Biology:Arrestin
| S-antigen; retina and pineal gland (arrestin) | |
|---|---|
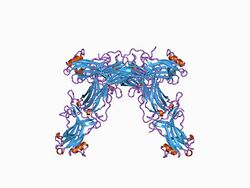
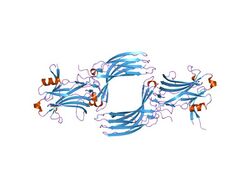
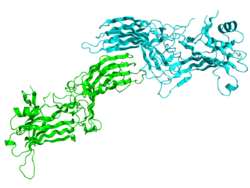 Crystallographic structure of the bovine arrestin-S.[1] | |
| Identifiers | |
| Symbol | SAG |
| Alt. symbols | arrestin-1 |
| NCBI gene | 6295 |
| HGNC | 10521 |
| OMIM | 181031 |
| RefSeq | NM_000541 |
| UniProt | P10523 |
| Other data | |
| Locus | Chr. 2 q37.1 |
| arrestin beta 1 | |
|---|---|
| Identifiers | |
| Symbol | ARRB1 |
| Alt. symbols | ARR1, arrestin-2 |
| NCBI gene | 408 |
| HGNC | 711 |
| OMIM | 107940 |
| RefSeq | NM_004041 |
| UniProt | P49407 |
| Other data | |
| Locus | Chr. 11 q13 |
| arrestin beta 2 | |
|---|---|
| Identifiers | |
| Symbol | ARRB2 |
| Alt. symbols | ARR2, arrestin-3 |
| NCBI gene | 409 |
| HGNC | 712 |
| OMIM | 107941 |
| RefSeq | NM_004313 |
| UniProt | P32121 |
| Other data | |
| Locus | Chr. 17 p13 |
| arrestin 3, retinal (X-arrestin) | |
|---|---|
| Identifiers | |
| Symbol | ARR3 |
| Alt. symbols | ARRX, arrestin-4 |
| NCBI gene | 407 |
| HGNC | 710 |
| OMIM | 301770 |
| RefSeq | NM_004312 |
| UniProt | P36575 |
| Other data | |
| Locus | Chr. X q |
Arrestins (abbreviated Arr) are a small family of proteins important for regulating signal transduction at G protein-coupled receptors.[2][3] Arrestins were first discovered in the late '80s as a part of a conserved two-step mechanism for regulating the activity of G protein-coupled receptors (GPCRs) in the visual rhodopsin system by Hermann Kühn, Scott Hall, and Ursula Wilden[4] and in the β-adrenergic system by Martin J. Lohse and co-workers.[5][6]
Function
In response to a stimulus, GPCRs activate heterotrimeric G proteins. In order to turn off this response, or adapt to a persistent stimulus, active receptors need to be desensitized. The first step in desensitization is phosphorylation of the receptor by a class of serine/threonine kinases called G protein coupled receptor kinases (GRKs). GRK phosphorylation specifically prepares the activated receptor for arrestin binding. Arrestin binding to the receptor blocks further G protein-mediated signaling and targets receptors for internalization, and redirects signaling to alternative G protein-independent pathways, such as β-arrestin signaling.[7][8][9][10][6] In addition to GPCRs, arrestins bind to other classes of cell surface receptors and a variety of other signaling proteins.[11]
Subtypes
Mammals express four arrestin subtypes and each arrestin subtype is known by multiple aliases. The systematic arrestin name (1–4) plus the most widely used aliases for each arrestin subtype are listed in bold below:
- Arrestin-1 was originally identified as the S-antigen (SAG) causing uveitis (autoimmune eye disease), then independently described as a 48 kDa protein that binds light-activated phosphorylated rhodopsin before it became clear that both are one and the same. It was later renamed visual arrestin, but when another cone-specific visual subtype was cloned the term rod arrestin was coined. This also turned out to be a misnomer: arrestin-1 expresses at comparable very high levels in both rod and cone photoreceptor cells.
- Arrestin-2 was the first non-visual arrestin cloned. It was first named β-arrestin simply because of the two GPCRs available in purified form at the time, rhodopsin and β2-adrenergic receptor, it showed preference for the latter.
- Arrestin-3. The second non-visual arrestin cloned was first termed β-arrestin-2 (retroactively changing the name of β-arrestin into β-arrestin-1), even though by that time it was clear that non-visual arrestins interact with hundreds of different GPCRs, not just with β2-adrenergic receptor. Systematic names, arrestin-2 and arrestin-3, respectively, were proposed soon after that.
- Arrestin-4 was cloned by two groups and termed cone arrestin, after photoreceptor type that expresses it, and X-arrestin, after the chromosome where its gene resides. In the HUGO database its gene is called arrestin-3.
