Biology:Sex-determination system

A sex-determination system is a biological system that determines the development of the organism's sex.[1] Most organisms that create their offspring using sexual reproduction have two common sexes, males and females, and in other species there are hermaphrodites, organisms that can function reproductively as either female or male, or both.[2]
There are also some species in which only one sex is present, temporarily or permanently. This can be due to parthenogenesis, the act of a female reproducing without fertilization, mostly seen in plant species. In some plants or algae the gametophyte stage may reproduce itself, thus producing more individuals of the same sex as the parent.
In some species, sex determination is genetic: males and females have different alleles or even different genes that specify their sexual morphology. In animals this is often accompanied by chromosomal differences, generally through combinations of XY, ZW, XO, ZO chromosomes, or haplodiploidy. The sexual differentiation is generally triggered by a main gene (a "sex locus"), with a multitude of other genes following in a domino effect.
In other cases, the sex of a fetus is determined by environmental variables (such as temperature). The details of some sex-determination systems are not yet fully understood.
Some species such as various plants and fish do not have a fixed sex and instead go through life cycles and change sex based on genetic cues during corresponding life stages of their type. This could be due to environmental factors such as seasons and temperature. In some gonochoric species, a few individuals may have conditions that cause a mix of different sex characteristics.[3]
Discovery
The scientific understanding of sex determination has evolved significantly over the centuries through a series of landmark discoveries across plants, animals, and insects.
In 1694, German botanist Rudolf Jakob Camerarius conducted pioneering experiments on pollination, during which he identified the existence of distinct male and female reproductive structures in plants, including maize. His findings laid the groundwork for understanding plant reproduction and sexual differentiation .[4]
In 1866, Gregor Mendel, an Austrian monk and scientist, published his seminal work on the inheritance of traits in pea plants. These findings—now known as Mendelian inheritance—introduced the concept of heritable units (genes) passed from two gametes. Mendel's principles eventually became the foundation of modern genetics.[5]
In 1902, American zoologist C.E. McClung proposed that sex chromosomes played a central role in determining sex, based on cytological studies in insects. This was one of the earliest steps toward a chromosomal theory of sex determination.[6]
In 1903, American geneticist Nettie Stevens made a groundbreaking discovery while studying the mealworm (Tenebrio molitor), demonstrating that sex is determined by specific chromosomes, now known as X and Y chromosomes. Her findings provided the first concrete evidence for chromosomal sex determination.[7][8]
In 1917, botanist Charles Elmer Allen extended this understanding to the plant kingdom by discovering sex chromosomes in plants, confirming that the chromosomal mechanism is not exclusive to animals.[9][failed verification]
In 1922, geneticist Calvin B. Bridges introduced the genie balance theory, based on experiments with Drosophila melanogaster (fruit flies). He proposed that the ratio of X chromosomes to sets of autosomes determines sexual development, adding complexity to the previously accepted XX/XY system.[10]
In 1928, Swiss biologist Fritz Baltzer was the first to describe environmental sex determination, showing that external environmental factors, such as temperature, could influence the development of sex in certain organisms. This discovery expanded the understanding of sex determination beyond purely genetic mechanisms.[11]
Chromosomal systems
Among animals, the most common chromosomal sex determination systems are XY, XO, ZW, ZO, but with numerous exceptions.
