various histologies, but virtually never in normal cells.[3] ecDNA are thought to be produced through double-strand breaks in chromosomes or over-replication of DNA in an organism. Studies show that in cases of cancer and other genomic instability, higher levels of EEs can be observed.[5]
Mitochondrial DNA can play a role in the onset of disease in a variety of ways. Point mutations in or alternative gene arrangements of mtDNA have been linked to several diseases that affect the heart, central nervous system, endocrine system, gastrointestinal tract, eye, and kidney.[22] Loss of the amount of mtDNA present in the mitochondria can lead to a whole subset of diseases known as mitochondrial depletion syndromes (MDDs) which affect the liver, central and peripheral nervous systems, smooth muscle and hearing in humans.[23] There have been mixed, and sometimes conflicting, results in studies that attempt to link mtDNA copy number to the risk of developing certain cancers. Studies have been conducted that show an association between both increased and decreased mtDNA levels and the increased risk of developing breast cancer. A positive association between increased mtDNA levels and an increased risk for developing kidney tumors has been observed but there does not appear to be a link between mtDNA levels and the development of stomach cancer.[47]
Extrachromosomal DNA is found in Apicomplexa, which is a group of protozoa. The malaria parasite (genus Plasmodium), the AIDS-related pathogen (Toxoplasma and Cryptosporidium) are both members of the Apicomplexa group. Mitochondrial DNA (mtDNA) was found in the malaria parasite.[48] There are two forms of extrachromosomal DNA found in the malaria parasites. One of these is 6-kb linear DNA and the second is 35-kb circular DNA. These DNA molecules have been researched as potential nucleotide target sites for antibiotics.[49]
Role of ecDNA in cancer
Gene amplification is among the most common mechanisms of oncogene activation. Gene amplifications in cancer are often on extrachromosomal, circular elements.[50][4] One of the primary functions of ecDNA in cancer is to enable the tumor to rapidly reach high copy numbers, while also promoting rapid, massive cell-to-cell genetic heterogeneity.[3][8] The most commonly amplified oncogenes in cancer are found on ecDNA and have been shown to be highly dynamic, re-integrating into non-native chromosomes as homogeneous staining regions (HSRs)[51][3] and altering copy numbers and composition in response to various drug treatments.[52][7][53]
The circular shape of ecDNA differs from the linear structure of chromosomal DNA in meaningful ways that influence cancer pathogenesis.[54] Oncogenes encoded on ecDNA have massive transcriptional output, ranking in the top 1% of genes in the entire transcriptome. In contrast to bacterial plasmids or mitochondrial DNA, ecDNA are chromatinized, containing high levels of active histone marks, but a paucity of repressive histone marks. The ecDNA chromatin architecture lacks the higher-order compaction that is present on chromosomal DNA and is among the most accessible DNA in the entire cancer genome. This less-compact organization permits greater access for transcription factors and transcription machinery, directly contributing to the increased expression of oncogenes.[54]
Furthermore, research on the three-dimensional genomic landscape of cancer has shown that ecDNAs frequently cluster together within the nucleus, forming "ecDNA hubs."[55] Spatially, these hubs facilitate intermolecular enhancer–promoter interactions to promote oncogene overexpression. Research has also demonstrated that cancers containing ecDNA generate the a higher frequency of new enhancer-promoter interactions compared to other types of structural variations, making ecDNA a potent driver of epigenetic rewiring.[56] This "enhancer rewiring" mechanism allows oncogenes to interact with multiple distant regulatory elements, increasing transcriptional activity, cancer growth, and heterogeneity.
Consequently, the presence of ecDNA in cancerous tumors is associated with a poor prognosis and significantly shorter survival.[50] This aggressiveness is attributed to massive oncogene copy number increases, regulatory rewiring, and intratumoral heterogeneity, which together accelerate tumor evolution and resistance to various cancer treatments.[57][3]
References
- ↑ 1.0 1.1 1.2 "Extrachromosomal DNA in eucaryotes". Plasmid 14 (3): 177–91. November 1985. doi:10.1016/0147-619X(85)90001-0. PMID 3912782.