Other arrestin-like proteins include mammalian ARRDC1–5 and thioredoxin-interacting protein (TXNIP), in addition to VPS26A and VPS26B, which fold like the true arrestins but share only roughly 10% amino acid identity and 20% homology overall.[12]
Fish and other vertebrates appear to have only three arrestins: no equivalent of arrestin-2, which is the most abundant non-visual subtype in mammals, was cloned so far. The proto-chordate Ciona intestinalis (sea squirt) has only one arrestin, which serves as visual in its mobile larva with highly developed eyes, and becomes generic non-visual in the blind sessile adult. Conserved positions of multiple introns in its gene and those of our arrestin subtypes suggest that they all evolved from this ancestral arrestin.[13] Lower invertebrates, such as roundworm Caenorhabditis elegans, also have only one arrestin. Insects have arr1 and arr2, originally termed visual arrestins because they are expressed in photoreceptors, and one non-visual subtype (kurtz in Drosophila). Later arr1 and arr2 were found to play an important role in olfactory neurons and renamed sensory. Fungi have distant arrestin relatives involved in pH sensing.
Tissue distribution
Mechanism
Arrestins block GPCR coupling to G proteins in two ways. First, arrestin binding to the cytoplasmic face of the receptor occludes the binding site for heterotrimeric G-protein, preventing its activation (desensitization).[14] Second, arrestin links the receptor to elements of the internalization machinery, clathrin and clathrin adaptor AP2, which promotes receptor internalization via coated pits and subsequent transport to internal compartments, called endosomes. Subsequently, the receptor could be either directed to degradation compartments (lysosomes) or recycled back to the plasma membrane where it can again signal. The strength of arrestin-receptor interaction plays a role in this choice: tighter complexes tend to increase the probability of receptor degradation (Class B), whereas more transient complexes favor recycling (Class A), although this rule is far from absolute.[2] More recently direct interactions between Gi/o family G proteins and Arrestin were discovered downstream of multiple receptors, regardless of canonical G protein coupling.[15] These recent findings introduce a GPCR signaling mechanism distinct from canonical G protein activation and β-arrestin desensitization in which GPCRs cause the formation of Gαi:β-arrestin signaling complexes.
Structure
Arrestins are elongated molecules, in which several intra-molecular interactions hold the relative orientation of the two domains. Unstimulated cell arrestins are localized in the cytoplasm in a basal inactive conformation. Active phosphorylated GPCRs recruit arrestin to the plasma membrane. Receptor binding induces a global conformational change that involves the movement of the two arrestin domains and the release of its C-terminal tail that contains clathrin and AP2 binding sites. Increased accessibility of these sites in receptor-bound arrestin targets the arrestin-receptor complex to the coated pit. Arrestins also bind microtubules (part of the cellular skeleton), where they assume yet another conformation, different from both free and receptor-bound form. Microtubule-bound arrestins recruit certain proteins to the cytoskeleton, which affects their activity and/or redirects it to microtubule-associated proteins.
Arrestins shuttle between cell nucleus and cytoplasm. Their nuclear functions are not fully understood, but it was shown that all four mammalian arrestin subtypes remove some of their partners, such as protein kinase JNK3 or the ubiquitin ligase Mdm2, from the nucleus. Arrestins also modify gene expression by enhancing transcription of certain genes.
Application
S-Arrestin is a protein found in mice that binds to rhodopsin to stop its activity, preventing further signaling. S-arrestin binds to G protein-coupled receptors (GPCRs), like rhodopsin, following receptor activation and phosphorylation by G protein-coupled receptor kinases (GRKs). Rhodopsin is found in rod cells of the retina, essential for vision. It detects light and initiates a signaling cascade called phototransduction. However, excessive activation can be harmful, so it must be carefully regulated. The phosphorylation of the receptor's intracellular loops and C-terminal tail creates a high-affinity binding site for S-arrestin. S-arrestin then sterically hinders further G protein coupling, effectively desensitizing the receptor and directing it towards alternative signaling pathways or internalization via clathrin-mediated endocytosis.
Binding to rhodopsin
The binding of S-arrestin to rhodopsin is specific and involves changes that occur in rhodopsin after activation. Important serine (Ser) and threonine (Thr) residues in rhodopsin's tail, particularly Thr-340 and Ser-343, are phosphorylated by enzymes called GRKs. These phosphorylated residues strongly attract S-arrestin, helping it bind tightly and effectively shut down rhodopsin's signaling.[16]
Additionally, studies of the protein structure have shown that during activation, rhodopsin's transmembrane helix 7 (TM7) and helix 8 change shape. These changes expose a binding site that interacts with a specific part of arrestin called the "finger loop." This interaction, clearly seen in the crystal structure (PDB ID: 4ZWJ), shows how arrestin fits precisely onto activated and phosphorylated rhodopsin, efficiently stopping the visual signal.[17]

|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
References
- ↑ 1.0 1.1 PDB: 1CF1; "The 2.8 A crystal structure of visual arrestin: a model for arrestin's regulation". Cell 97 (2): 257–69. April 1999. doi:10.1016/S0092-8674(00)80735-7. PMID 10219246.