According to the Tree of Sex database[12] (as of 2023), the known sex determination systems are:[13]
| Taxonomic group | XY | XO | ZW | ZO | Other1 | XO/XY ratio | ZO/ZW ratio |
|---|---|---|---|---|---|---|---|
| Vertebrates | 722 | 15 | 480 | 3 | 254 | 0.02 | 0.01 |
| Insects | 4415 | 1857 | 37 | 25 | 156 | 0.42 | 0.68 |
| Angiosperms | 23 | 0 | 1 | 0 | 19 | 0.00 | 0.00 |
| Total | 5160 | 1872 | 518 | 28 | 429 | 0.36 | 0.05 |
1. complex sex chromosomes, homomorphic sex chromosomes, or others
XX/XY sex chromosomes
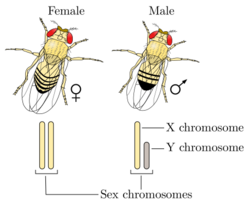
The XX/XY sex-determination system is perhaps the most familiar as it is found in humans and most other mammals, as well as in some insect species. In the XX/XY system, karyotypic females usually have two X chromosomes (XX), while karyotypic males usually have a single X and a single Y chromosome (XY). The X and Y sex chromosomes are different in shape and size from each other, unlike other chromosome pairs (autosomes), and are sometimes called allosomes. In some species, including humans, individuals remain phenotypically undifferentiated for some time during development (embryogenesis); in others, however, such as fruit flies, sexual differentiation occurs as soon fertilization occurs.[14]
Y-centered sex determination
Some species (including humans) have a gene SRY on the Y chromosome that triggers development of the male phenotype. Members of SRY-reliant species can also have other chromosomal combinations such as XXY.[14] In humans, karyotypic sex is generally determined by the presence or absence of a Y chromosome with a functional SRY gene. If the SRY gene is present and activated during fetal development, cells create testosterone and anti-müllerian hormone which typically leads to male phenotypic development.[14] In embryos that lack a functioning SRY gene, such as most XX individuals, the individual develops phenotypically female.
In Y-centered sex determination, the SRY gene is the gene that triggers male phenotype development, however multiple genes are required for this process to complete. In XY mice, lack of the gene DAX1 on the X chromosome results in sterility, but in humans it causes adrenal hypoplasia congenita.[15] However, when an extra DAX1 gene is placed on the X chromosome, the result is phenotypically female, despite the existence of SRY, since it overrides the effects of SRY.[16] Even when there are functional X chromosomes in XX females, duplication or expression of SOX9 causes testes to develop.[17][18] Gradual sex reversal in developed mice can also occur when the gene FOXL2 is removed from females.[19] Even though the gene DMRT1 is used by birds as their sex locus, species who have XY chromosomes also rely upon DMRT1, contained on chromosome 9, for sexual differentiation at some point in their formation.[14]
X-centered sex determination
Some species, such as fruit flies, use the presence of two X chromosomes to determine femaleness.[20] Species that use the number of Xs to determine sex are nonviable with an extra X chromosome.
Other variants of XX/XY sex determination
Some fish have variants of the XY sex-determination system, as well as the regular system. For example, while having an XY format, Xiphophorus nezahualcoyotl and X. milleri also have a second Y chromosome, known as Y', that creates XY' females and YY' males.[21]
At least one monotreme, the platypus, presents a particular sex determination scheme that in some ways resembles that of the ZW sex chromosomes of birds and lacks the SRY gene. The platypus has sex chromosomes . The males have , while females have . During meiosis, 5 of X form one chain, and 5 of Y form another chain. Thus, they behave effectively as a typical XY chromosomal system, except each of X and Y is broken into 5 parts, with the effect that recombinations occur very frequently at 4 particular points.[22] One of the X chromosomes is homologous to the human X chromosome, and another is homologous to the bird Z chromosome.[23]
Although it is an XY system, the platypus' sex chromosomes share no homologues with eutherian sex chromosomes.[24] Instead, homologues with eutherian sex chromosomes lie on the platypus chromosome 6, which means that the eutherian sex chromosomes were autosomes at the time that the monotremes diverged from the therian mammals (marsupials and eutherian mammals). However, homologues to the avian DMRT1 gene on platypus sex chromosomes X3 and X5 suggest that it is possible the sex-determining gene for the platypus is the same one that is involved in bird sex-determination. More research must be conducted in order to determine the exact sex determining gene of the platypus.[25]
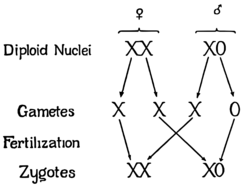
XX/X0 sex chromosomes
