Biology:PA clan of proteases
| PA clan of proteases | |
|---|---|
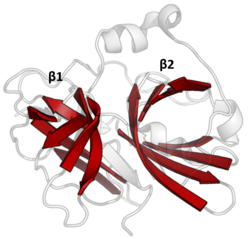 The double β-barrels that characterise the PA clan are highlighted in red. (TEV protease, PDB: 1lvm) | |
| Identifiers | |
| Symbol | N/A |
| Pfam clan | CL0124 |
| InterPro | IPR009003 |
| SCOP2 | 50494 / SCOPe / SUPFAM |
| Membranome | 319 |
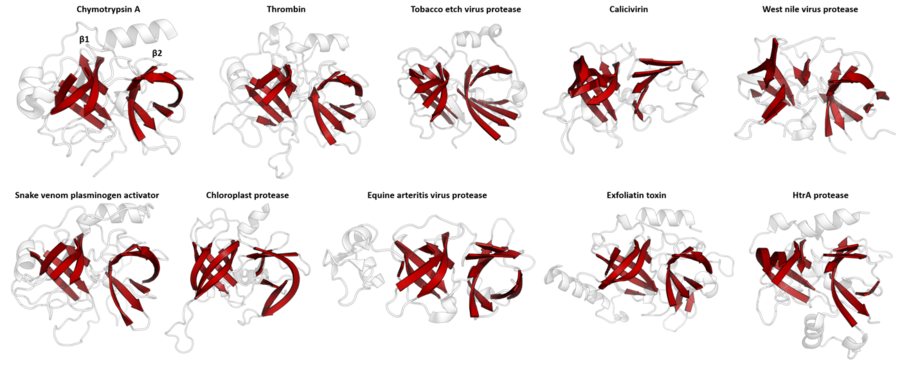
The PA clan (Proteases of mixed nucleophile, superfamily A) is the largest group of proteases with common ancestry as identified by structural homology. Members have a chymotrypsin-like fold and similar proteolysis mechanisms but can have identity of <10%. The clan contains both cysteine and serine proteases (different nucleophiles).[1][2] PA clan proteases can be found in plants,[3] animals,[3] fungi,[3] eubacteria,[4] archaea[5][6] and viruses.[2]
The common use of the catalytic triad for hydrolysis by multiple clans of proteases, including the PA clan, represents an example of convergent evolution.[7] The differences in the catalytic triad within the PA clan is also an example of divergent evolution of active sites in enzymes.[2]
History
In the 1960s, the sequence similarity of several proteases indicated that they were evolutionarily related.[8] These were grouped into the chymotrypsin-like serine proteases[9] (now called the S1 family). As the structures of these, and other proteases were solved by X-ray crystallography in the 1970s and 80s, it was noticed that several viral proteases such as Tobacco Etch Virus protease showed structural homology despite no discernible sequence similarity and even a different nucleophile.[2][10][11] Based on structural homology, a superfamily was defined and later named the PA clan (by the MEROPS classification system). As more structures are solved, more protease families have been added to the PA clan superfamily.[12][13]
Etymology
The P refers to Proteases of mixed nucleophile. The A indicates that it was the first such clan to be identified (there also exist the PB, PC, PD and PE clans).[1]
Structure

Despite retaining as little as 10% sequence identity, PA clan members isolated from viruses, prokaryotes and eukaryotes show structural homology and can be aligned by structural similarity (e.g. with DALI).[3]
Double β-barrel
PA clan proteases all share a core motif of two β-barrels with covalent catalysis performed by an acid-histidine-nucleophile catalytic triad motif. The barrels are arranged perpendicularly beside each other with hydrophobic residues holding them together as the core scaffold for the enzyme. The triad residues are split between the two barrels so that catalysis takes place at their interface.[14]
Viral protease loop
In addition to the double β-barrel core, some viral proteases (such as TEV protease) have a long, flexible C-terminal loop that forms a lid that completely covers the substrate and create a binding tunnel. This tunnel contains a set of tight binding pockets such that each side chain of the substrate peptide (P6 to P1’) is bound in a complementary site (S6 to S1’) and specificity is endowed by the large contact area between enzyme and substrate.[11] Conversely, cellular proteases that lack this loop, such as trypsin have broader specificity.
Evolution and function
Catalytic activity

Structural homology indicates that the PA clan members are descended from a common ancestor of the same fold. Although PA clan proteases use a catalytic triad perform 2-step nucleophilic catalysis,[7] some families use serine as the nucleophile whereas others use cysteine.[2] The superfamily is therefore an extreme example of divergent enzyme evolution since during evolutionary history, the core catalytic residue of the enzyme has switched in different families.[15] In addition to their structural similarity, directed evolution has been shown to be able to convert a cysteine protease into an active serine protease.[16] All cellular PA clan proteases are serine proteases, however there are both serine and cysteine protease families of viral proteases.[7] The majority are endopeptidases, with the exception being the S46 family of exopeptidases.[17][18]
Biological role and substrate specificity
In addition to divergence in their core catalytic machinery, the PA clan proteases also show wide divergent evolution in function. Members of the PA clan can be found in eukaryotes, prokaryotes and viruses and encompass a wide range of functions. In mammals, some are involved in blood clotting (e.g. thrombin) and so have high substrate specificity as well as digestion (e.g. trypsin) with broad substrate specificity. Several snake venoms are also PA clan proteases, such as pit viper haemotoxin and interfere with the victim's blood clotting cascade. Additionally, bacteria such as Staphylococcus aureus secrete exfoliative toxin which digest and damage the host's tissues. Many viruses express their genome as a single, massive polyprotein and use a PA clan protease to cleave this into functional units (e.g. polio, norovirus, and TEV proteases).[19][20]
There are also several pseudoenzymes in the superfamily, where the catalytic triad residues have been mutated and so function as binding proteins.[21] For example, the heparin-binding protein Azurocidin has a glycine in place of the nucleophile and a serine in place of the histidine.[22]
Families
Within the PA clan (P=proteases of mixed nucleophiles), families are designated by their catalytic nucleophile (C=cysteine proteases, S=serine proteases). Despite the lack of sequence homology for the PA clan as a whole, individual families within it can be identified by sequence similarity.
| Family | Examples | Known structure? |
|---|---|---|
| C03 | polio-type picornain 3C (poliovirus) | Yes |
| C04 | tobacco etch virus protease (tobacco etch virus ) | Yes |
| C24 | rabbit hemorrhagic disease virus 3C-like peptidase (rabbit hemorrhagic disease virus) | No |
| C30 | porcine transmissible gastroenteritis virus-type main peptidase (transmissible gastroenteritis virus) | Yes |
| C37 | calicivirin (Southampton virus) | Yes |
| C62 | gill-associated virus 3C-like peptidase (gill-associated virus) | No |
| C74 | pestivirus NS2 peptidase (bovine viral diarrhea virus 1) | No |
| C99 | iflavirus processing peptidase (Ectropis obliqua picorna-like virus) | No |
| C107 | alphamesonivirus 3C-like peptidase (Cavally virus) | No |
| S01 | chymotrypsin A (Bos taurus) | Yes |
| S03 | togavirin (Sindbis virus) | Yes |
| S06 | IgA specific serine endopeptidase (Neisseria gonorrhoeae) | Yes |
| S07 | flavivirin (yellow fever virus) | No |
| S29 | hepacivirin (hepatitis C virus) | Yes |
| S30 | potyvirus P1 peptidase (plum pox virus) | No |
| S31 | pestivirus NS3 polyprotein peptidase (bovine viral diarrhea virus 1) | No |
| S32 | equine arterivirus serine peptidase (equine arteritis virus) | Yes |
| S39 | sobemovirus peptidase (cocksfoot mottle virus) | Yes |
| S46 | dipeptidyl-peptidase 7 (Porphyromonas gingivalis) | No |
| S55 | SpoIVB peptidase (Bacillus subtilis) | No |
| S64 | Ssy5 peptidase (Saccharomyces cerevisiae) | No |
| S65 | picornain-like cysteine peptidase (Breda-1 torovirus) | No |
| S75 | White bream virus serine peptidase (White bream virus) | No |
See also
- Protease
- cysteine-
- serine-
- threonine-
- aspartic-
- metallo-
- Catalytic triad
- Homology
- MEROPS
- Protein family
- Protein superfamily
- Protein structure
- Structural alignment
References
- ↑ 1.0 1.1 "MEROPS: the database of proteolytic enzymes, their substrates and inhibitors". Nucleic Acids Research 40 (Database issue): D343-50. January 2012. doi:10.1093/nar/gkr987. PMID 22086950.
- ↑ 2.0 2.1 2.2 2.3 2.4 "Viral cysteine proteases are homologous to the trypsin-like family of serine proteases: structural and functional implications". Proceedings of the National Academy of Sciences of the United States of America 85 (21): 7872–6. November 1988. doi:10.1073/pnas.85.21.7872. PMID 3186696. Bibcode: 1988PNAS...85.7872B.
- ↑ 3.0 3.1 3.2 3.3 "Modeling and structural analysis of PA clan serine proteases". BMC Research Notes 5. May 2012. doi:10.1186/1756-0500-5-256. PMID 22624962.