- ↑ 2.0 2.1 "Regulation of receptor trafficking by GRKs and arrestins". Annual Review of Physiology 69: 451–82. 2007. doi:10.1146/annurev.physiol.69.022405.154712. PMID 17037978.
- ↑ "Transduction of receptor signals by beta-arrestins". Science 308 (5721): 512–7. April 2005. doi:10.1126/science.1109237. PMID 15845844. Bibcode: 2005Sci...308..512L.
- ↑ "Phosphodiesterase activation by photoexcited rhodopsin is quenched when rhodopsin is phosphorylated and binds the intrinsic 48-kDa protein of rod outer segments". Proceedings of the National Academy of Sciences of the United States of America 83 (5): 1174–8. March 1986. doi:10.1073/pnas.83.5.1174. PMID 3006038. Bibcode: 1986PNAS...83.1174W.
- ↑ "beta-Arrestin: a protein that regulates beta-adrenergic receptor function". Science 248 (4962): 1547–50. June 1990. doi:10.1126/science.2163110. PMID 2163110. Bibcode: 1990Sci...248.1547L.
- ↑ 6.0 6.1 "The structural basis of arrestin-mediated regulation of G-protein-coupled receptors". Pharmacology & Therapeutics 110 (3): 465–502. June 2006. doi:10.1016/j.pharmthera.2005.09.008. PMID 16460808.
- ↑ "Biased signalling: from simple switches to allosteric microprocessors". Nature Reviews. Drug Discovery 17 (4): 243–260. January 2018. doi:10.1038/nrd.2017.229. PMID 29302067.
- ↑ "Distinct conformations of GPCR-β-arrestin complexes mediate desensitization, signaling, and endocytosis". Proceedings of the National Academy of Sciences of the United States of America 114 (10): 2562–2567. February 2017. doi:10.1073/pnas.1701529114. PMID 28223524. Bibcode: 2017PNAS..114.2562C.
- ↑ "Functional competence of a partially engaged GPCR-β-arrestin complex". Nature Communications 7. November 2016. doi:10.1038/ncomms13416. PMID 27827372. Bibcode: 2016NatCo...713416K.
- ↑ "Core engagement with β-arrestin is dispensable for agonist-induced vasopressin receptor endocytosis and ERK activation". Molecular Biology of the Cell 28 (8): 1003–10. April 2017. doi:10.1091/mbc.E16-12-0818. PMID 28228552.
- ↑ "The molecular acrobatics of arrestin activation". Trends in Pharmacological Sciences 25 (2): 105–11. February 2004. doi:10.1016/j.tips.2003.12.008. PMID 15102497.
- ↑ "Chapter Two - True Arrestins and Arrestin-Fold Proteins: A Structure-Based Appraisal". Progress in Molecular Biology and Translational Science. 118. Academic Press. 2013. pp. 21-56. doi:10.1016/B978-0-12-394440-5.00002-4. ISBN 978-0-12-394440-5.
- ↑ "Arrestins: ubiquitous regulators of cellular signaling pathways". Genome Biology 7 (9): 236. 2006. doi:10.1186/gb-2006-7-9-236. PMID 17020596.
- ↑ "Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser". Nature 523 (7562): 561–7. July 2015. doi:10.1038/nature14656. PMID 26200343. Bibcode: 2015Natur.523..561K.
- ↑ "Noncanonical scaffolding of Gαi and β-arrestin by G protein–coupled receptors". Science 371 (Ahead of print). 2021. doi:10.1126/science.aay1833. PMID 33479120.
- ↑ Zhang, L (1997). "Rhodopsin phosphorylation sites and their role in S-arrestin binding". J Biol Chem 272 (23): 14762–8. doi:10.1074/jbc.272.23.14762. PMID 9169442.
- ↑ Kang, H (2015). "Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser". Nature 523 (7562): 561–567. doi:10.1038/nature14656. PMID 26200343. Bibcode: 2015Natur.523..561K.
- ↑ "Crystal structure of beta-arrestin at 1.9 A: possible mechanism of receptor binding and membrane Translocation". Structure 9 (9): 869–80. September 2001. doi:10.1016/S0969-2126(01)00644-X. PMID 11566136.
External links
- Arrestin at the US National Library of Medicine Medical Subject Headings (MeSH)
 |