In this variant of the XY system, females have two copies of the sex chromosome (XX) but males have only one (X0). The 0 denotes the absence of a second sex chromosome. Generally in this method, the sex is determined by amount of genes expressed across the two chromosomes. This system is observed in a number of insects, including the grasshoppers and crickets of order Orthoptera and in cockroaches (order Blattodea). A small number of mammals also lack a Y chromosome. These include the Amami spiny rat (Tokudaia osimensis) and the Tokunoshima spiny rat (Tokudaia tokunoshimensis) and Sorex araneus, a shrew species. Transcaucasian mole voles (Ellobius lutescens) also have a form of XO determination, in which both sexes lack a second sex chromosome.[16] The mechanism of sex determination is not yet understood.[26]
The nematode C. elegans is male with one sex chromosome (X0); with a pair of chromosomes (XX) it is a hermaphrodite.[27] Its main sex gene is XOL, which encodes XOL-1 and also controls the expression of the genes TRA-2 and HER-1. These genes reduce male gene activation and increase it, respectively.[28]
ZW/ZZ sex chromosomes
The ZW sex-determination system is found in birds, some reptiles, and some insects and other organisms. The ZW sex-determination system is reversed compared to the XY system: females have two different kinds of chromosomes (ZW), and males have two of the same kind of chromosomes (ZZ). In the chicken, this was found to be dependent on the expression of DMRT1.[29] In birds, the genes FET1 and ASW are found on the W chromosome for females, similar to how the Y chromosome contains SRY.[14] However, not all species depend upon the W for their sex. For example, there are moths and butterflies that are ZW, but some have been found female with ZO, as well as female with ZZW.[27] Also, while mammals deactivate one of their extra X chromosomes when female, it appears that in the case of Lepidoptera, the males produce double the normal amount of enzymes, due to having two Z's.[27] Because the use of ZW sex determination is varied, it is still unknown how exactly most species determine their sex.[27] However, reportedly, the silkworm Bombyx mori uses a single female-specific piRNA as the primary determiner of sex.[30] Despite the similarities between the ZW and XY systems, these sex chromosomes evolved separately. In the case of the chicken, their Z chromosome is more similar to humans' autosome 9.[31] The chicken's Z chromosome also seems to be related to the X chromosome of the platypus.[32] When a ZW species, such as the Komodo dragon, reproduces parthenogenetically, usually only males are produced. This is due to the fact that the haploid eggs double their chromosomes, resulting in ZZ or WW. The ZZ become males, but the WW are not viable and are not brought to term.[33]
In both XY and ZW sex determination systems, the sex chromosome carrying the critical factors is often significantly smaller, carrying little more than the genes necessary for triggering the development of a given sex.[34]
ZZ/Z0 sex chromosomes
The ZZ/Z0 sex-determination system is found in some moths. In these insects there is one sex chromosome, Z. Males have two Z chromosomes, whereas females have one Z. Males are ZZ, while females are Z0.[35][36][37]
UV sex chromosomes
In some bryophyte and some algae species, the gametophyte stage of the life cycle, rather than being hermaphrodite, occurs as separate male or female individuals that produce male and female gametes respectively. When meiosis occurs in the sporophyte generation of the life cycle, the sex chromosomes known as U and V assort in spores that carry either the U chromosome and give rise to female gametophytes, or the V chromosome and give rise to male gametophytes.[38][39]
Mating types
The mating type in microorganisms is analogous to sex in multi-cellular organisms, and is sometimes described using those terms, though they are not necessarily correlated with physical body structures. Some species have more than two mating types. Tetrahymena, a type of ciliate, has 7 mating types
Mating types are extensively studied in fungi. Among fungi, mating type is determined by chromosomal regions called mating-type loci. Furthermore, it is not as simple as "two different mating types can mate", but rather, a matter of combinatorics. As a simple example, most basidiomycete have a "tetrapolar heterothallism" mating system: there are two loci, and mating between two individuals is possible if the alleles on both loci are different. For example, if there are 3 alleles per locus, then there would be 9 mating types, each of which can mate with 4 other mating types.[40] By multiplicative combination, it generates a vast number of mating types. For example, Schizophyllum commune, a type of fungus, has
mating types.