- ↑ 2.0 2.1 2.2 2.3 "Extrachromosomal oncogene amplification in tumour pathogenesis and evolution". Nature Reviews. Cancer 19 (5): 283–288. May 2019. doi:10.1038/s41568-019-0128-6. PMID 30872802.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 "Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity". Nature 543 (7643): 122–125. March 2017. doi:10.1038/nature21356. PMID 28178237. Bibcode: 2017Natur.543..122T.
- ↑ 4.0 4.1 4.2 "Cancer May Be Driven by DNA Outside of Chromosomes" (in en). https://www.the-scientist.com/features/cancer-may-be-driven-by-dna-outside-of-chromosomes-68590.
- ↑ 5.0 5.1 "Formation of non-random extrachromosomal elements during development, differentiation and oncogenesis". Seminars in Cancer Biology 17 (1): 56–64. February 2007. doi:10.1016/j.semcancer.2006.10.007. PMID 17116402.
- ↑ Alberts, Bruce; Bray, Dennis; Hopkin, Karen; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2014). Essential Cell Biology (Fourth ed.). New York, New York, USA: Garland Science. p. 449. ISBN 978-0-8153-4454-4.
- ↑ 7.0 7.1 "Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA". Science 343 (6166): 72–6. January 2014. doi:10.1126/science.1241328. PMID 24310612. Bibcode: 2014Sci...343...72N.
- ↑ 8.0 8.1 "Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma". Nature Genetics 50 (5): 708–717. May 2018. doi:10.1038/s41588-018-0105-0. PMID 29686388.
- ↑ "Cytoplasmic DNA is structurally different from nuclear DNA". Die Naturwissenschaften 70 (5): 252–4. May 1983. doi:10.1007/BF00405447. PMID 6877387. Bibcode: 1983NW.....70..252K.
- ↑ 10.0 10.1 Nelson, David (2008). Lehninger Principles of Biochemistry. New York: W. H. Freeman and Company. pp. 307–308. ISBN 978-0-7167-7108-1. https://archive.org/details/lehningerprincip00lehn_1/page/307.
- ↑ Watson, James (2007). Recombinant RNA: Genes and Genomes- A Short Course. New York: W. H. Freeman and Company. p. 81. ISBN 978-0-7167-2866-5.
- ↑ "Extrachromosomal genetic elements in Micrococcus". Applied Microbiology and Biotechnology 97 (1): 63–75. January 2013. doi:10.1007/s00253-012-4539-5. PMID 23138713.
- ↑ Barnum, Susan (2005). Biotechnology- An Introduction. California: Brooks / Cole. pp. 62–63. ISBN 978-0-495-11205-1.
- ↑ "From plasmids to protection: a review of DNA vaccines against infectious diseases". International Reviews of Immunology 25 (3–4): 99–123. 2006. doi:10.1080/08830180600785827. PMID 16818367.
- ↑ "Mitigating the looming vaccine crisis: production and delivery of plasmid-based vaccines". Critical Reviews in Biotechnology 31 (1): 32–52. March 2011. doi:10.3109/07388551.2010.483460. PMID 20879832. http://eprints.ums.edu.my/20570/1/Mitigating%20the%20Looming%20Vaccine%20crisis.pdf.
- ↑ 16.0 16.1 16.2 "Linear plasmids and chromosomes in bacteria". Molecular Microbiology 10 (5): 917–22. December 1993. doi:10.1111/j.1365-2958.1993.tb00963.x. PMID 7934868. https://zenodo.org/record/1230611.
- ↑ 17.0 17.1 "Microbial linear plasmids". Applied Microbiology and Biotechnology 47 (4): 329–36. April 1997. doi:10.1007/s002530050936. PMID 9163946.
- ↑ Dance, Amber (16 July 2021). "Massive DNA 'Borg' structures perplex scientists" (in en). Nature 595 (7869): 636. doi:10.1038/d41586-021-01947-3. Bibcode: 2021Natur.595..636D.
- ↑ Andrew, Shakespeare, William Gurr (30 July 2021). "Previously undiscovered DNA 'borgs' found on California wetlands" (in en). The Independent. https://www.independent.co.uk/climate-change/news/dna-wetlands-borgs-california-b1893312.html.
