Biology:Histone H4
| H4 histone, family 3 | |
|---|---|
| Identifiers | |
| Symbol | H4F3 |
| NCBI gene | 3023 |
| HGNC | 4780 |
| UniProt | P62805 |
| Other data | |
| Locus | Chr. 3 q13.13 |
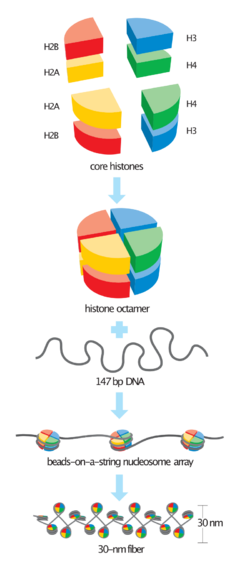
Histone H4 is one of the five main histone proteins involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and a long N-terminal tail, H4 is involved with the structure of the nucleosome of the 'beads on a string' organization. Histone proteins are highly post-translationally modified. Covalently bonded modifications include acetylation and methylation of the N-terminal tails. These modifications may alter expression of genes located on DNA associated with its parent histone octamer.[1][2] Histone H4 is an important protein in the structure and function of chromatin, where its sequence variants and variable modification states are thought to play a role in the dynamic and long term regulation of genes.
Genetics
Histone H4 is encoded in multiple genes at different loci including: HIST1H4A, HIST1H4B, HIST1H4C, HIST1H4D, HIST1H4E, HIST1H4F, HIST1H4G, HIST1H4H, HIST1H4I, HIST1H4J, HIST1H4K, HIST1H4L, HIST2H4A, HIST2H4B, HIST4H4.
Evolution
Histone proteins are among the most highly conserved eukaryotic proteins. For example, the amino acid sequence of histone H4 from a pea and cow differ at only 2 out of the 102 positions. This evolutionary conservation suggests that the functions of histone proteins involve nearly all of their amino acids so that any change is deleterious to the cell. Most changes in histone sequences are lethal; the few that are not lethal cause changes in the pattern of gene expression as well as other abnormalities.[3]
Structure
Histone H4 is a 102 to 135 amino acid protein which shares a structural motif, known as the histone fold, formed from three a-helices connected by two loops. Histone proteins H3 and H4 bind to form a H3-H4 dimer, two of these H3-H4 dimers combine to form a tetramer. This tetramer further combines with two H2a-H2b dimers to form the compact Histone octamer core.[3]
Sequence variants
Histone H4 is one of the slowest evolving proteins. There are H4 genes that are constitutively expressed throughout the cell cycle that encode for proteins that are identical in sequence to the major H4.[4] Variants in human histone H4 were only recently discovered and are very rare.[5][6][7]
Pathogenic de novo missense variants have been identified in six H4 genes (HIST1H4C, HIST1H4D, HIST1H4E, HIST1H4F, HIST1H4I, HIST1H4J) in 33 individuals total, all presenting with neurodevelopmental features of intellectual disability and motor and/or gross developmental delay, but with variable non-neurological features. Ten amino acids were affected, six of which were found recurrently. These mutations were located within either the H4 core globular domain (involved in protein-protein interaction) or C-terminal tail (involved in post-translational modification).[5][6][7]
Alternative translation
The Osteogenic Growth Peptide (OGP) is a 14-aa peptide produced from alternative translation of histone H4 mRNA, sharing the C-terminal sequence ALKRQGRTLYGFGG of histone H4. Translation is initiated at the 85th amino acid of the histone H4 mRNA, resulting in a 19-aa peptide (preOGP). This is converted into OGP through the cleavage of 5 amino-terminal residues.[8] It is found in human and rat circulation as well as regenerating bone marrow. In blood serum it is bound to α2M along with two other binding proteins that are not clearly identified. A specific receptor has not been identified, but some signaling pathways involved in its bone-regenaration function has been elucidated.[9]
Post-translational modifications
Eukaryotic organisms can produce small amounts of specialized variant core histones that differ in amino acid sequence from the main ones. These variants with a variety of covalent modifications on the N-terminal can be added to histones making possible different chromatin structures that are required for DNA function in higher eukaryotes. Potential modifications include methylation (mono-, di-, or tri-methylation) or acetylation on the tails.[3]
Methylation
Histone methylation occurs on arginine, lysine and histidine amino acids residues. Mono-, di- or tri-methylation has been discovered on histone H2A, H3 and H4.[10] Histone methylation has been associated with various cellular functions such as transcription, DNA replication, and DNA damage response including repair, heterochromatin formation, and somatic cell reprogramming. Among these biological functions, transcriptional repression and activation are the most studied.[10] Studies have shown that H4R3 methylation by PRMT1 (a histone methyltransferase) appears to be essential in vivo for the establishment or maintenance of a wide range of “active” chromatin modifications. Also methylation of histone H4 by PRMT1 was sufficient to permit subsequent acetylation on the N-terminal tail. However, acetylation of H4 inhibits its methylation by PRMT1.[11]
Acetylation
Acetylation of histones is thought to relax condensed heterochromatin as the negative charge of acetyl groups can repel the DNA phosphate backbone charges, thus reducing the histone binding affinity for DNA. This hypothesis was validated by the discovery of the histone acetyltransferase (HAT) activity of several transcriptional activator complexes.[10] Histone acetylation influences chromatin structure in several ways. First, it can provide a tag for the binding of proteins that contain areas which recognize the acetylated tails. Secondly, it can block the function of chromatin remodelers.[12] Thirdly, it neutralizes the positive charge on lysines.[12] Acetylation of histone H4 on lysine 16 (H4K16ac) is especially important for chromatin structure and function in a variety of eukaryotes and is catalyzed by specific histone lysine acetyltransferases (HATs). H4K16 is particularly interesting because this is the only acetylatable site of the H4 N-terminal tail, and can influence the formation of a compact higher-order chromatin structure.[12] Hypoacetylation of H4K16 appears to cause delayed recruitment of DNA repair proteins to sites of DNA damage in a mouse model of the premature aging syndrome Hutchinson Gilford progeria.[13] H4K16Ac also has roles in transcriptional activation and the maintenance of euchromatin.[14]
List of H4 modifications
H4S1p
H4R3me
H4K16adp
H4S47o-p
H4K91ub[15]
See also
References
- ↑ "Recognition and classification of histones using support vector machine". Journal of Computational Biology 13 (1): 102–112. 2006. doi:10.1089/cmb.2006.13.102. PMID 16472024. https://eprints.ucm.es/9328/1/31.Reche_etal_MI_2006.pdf.