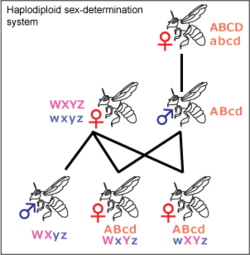
Haplodiploidy
Haplodiploidy is found in insects belonging to Hymenoptera, such as ants and bees. Sex determination is controlled by the zygosity of a complementary sex determiner (csd) locus. Unfertilized eggs develop into haploid individuals which have a single, hemizygous copy of the csd locus and are therefore males. Fertilized eggs develop into diploid individuals which, due to high variability in the csd locus, are generally heterozygous females. In rare instances diploid individuals may be homozygous, these develop into sterile males. The gene acting as a csd locus has been identified in the honeybee and several candidate genes have been proposed as a csd locus for other Hymenopterans.[41][42][43] Most females in the Hymenoptera order can decide the sex of their offspring by holding received sperm in their spermatheca and either releasing it into their oviduct or not. This allows them to create more workers, depending on the status of the colony.[44]
Polygenic sex determination
Polygenic sex determination is when the sex is primarily determined by genes that occur on multiple non-homologous chromosomes. The environment may have a limited, minor influence on sex determination. Examples include African cichlid fish (Metriaclima spp.), lemmings (Myopus schisticolor), green swordtail,[21] medaka,[21] etc. In such systems, there is typically a dominance hierarchy, where one system is dominant over another if in conflict. For example, in some species of cichlid fish from Lake Malawi, if an individual has both the XY locus (on one chromosome pair) and the WZ locus (on another chromosome pair), then the W is dominant and the individual has a female phenotype.[45]
The sex-determination system of zebrafish is polygenic. Juvenile zebrafishes (0–30 days after hatching) have both ovary-like tissue to testis tissue. They then develop into male or female adults, with the determination based on a complex interaction genes on multiple chromosomes, but not affected by environmental variations.[46][47]
Sex determination by non-coding RNA
In many insects, sex is determined by a long non-coding RNA (lncRNA), ANTSR (pronounced ant-ser) that was discovered in ants.[48] Ants that are heterozygous at the ANTSR locus become females, while homozygous (and hemizygous) embryos become males (in Hymenoptera, e.g. ants, bees, wasps, sawflies, females arise from fertilized, diploid eggs, while males develop from unfertilized, haploid eggs). The ANTSR system has been maintained for over 150 million years but still evolves rapidly and thus has lost sequence similarity among many species that use it.[49]
Other chromosomal systems
In systems with two sex chromosomes, they can be heteromorphic or homomorphic. Homomorphic sex chromosomes are almost identical in size and gene content. The two familiar kinds of sex chromosome pairs (XY and ZW) are heteromorphic. Homomorphic sex chromosomes exist among pufferfish, ratite birds, pythons, and European tree frogs. Some are quite old, meaning that there is some evolutionary force that resists their differentiation.[50] For example, three species of European tree frogs have homologous, homomorphic sex chromosomes, and this homomorphism was maintained for at least 5.4 million years by occasional recombination.[51]
The Nematocera, particularly the Simuliids and Chironomus, have sex determination regions that are labile, meaning that one species may have the sex determination region in one chromosome, but a closely related species might have the same region moved to a different non-homologous chromosome. Some species even have the sex determination region different among individuals within the same species (intraspecific variation).[52][53][54] In some species, some populations have homomorphic sex chromosomes while other populations have heteromorphic sex chromosomes.