- ↑ Al-Shayeb, Basem; Schoelmerich, Marie C.; West-Roberts, Jacob; Valentin-Alvarado, Luis E.; Sachdeva, Rohan; Mullen, Susan; Crits-Christoph, Alexander; Wilkins, Michael J.; Williams, Kenneth H.; Doudna, Jennifer A.; Banfield, Jillian F. (10 July 2021). "Borgs are giant extrachromosomal elements with the potential to augment methane oxidation". bioRxiv 10.1101/2021.07.10.451761.
- ↑ 21.0 21.1 21.2 Lodish, Harvey (2013). Molecular Cell Biology, 7th edition. New York: W.H. Freeman and Company. pp. 245–251. ISBN 978-1-4641-2398-6.
- ↑ 22.0 22.1 22.2 "Mitochondrial DNA and disease". Lancet 354 Suppl 1 (9176): SI17-21. July 1999. doi:10.1016/S0140-6736(99)90244-1. PMID 10437851.
- ↑ 23.0 23.1 23.2 "Quantitative evaluation of the mitochondrial DNA depletion syndrome". Clinical Chemistry 56 (7): 1119–27. July 2010. doi:10.1373/clinchem.2009.141549. PMID 20448188.
- ↑ "Mitochondrial DNA repair pathways". Journal of Bioenergetics and Biomembranes 31 (4): 391–8. August 1999. doi:10.1023/A:1005484004167. PMID 10665528.
- ↑ "Mitochondrial DNA, chloroplast DNA and the origins of development in eukaryotic organisms". Biology Direct 5 (42): 42. June 2010. doi:10.1186/1745-6150-5-42. PMID 20587059.
- ↑ "Genetic aspects of mitochondrial genome evolution". Molecular Phylogenetics and Evolution 69 (2): 328–38. November 2013. doi:10.1016/j.ympev.2012.10.020. PMID 23142697. Bibcode: 2013MolPE..69..328B.
- ↑ "Rates and patterns of chloroplast DNA evolution". Proceedings of the National Academy of Sciences of the United States of America 91 (15): 6795–801. July 1994. doi:10.1073/pnas.91.15.6795. PMID 8041699. Bibcode: 1994PNAS...91.6795C.
- ↑ "Cytogenomic analyses reveal the structural plasticity of the chloroplast genome in higher plants". The Plant Cell 13 (2): 245–54. February 2001. doi:10.1105/tpc.13.2.245. PMID 11226183. Bibcode: 2001PlanC..13..245L.
- ↑ Aronsson, Henrik; Sandelius, Anna Stina (2009). The chloroplast interactions with the environment ([Online-Ausg.]. ed.). Berlin: Springer. p. 18. ISBN 978-3-540-68696-5.
- ↑ "Circular chloroplast chromosomes: the grand illusion". The Plant Cell 16 (7): 1661–6. July 2004. doi:10.1105/tpc.160771. PMID 15235123. Bibcode: 2004PlanC..16.1661B.
- ↑ "Motif analysis unveils the possible co-regulation of chloroplast genes and nuclear genes encoding chloroplast proteins". Plant Molecular Biology 80 (2): 177–87. September 2012. doi:10.1007/s11103-012-9938-6. PMID 22733202. Bibcode: 2012PMolB..80..177W.
- ↑ "Essential nucleoid proteins in early chloroplast development". Trends in Plant Science 18 (4): 186–94. April 2013. doi:10.1016/j.tplants.2012.11.003. PMID 23246438. Bibcode: 2013TPS....18..186P.
- ↑ "The loss of DNA from chloroplasts as leaves mature: fact or artefact?". Journal of Experimental Botany 60 (11): 3005–10. 2009. doi:10.1093/jxb/erp158. PMID 19454766.
- ↑ 34.0 34.1 "Extrachromosomal circular DNA derived from tandemly repeated genomic sequences in plants". The Plant Journal 53 (6): 1027–34. March 2008. doi:10.1111/j.1365-313X.2007.03394.x. PMID 18088310. Bibcode: 2008PlJ....53.1027C.
- ↑ "Replication independent formation of extrachromosomal circular DNA in mammalian cell-free system". PLOS ONE 4 (7). July 2009. doi:10.1371/journal.pone.0006126. PMID 19568438. Bibcode: 2009PLoSO...4.6126C.
- ↑ "Extrachromosomal circles of satellite repeats and 5S ribosomal DNA in human cells". Mobile DNA 1 (1): 11. March 2010. doi:10.1186/1759-8753-1-11. PMID 20226008.
- ↑ "The curious DNA circles that make treating cancer so hard". https://cen.acs.org/pharmaceuticals/oncology/curious-DNA-circles-make-treating/98/i40.
- ↑ "Viral mutation rates". Journal of Virology 84 (19): 9733–48. October 2010. doi:10.1128/JVI.00694-10. PMID 20660197. Bibcode: 2010JVir...84.9733S.
- ↑ Silverthorn, Dee Unglaub (2007). Human Physiology. Peason/Benjamin Cummings.
- ↑ "Viral Genomes". https://www.ncbi.nlm.nih.gov/labs/virus/vssi/.
- ↑ "Inactivation of p53 rescues the maintenance of high risk HPV DNA genomes deficient in expression of E6". PLOS Pathogens 9 (10). October 2013. doi:10.1371/journal.ppat.1003717. PMID 24204267.
- ↑ "Cytoplasmic DNA innate immune pathways". Immunological Reviews 243 (1): 99–108. September 2011. doi:10.1111/j.1600-065X.2011.01051.x. PMID 21884170.
- ↑ 44.0 44.1 "Innate immune DNA sensing pathways: STING, AIMII and the regulation of interferon production and inflammatory responses". Current Opinion in Immunology 23 (1): 10–20. February 2011. doi:10.1016/j.coi.2010.12.015. PMID 21239155.
- ↑ An Introduction to Genetic Analysis. New York: W.H.Freeman. 2000. https://www.ncbi.nlm.nih.gov/books/NBK22059/.
- ↑ 46.0 46.1 "Maternal inheritance of mitochondrial DNA by diverse mechanisms to eliminate paternal mitochondrial DNA". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1833 (8): 1979–84. August 2013. doi:10.1016/j.bbamcr.2013.03.010. PMID 23524114.
- ↑ "Mitochondrial DNA copy number is associated with breast cancer risk". PLOS ONE 8 (6). June 2013. doi:10.1371/journal.pone.0065968. PMID 23776581. Bibcode: 2013PLoSO...865968T.
- ↑ "Extrachromosomal DNA in the Apicomplexa". Microbiology and Molecular Biology Reviews 61 (1): 1–16. March 1997. doi:10.1128/mmbr.61.1.1-16.1997. PMID 9106361.
- ↑ "Maternal inheritance of extrachromosomal DNA in malaria parasites". Molecular and Biochemical Parasitology 65 (1): 95–8. May 1994. doi:10.1016/0166-6851(94)90118-X. PMID 7935632.
- ↑ 50.0 50.1 "Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers". Nature Genetics 52 (8): 891–897. August 2020. doi:10.1038/s41588-020-0678-2. PMID 32807987.
- ↑ "Gene amplification as double minutes or homogeneously staining regions in solid tumors: origin and structure". Genome Research 20 (9): 1198–206. September 2010. doi:10.1101/gr.106252.110. PMID 20631050.
- ↑ "Elimination of extrachromosomally amplified MYC genes from human tumor cells reduces their tumorigenicity". Proceedings of the National Academy of Sciences of the United States of America 89 (17): 8165–9. September 1992. doi:10.1073/pnas.89.17.8165. PMID 1518843. Bibcode: 1992PNAS...89.8165V.
- ↑ "Structure and evolution of double minutes in diagnosis and relapse brain tumors". Acta Neuropathologica 137 (1): 123–137. January 2019. doi:10.1007/s00401-018-1912-1. PMID 30267146.
- ↑ 54.0 54.1 "Circular ecDNA promotes accessible chromatin and high oncogene expression". Nature 575 (7784): 699–703. November 2019. doi:10.1038/s41586-019-1763-5. PMID 31748743. Bibcode: 2019Natur.575..699W.
- ↑ Hung, King L.; Yost, Kathryn E.; Xie, Liangqi; Shi, Quanming; Helmsauer, Konstantin; Luebeck, Jens; Schöpflin, Robert; Lange, Joshua T. et al. (December 2021). "ecDNA hubs drive cooperative intermolecular oncogene expression" (in en). Nature 600 (7890): 731–736. doi:10.1038/s41586-021-04116-8. ISSN 1476-4687. PMID 34819668. Bibcode: 2021Natur.600..731H.
- ↑ Yost, Kathryn E.; Zhao, Yanding; Hung, King L.; Zhu, Kaiyuan; Xu, Duo; Corces, M. Ryan; Shams, Shadi; Louie, Bryan H. et al. (May 2025). "Three-dimensional genome landscape of primary human cancers" (in en). Nature Genetics 57 (5): 1189–1200. doi:10.1038/s41588-025-02188-0. ISSN 1546-1718. PMID 40355593.
- ↑ McKie, Robin; Rose, J. C.; Chen, C. Y.; Pichugin, Y.; Xie, L.; Tang, J.; Hung, K. L.; Yost, K. E. et al. (February 18, 2023). 'Bond villain' DNA could transform cancer treatment, scientists say. 54. pp. 1527–1533. doi:10.1038/s41588-022-01177-x. PMID 36123406. PMC 9534767. https://www.theguardian.com/science/2023/feb/18/bond-villain-dna-could-transform-cancer-treatment-scientists-say.
Further reading
- "Mitochondrial DNA nucleoid structure". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 1819 (9–10): 914–20. 2012. doi:10.1016/j.bbagrm.2011.11.005. PMID 22142616.
- "New insight on the role of extrachromosomal retroviral DNA". Leukemia 11 (9): 1395–9. September 1997. doi:10.1038/sj.leu.2400776. PMID 9305590.
- "Extrachromosomal circular DNA of tandemly repeated genomic sequences in Drosophila". Genome Research 13 (6A): 1133–45. June 2003. doi:10.1101/gr.907603. PMID 12799349.
- "Formation of extrachromosomal circles from telomeric DNA in Xenopus laevis". EMBO Reports 3 (12): 1168–74. December 2002. doi:10.1093/embo-reports/kvf240. PMID 12446568.
- "Extrachromosomal genes: a powerful tool in gene targeting approaches". Gene Therapy 9 (11): 679–82. June 2002. doi:10.1038/sj.gt.3301749. PMID 12032686.
- Cummings, Donald (1979). Extrachromosomal DNA. New York: Academic Press Inc..
- "Studies on extrachromosomal DNA elements. Replication of the colicinogenic factor Col E1 in two temperature sensitive mutants of Escherichia coli defective in DNA replication". European Journal of Biochemistry 15 (2): 311–20. August 1970. doi:10.1111/j.1432-1033.1970.tb01009.x. PMID 4926129.
- "Extrachromosomal plasmids in the plant pathogenic fungus Rhizoctonia solani". Current Genetics 25 (5): 423–31. May 1994. doi:10.1007/BF00351781. PMID 8082188.
- "Extrachromosomal inheritance: hereditary symbionts, mitochondria, chiloroplasts". Annual Review of Genetics 5: 361–406. 1971. doi:10.1146/annurev.ge.05.120171.002045. PMID 16097660.
- "Extrachromosomal microDNAs and chromosomal microdeletions in normal tissues". Science 336 (6077): 82–6. April 2012. doi:10.1126/science.1213307. PMID 22403181. Bibcode: 2012Sci...336...82S.
- "One ring to rule them all? Genome sequencing provides new insights into the 'master circle' model of plant mitochondrial DNA structure". The New Phytologist 200 (4): 978–85. December 2013. doi:10.1111/nph.12395. PMID 24712049. Bibcode: 2013NewPh.200..978S.
- "Sociobiological control of plasmid copy number in bacteria". PLOS ONE 5 (2). February 2010. doi:10.1371/journal.pone.0009328. PMID 20195362. Bibcode: 2010PLoSO...5.9328W.
 |
