Biology:Hydrogenosome
A hydrogenosome is a membrane-enclosed organelle found in some anaerobic ciliates, flagellates, fungi, and three species of loriciferans.[1] Hydrogenosomes are highly variable organelles that have presumably evolved from protomitochondria to produce molecular hydrogen and ATP in anaerobic conditions.[2]
Hydrogenosomes were discovered in 1973 by D. G. Lindmark and M. Müller. Because hydrogenosomes hold evolutionary lineage significance for organisms living in anaerobic or oxygen-stressed environments, many research institutions have since documented their findings on how the organelle differs in various sources.[3]
History
Hydrogenosomes were isolated, purified, biochemically characterized and named in the early 1970s by Lindmark and Müller at Rockefeller University. In addition to this seminal study on hydrogenosomes, they also demonstrated for the first time the presence of pyruvate:ferredoxin oxido-reductase and hydrogenase in eukaryotes.[3] Further studies were subsequently conducted on the biochemical cytology and subcellular organization of several anaerobic protozoan parasites (ex:Trichomonas vaginalis, Tritrichomonas foetus, Giardia lamblia, and Entamoeba sp.).[2]
Using information obtained from hydrogenosomal and biochemical cytology studies these researchers determined the mode of action of metronidazole (Flagyl). Today, metronidazole is recognized as a standard chemotherapeutic agent for the treatment of anaerobic infections.[4][5]
Since their discovery, hydrogenosomes have been found in a variety of anaerobic unicellular ciliates, flagellates, and fungi. The most notable amongst these is the parasitic Trichomonas vaginalis.[6]
Description
Hydrogenosomes are organelles that are speculated to have evolved from mitochondria to provide a different mechanism for anaerobic ATP synthesis utilizing pyruvate. The reaction results in the production of molecular hydrogen, from which the organelle receives its name.[3]
Hydrogenosomes range from 0.5-2 micrometers and are bound by a double membrane. They are most often dumbell-shaped and found in large complexes of stacked hydrogenosomes. These stacks range from 4 or 5 (called juvenile complexes) to 20 or more hydrogenosomes.[2]
In most cases, hydrogenosomes are genomeless, as a majority of the mitochondrial genome was transferred to the nucleus; because of this, all hydrogenosomal proteins are imported to the organelle.[7][8] However, a hydrogenosomal genome has been detected in the cockroach ciliate Nyctotherus ovalis, and the stramenopile Blastocystis.[9]
Due to the fact that many organisms have evolved to fit their anaerobic environments, a multitude of organisms have independently evolved hydrogenosomes or structures with similar functions. The similarity between Nyctotherus and Blastocystis, which are only distantly related, is believed to be the result of convergent evolution, and calls into question whether there is a clear-cut distinction between mitochondria, hydrogenosomes, and mitosomes (another kind of degenerate mitochondria).[2][9]
Source organisms
A non-exhaustive list of organisms containing hydrogenosomes includes:
- parabasalid flagellates (e.g. Trichomonas vaginalis, Tritrichomonas foetus, Histomonas meleagridis)
- preaxostylid flagellates (e.g. Trimastix pyriformis)
- heterolobosean amoeboflagellates (e.g. Psalteriomonas lanterna)
- anaerobic ciliates (e.g. Nyctotherus ovalis, Metopus palaeformis, Trimyema compressum, Caenomorpha uniserialis, Dasytricha ruminantium)
- anaerobic chytridiomycete fungi (e.g. Neocallimastix spp., Piromyces spp.)