The New Zealand frog, Leiopelma hochstetteri, uses a supernumerary sex chromosome. With zero of that chromosome, the frog develops into a male. With one or more, the frog develops into a female. One female had as many as 16 of that chromosome.[55]
Different populations of the Japanese frog Rana rugosa uses different systems. Two use homomorphic male heterogamety, one uses XX/XY, one uses ZZ/ZW. Remarkably, the X and Z chromosomes are homologous, and the Y and W as well. Dmrt1 is on autosome 1 and not sex-linked. This means that an XX female individual is genetically similar to a ZZ male individual, and an XY male individual is to a ZW female individual. The mechanism behind this is yet unclear, but it is hypothesized that during its recent evolution, the XY-to-ZW transition occurred twice.[56][57]
Clarias gariepinus uses both XX/XY and ZW/ZZ system within the species, with some populations using homomorphic XX/XY while others using heteromorphic ZW/ZZ. A population in Thailand appears to use both systems simultaneously, possibly because C. gariepinus were not native to Thailand, and were introduced from different source populations which resulted in a mixture.[58]
Multiple sex chromosomes like those of platypus also occurs in bony fish.[59] Some moths and butterflies have or .[60]
The Southern platyfish has a complex sex determination system involving 3 sex chromosomes and 4 autosomal alleles.[61][62]
Gastrotheca pseustes has C-banding heteromorphism, meaning that both males and females have XY chromosomes, but their Y chromosomes are different on one or more C-bands. Eleutherodactylus maussi has a system.[63][64]
Evolution
See [65] for a review.
Origin of sex chromosomes
Sexual chromosome pairs can arise from an autosomal pair that, for various reasons, stopped recombination, allowing for their divergence. The rate at which recombination is suppressed, and therefore the rate of sex chromosome divergence, is very different across clades.[50]
In analogy with geological strata, historical events in the evolution of sex chromosomes are called evolutionary strata. The human Y-chromosome has had about 5 strata since the origin of the X and Y chromosomes about 300 Mya from a pair of autosomes. Each stratum was formed when a pseudoautosomal region (PAR) of the Y chromosome is inverted, stopping it from recombination with the X chromosome. Over time, each inverted region decays, possibly due to Muller's ratchet.[66][67] Primate Y-chromosome evolution was rapid, with multiple inversions and shifts of the boundary of PAR.[68]
Among many species of the salamanders, the two chromosomes are only distinguished by a pericentric inversion, so that the banding pattern of the X chromosome is the same as that of Y, but with a region near the centromere reversed. (fig 7 [69]) In some species, the X is pericentrically inverted and the Y is ancestral. In other species it is the opposite. (p. 15 [69])
The gene content of the X chromosome is almost identical among placental mammals. This is hypothesized to be because the X inactivation means any change would cause serious disruption, thus subjecting it to strong purifying selection. Similarly, birds have highly conserved Z chromosomes.[57]
Neo-sex chromosomes
Neo-sex chromosomes are currently existing sex chromosomes that formed when an autosome pair fused to the previously existing sex chromosome pair. Following this fusion, the autosomal portion undergoes recombination suppression, allowing them to differentiate. Such systems have been observed in insects, reptiles, birds, and mammals. They are useful to the study of the evolution of Y chromosome degeneration and dosage compensation.[70][71]
Sex-chromosome turnover
The sex-chromosome turnover is an evolutionary phenomenon where sex chromosomes disappear or become autosomal, and autosomal chromosomes become sexual, repeatedly over evolutionary time. Some lineages have extensive turnover, but others don't. Generally, in an XY system, if the Y chromosome is degenerate, mostly different from the X chromosome, and has X dosage compensation, then turnover is unlikely. In particular, this applies to humans.[72][65][73]
The ZW and XY systems can evolve into to each other due to sexual conflict.[74]
Homomorphism and the fountain of youth
It is an evolutionary puzzle why certain sex chromosomes remain homomorphic over millions of years, especially among lineages of fishes, amphibians, and nonavian reptiles. The fountain-of-youth model states that heteromorphy results from recombination suppression, and recombination suppression results from the male phenotype, not the sex chromosomes themselves. Therefore, if some XY sex-reversed females are fertile and adaptive under some circumstances, then the X and Y chromosomes would recombine in these individuals, preventing Y chromosome decay and maintaining long-term homomorphism.[75]