- ↑ Basic Genetics. Boston: Jones and Bartlett Publishers. 1988. ISBN 978-0-86720-090-4. https://archive.org/details/basicgenetics0000hart.
- ↑ 3.0 3.1 3.2 Molecular Biology of the Cell (5th ed.). 2008. ISBN 978-0-8153-4105-5. OCLC 82473851.
- ↑ "Histone variants: deviants?". Genes & Development 19 (3): 295–310. February 2005. doi:10.1101/gad.1272805. PMID 15687254.
- ↑ 5.0 5.1 "Germline mutations affecting the histone H4 core cause a developmental syndrome by altering DNA damage response and cell cycle control". Nature Genetics 49 (11): 1642–1646. November 2017. doi:10.1038/ng.3956. PMID 28920961. https://www.nature.com/articles/ng.3956.
- ↑ 6.0 6.1 "A de novo variant in the human HIST1H4J gene causes a syndrome analogous to the HIST1H4C-associated neurodevelopmental disorder". European Journal of Human Genetics 28 (5): 674–678. May 2020. doi:10.1038/s41431-019-0552-9. PMID 31804630.
- ↑ 7.0 7.1 "Recurrent de novo missense variants across multiple histone H4 genes underlie a neurodevelopmental syndrome". American Journal of Human Genetics 109 (4): 750–758. April 2022. doi:10.1016/j.ajhg.2022.02.003. PMID 35202563.
- ↑ "Biosynthesis of osteogenic growth peptide via alternative translational initiation at AUG85 of histone H4 mRNA". The Journal of Biological Chemistry 274 (20): 14474–14481. May 1999. doi:10.1074/jbc.274.20.14474. PMID 10318873.
- ↑ "Role of Osteogenic Growth Peptide (OGP) and OGP(10-14) in Bone Regeneration: A Review". International Journal of Molecular Sciences 17 (11): 1885. November 2016. doi:10.3390/ijms17111885. PMID 27879684.
- ↑ 10.0 10.1 10.2 "Epigenetic mechanisms in mammals". Cellular and Molecular Life Sciences 66 (4): 596–612. February 2009. doi:10.1007/s00018-008-8432-4. PMID 18985277.
- ↑ "Methylation of histone H4 by arginine methyltransferase PRMT1 is essential in vivo for many subsequent histone modifications". Genes & Development 19 (16): 1885–1893. August 2005. doi:10.1101/gad.1333905. PMID 16103216.
- ↑ 12.0 12.1 12.2 "H4K16 acetylation marks active genes and enhancers of embryonic stem cells, but does not alter chromatin compaction". Genome Research 23 (12): 2053–2065. December 2013. doi:10.1101/gr.155028.113. PMID 23990607.
- ↑ "Histone H4 lysine 16 hypoacetylation is associated with defective DNA repair and premature senescence in Zmpste24-deficient mice". Proceedings of the National Academy of Sciences of the United States of America 108 (30): 12325–12330. July 2011. doi:10.1073/pnas.1102789108. PMID 21746928. Bibcode: 2011PNAS..10812325K.
- ↑ "Histone H4-K16 acetylation controls chromatin structure and protein interactions". Science 311 (5762): 844–847. February 2006. doi:10.1126/science.1124000. PMID 16469925. Bibcode: 2006Sci...311..844S. https://semanticscholar.org/paper/2a89fbf0477731f32bd48c8b12bea614b2f5fea9.
- ↑ "Epigenetic modifications poster | Abcam". https://www.abcam.com/pathways/epigenetic-modifications-poster.
See also
- nucleosome
- histone
- chromatin
- Other histone proteins involved in chromatin:
