Biology:Halspiviridae
| Salterprovirus | |
|---|---|
| |
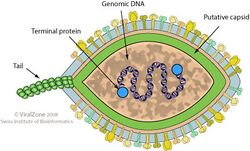
| Diagram of virion structure | |
| |
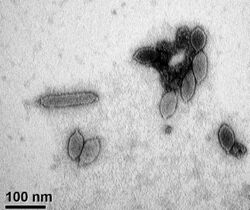
| Transmission electron micrograph of virions, negatively stained with uranyl acetate. Scale bar = 100 nm. | |
| Virus classification | |
| (unranked): | Virus |
| Family: | Halspiviridae |
| Genus: | Salterprovirus |
| Species | |
| |
| Lua error in Module:Location_map at line 522: Unable to find the specified location map definition: "Module:Location map/data/Victoria" does not exist. | |
| Locality of isolation: saltern crystalliser in Avalon, Victoria, Australia | |
| Synonyms | |
|
Salterprovirus
Salterprovirus His1
| |
Halspiviridae is a family of viruses that consists of a single genus, Salterprovirus, which consists of a single recognised species; Salterprovirus His1 (hereafter, 'His1'). This virus was isolated from hypersaline water in Australia and was able to be cultured on the halophilic archaeon Haloarcula hispanica. Like many other archaeoviruses, His1 has an approximately limoniform (lemon-shaped) virion.[1][2][3][4]
Etymology
The family name, Halspiviridae, is derived from halophilic and spindle-shaped, in reference to the habitat and virion morphology, respectively. The genus name, Salterprovirus, is derived from salty terminal protein virus, as the linear dsDNA genome has proteins attached to the 5′ termini.[1]
Taxonomy
The virion has a spindle-shaped morphology and is similar in shape to that of viruses infecting thermophilic archaea, the Fuselloviridae, and His1 was originally described as a probable member of that group.[1] However, it was later found that there is no genetic relationship and their replication strategies are entirely different, and so His1 was classified into a new group, genus Salterprovirus within the family Halspiviridae.[5] Halspiviridae has not been classified within any higher-ranked taxa.[citation needed]
Environmental DNA sequences derived from Namib salt pans indicate the presence of currently unrecognised, distant relatives of His1.[6]
Another species of virus, now named Gammapleolipovirus His2 (hereafter 'His2'), was originally considered to be related to His1,[2][3] but later analysis of the His2 virion revealed that this species actually belongs to the family Pleolipoviridae.[7]
Structure
The virus is enveloped, with limoniform or spindle-shaped morphology. Genomes are linear, around 14.5kb in length. The genome has 35 open reading frames.[3] A negatively stained electron microscope (EM) picture of His1 virions is shown on the right of this page. There is some variation in particle length (e.g. example seen left of centre), but most display the typical limoniform capsid with a short tail. High resolution micrographs and cryoEM reconstructions have been published by Hong et al. (2015),[8] who gave average dimensions of 92 x 40 nm with a 12 nm tail.
| Structure | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|
| Limoniform | Protein | Linear | Monopartite |
Replication cycle
Viral replication is cytoplasmic. Entry into the host cell is achieved by virus attachment to the host cell. An adsorption rate constant for His1 of 1.9 x 10−12 ml min−1 has been experimentally determined by Pietilä et al. (2013).[9] DNA-templated transcription is the method of transcription. Haloarcula hispanica may serve as a host. Transmission occurs via passive diffusion.[3]
| Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|
| Archea: Haloarcula hispanica | None | Injection | Lytic | Cytoplasm | Cytoplasm | Passive diffusion |
Genome
| NCBI genome ID | NC_007914 |
|---|---|
| Genome size | 14,464 nucleotides |
| Year of completion | 2020 |
The linear, dsDNA genome of His1 consists of 14,464 base-pairs (bp), has imperfect inverted terminal repeat sequences of 105 bp, and is annotated to carry 35 protein coding genes, including a gene specifying a protein-primed DNA polymerase (B-family). The ends of the genome have a protein attached.[2] The protein sequence of the polymerase is 42% identical to the polymerase specified by the gammapleolipovirus His2, even though the two viruses belong to very different taxonomic groups.[citation needed]
References
- ↑ 1.0 1.1 1.2 "His1, an archaeal virus of the Fuselloviridae family that infects Haloarcula hispanica". Journal of Virology 72 (11): 9392–5. November 1998. doi:10.1128/JVI.72.11.9392-9395.1998. PMID 9765495.
- ↑ 2.0 2.1 2.2 "His1 and His2 are distantly related, spindle-shaped haloviruses belonging to the novel virus group, Salterprovirus". Virology 350 (1): 228–39. June 2006. doi:10.1016/j.virol.2006.02.005. PMID 16530800.
- ↑ 3.0 3.1 3.2 3.3 "Viral Zone". ExPASy. http://viralzone.expasy.org/all_by_species/190.html.
- ↑ ICTV. "Virus Taxonomy: 2014 Release". http://ictvonline.org/virusTaxonomy.asp.
- ↑ "Taxonomy of prokaryotic viruses: 2018-2019 update from the ICTV Bacterial and Archaeal Viruses Subcommittee". Archives of Virology 165 (5): 1253–1260. May 2020. doi:10.1007/s00705-020-04577-8. PMID 32162068.
- ↑ "Metaviromics of Namib Desert Salt Pans: A Novel Lineage of Haloarchaeal Salterproviruses and a Rich Source of ssDNA Viruses". Viruses 8 (1): 14. January 2016. doi:10.3390/v8010014. PMID 26761024.
- ↑ "ICTV Virus Taxonomy Profile: Pleolipoviridae". The Journal of General Virology 98 (12): 2916–2917. December 2017. doi:10.1099/jgv.0.000972. PMID 29125455.
- ↑ "Lemon-shaped halo archaeal virus His1 with uniform tail but variable capsid structure". Proceedings of the National Academy of Sciences of the United States of America 112 (8): 2449–54. February 2015. doi:10.1073/pnas.1425008112. PMID 25675521. Bibcode: 2015PNAS..112.2449H.
- ↑ "Modified coat protein forms the flexible spindle-shaped virion of haloarchaeal virus His1". Environmental Microbiology 15 (6): 1674–86. June 2013. doi:10.1111/1462-2920.12030. PMID 23163639.
External links
Wikidata ☰ Q7406092 entry
 |