The vast variety of source organisms can be accredited to the theorized convergent evolution of hydrogenosomes from mitochondria to fit an anaerobic environment.[2][7][9]
In 2010, scientists have also reported their discovery of the first known anaerobic metazoans with hydrogenosome-like organelles. Three multicellular species of Loricifera — Spinoloricus nov. sp., Rugiloricus nov. sp. and Pliciloricus nov. sp. — have been found deep in Mediterranean sediments, and use hydrogenosomes in their anaerobic metabolism cycle.[10]
ATP synthesis
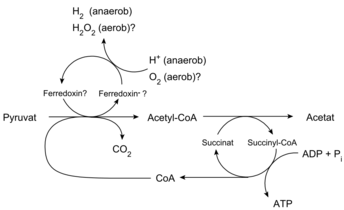
The hydrogenosomes of trichomonads (the most studied of the hydrogenosome-containing microorganisms) produce molecular hydrogen, acetate, carbon dioxide and ATP by the combined actions of pyruvate:ferredoxin oxido-reductase, hydrogenase, acetate:succinate CoA transferase and succinate thiokinase. Superoxide dismutase, malate dehydrogenase (decarboxylating), ferredoxin, adenylate kinase and NADH:ferredoxin oxido-reductase are also localized in the hydrogenosome.[11]
See also
References
- ↑ "ScienceShot: Animals That Live Without Oxygen" (in en). https://www.science.org/content/article/scienceshot-animals-live-without-oxygen.
- ↑ 2.0 2.1 2.2 2.3 2.4 "The hydrogenosomes of Psalteriomonas lanterna". BMC Evolutionary Biology 9 (1): 287. December 2009. doi:10.1186/1471-2148-9-287. PMID 20003182. Bibcode: 2009BMCEE...9..287D.
- ↑ 3.0 3.1 3.2 Lindmark, Donald G.; Müller, Miklós (1973-11-25). "Hydrogenosome, a Cytoplasmic Organelle of the Anaerobic Flagellate Tritrichomonas foetus, and Its Role in Pyruvate Metabolism" (in en). Journal of Biological Chemistry 248 (22): 7724–7728. doi:10.1016/S0021-9258(19)43249-3. ISSN 0021-9258. PMID 4750424.
- ↑ "Flagyl, Flagyl ER (metronidazole) dosing, indications, interactions, adverse effects, and more". https://reference.medscape.com/drug/flagyl-metronidazole-342566#showall.
- ↑ "Alternative pathway of metronidazole activation in Trichomonas vaginalis hydrogenosomes". Antimicrobial Agents and Chemotherapy 49 (12): 5033–6. December 2005. doi:10.1128/AAC.49.12.5033-5036.2005. PMID 16304169.
- ↑ "The Trichomonas vaginalis hydrogenosome proteome is highly reduced relative to mitochondria, yet complex compared with mitosomes". International Journal for Parasitology 41 (13–14): 1421–34. November 2011. doi:10.1016/j.ijpara.2011.10.001. PMID 22079833.
- ↑ 7.0 7.1 "The core components of organelle biogenesis and membrane transport in the hydrogenosomes of Trichomonas vaginalis". PLOS ONE 6 (9). 2011-09-15. doi:10.1371/journal.pone.0024428. PMID 21935410. Bibcode: 2011PLoSO...624428R.
- ↑ "Hsp60 is targeted to a cryptic mitochondrion-derived organelle ("crypton") in the microaerophilic protozoan parasite Entamoeba histolytica". Molecular and Cellular Biology 19 (3): 2198–205. March 1999. doi:10.1128/MCB.19.3.2198. PMID 10022906.
- ↑ 9.0 9.1 9.2 "Organelles in Blastocystis that blur the distinction between mitochondria and hydrogenosomes". Current Biology 18 (8): 580–5. April 2008. doi:10.1016/j.cub.2008.03.037. PMID 18403202. Bibcode: 2008CBio...18..580S.
- ↑ "The first metazoa living in permanently anoxic conditions". BMC Biology 8: 30. April 2010. doi:10.1186/1741-7007-8-30. PMID 20370908.
- ↑ "Diversity and reductive evolution of mitochondria among microbial eukaryotes". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 365 (1541): 713–27. March 2010. doi:10.1098/rstb.2009.0224. PMID 20124340.
 |