- ↑ "Novel features of serine protease active sites and specificity pockets: sequence analysis and modelling studies of glutamate-specific endopeptidases and epidermolytic toxins". Protein Engineering 9 (7): 591–601. July 1996. doi:10.1093/protein/9.7.591. PMID 8844831.
- ↑ "MEROPS - Archaeal S01 proteases". http://merops.sanger.ac.uk/cgi-bin/organism_famdist?family=S01;taxon=Archaea;id=peptidase.
- ↑ "Bacterial serine proteases secreted by the autotransporter pathway: classification, specificity, and role in virulence". Cellular and Molecular Life Sciences 71 (5): 745–70. March 2014. doi:10.1007/s00018-013-1355-8. PMID 23689588.
- ↑ 7.0 7.1 7.2 "Intrinsic evolutionary constraints on protease structure, enzyme acylation, and the identity of the catalytic triad". Proceedings of the National Academy of Sciences of the United States of America 110 (8): E653-61. February 2013. doi:10.1073/pnas.1221050110. PMID 23382230. Bibcode: 2013PNAS..110E.653B.
- ↑ "The phylogeny of trypsin-related serine proteases and their zymogens. New methods for the investigation of distant evolutionary relationships". Journal of Molecular Biology 92 (2): 225–59. February 1975. doi:10.1016/0022-2836(75)90225-9. PMID 1142424.
- ↑ "Conservation and variability in the structures of serine proteinases of the chymotrypsin family". Journal of Molecular Biology 258 (3): 501–37. May 1996. doi:10.1006/jmbi.1996.0264. PMID 8642605.
- ↑ "Poliovirus-encoded proteinase 3C: a possible evolutionary link between cellular serine and cysteine proteinase families". FEBS Letters 194 (2): 253–7. January 1986. doi:10.1016/0014-5793(86)80095-3. PMID 3000829. Bibcode: 1986FEBSL.194..253G.
- ↑ 11.0 11.1 "Structural basis for the substrate specificity of tobacco etch virus protease". The Journal of Biological Chemistry 277 (52): 50564–72. December 2002. doi:10.1074/jbc.M207224200. PMID 12377789.
- ↑ "Picornaviral 3C cysteine proteinases have a fold similar to chymotrypsin-like serine proteinases". Nature 369 (6475): 72–6. May 1994. doi:10.1038/369072a0. PMID 8164744. Bibcode: 1994Natur.369...72A.
- ↑ "The arterivirus nsp4 protease is the prototype of a novel group of chymotrypsin-like enzymes, the 3C-like serine proteases". The Journal of Biological Chemistry 271 (9): 4864–71. March 1996. doi:10.1074/jbc.271.9.4864. PMID 8617757.
- ↑ "Characterization of the catalytic residues of the tobacco etch virus 49-kDa proteinase". Virology 172 (1): 302–10. September 1989. doi:10.1016/0042-6822(89)90132-3. PMID 2475971.
- ↑ "Modeling and structural analysis of PA clan serine proteases" (in En). BMC Research Notes 5 (1). May 2012. doi:10.1186/1756-0500-5-256. PMID 22624962.
- ↑ "Handicap-Recover Evolution Leads to a Chemically Versatile, Nucleophile-Permissive Protease". ChemBioChem 16 (13): 1866–1869. September 2015. doi:10.1002/cbic.201500295. PMID 26097079.
- ↑ "Identification of the catalytic triad of family S46 exopeptidases, closely related to clan PA endopeptidases". Scientific Reports 4. March 2014. doi:10.1038/srep04292. PMID 24598890. Bibcode: 2014NatSR...4.4292S.
- ↑ "S46 peptidases are the first exopeptidases to be members of clan PA". Scientific Reports 4. May 2014. doi:10.1038/srep04977. PMID 24827749. Bibcode: 2014NatSR...4.4977S.
- ↑ Salvesen, Guy (2013). Rawlings, Neil. ed. Handbook of proteolytic enzymes. Boston: Academic Press. ISBN 9780123822192.
- ↑ "The catalytic triad of serine peptidases". Cellular and Molecular Life Sciences 62 (19–20): 2161–72. October 2005. doi:10.1007/s00018-005-5160-x. PMID 16003488.
- ↑ "Sequence and structural differences between enzyme and nonenzyme homologs". Structure 10 (10): 1435–51. October 2002. doi:10.1016/s0969-2126(02)00861-4. PMID 12377129.
- ↑ "Structure of HBP, a multifunctional protein with a serine proteinase fold". Nature Structural Biology 4 (4): 265–8. April 1997. doi:10.1038/nsb0497-265. PMID 9095193.
External links
- MEROPS - Comprehensive protease database
- Superfamily - A database of protein folds
 |