Sex reversal denotes a situation where the phenotypic sex is different from the genotypic sex. While in humans, sex reversal (such as the XX male syndrome) are often infertile, sex-reversed individuals of some species are fertile under some conditions. For example, some XY-individuals in population of Chinook salmon in the Columbia River became fertile females, producing YY sons. Since Chinook salmons have homomorphic sex chromosomes, such YY sons are healthy. When YY males mate with XX females, all their progeny would be XY male if grown under normal conditions.[76]
Support for the hypothesis is found in the common frog, for which XX males and XY males both suppresses sex chromosome recombination, but XX and XY females both recombine at the same rate.[77]
Environmental systems
Temperature-dependent

Many other sex-determination systems exist. In some species of reptiles, including alligators, some turtles, and the tuatara, sex is determined by the temperature at which the egg is incubated during a temperature-sensitive period. There are no examples of temperature-dependent sex determination (TSD) in birds. Megapodes had formerly been thought to exhibit this phenomenon, but were found to actually have different temperature-dependent embryo mortality rates for each sex.[78] For some species with TSD, sex determination is achieved by exposure to hotter temperatures resulting in the offspring being one sex and cooler temperatures resulting in the other. This type of TSD is called Pattern I. For others species using TSD, it is exposure to temperatures on both extremes that results in offspring of one sex, and exposure to moderate temperatures that results in offspring of the opposite sex, called Pattern II TSD. The specific temperatures required to produce each sex are known as the female-promoting temperature and the male-promoting temperature.[79] When the temperature stays near the threshold during the temperature sensitive period, the sex ratio is varied between the two sexes.[80] Some species' temperature standards are based on when a particular enzyme is created. These species that rely upon temperature for their sex determination do not have the SRY gene, but have other genes such as DAX1, DMRT1, and SOX9 that are expressed or not expressed depending on the temperature.[79] The sex of some species, such as the Nile tilapia, Australian skink lizard, and Australian dragon lizard, has an initial bias, set by chromosomes, but can later be changed by the temperature of incubation.[21]
It is unknown how exactly temperature-dependent sex determination evolved.[81] It could have evolved through certain sexes being more suited to certain areas that fit the temperature requirements. For example, a warmer area could be more suitable for nesting, so more females are produced to increase the amount that nest next season.[81] In amniotes, environmental sex determination preceded the genetically determined systems of birds and mammals; it is thought that a temperature-dependent amniote was the common ancestor of amniotes with sex chromosomes.[82]
Other environmental systems
There are other environmental sex determination systems including location-dependent determination systems as seen in the marine worm Bonellia viridis – larvae become males if they make physical contact with a female, and females if they end up on the bare sea floor. This is triggered by the presence of a chemical produced by the females, bonellin.[83] Some species, such as some snails, practice sex change: adults start out male, then become female. In tropical clownfish, the dominant individual in a group becomes female while the other ones are male, and bluehead wrasses (Thalassoma bifasciatum) are the reverse.
Clownfish live in colonies of several small undifferentiated fish and two large fish (male and female). The male and female are the only sexually mature fish to reproduce. Clownfish are protandrous hermaphrodites, which means after they mature into males, they eventually can transform into females. They develop undifferentiated until they are needed to fill a certain role in their environment, i.e., if they receive the social and environmental cues to do so.[84]
Some species, however, have no sex-determination system. Hermaphrodite species include the common earthworm and certain species of snails. A few species of fish, reptiles, and insects reproduce by parthenogenesis and are female altogether. There are some reptiles, such as the boa constrictor and Komodo dragon that can reproduce both sexually and asexually, depending on whether a mate is available.[85]
Others
There are exceptional sex-determination systems, neither genetic nor environmental.
Cytoplasmic sex determination
The Wolbachia genus of parasitic bacteria lives inside the cytoplasm of its host, and is vertically transmitted from parents to children. They primarily infect arthropods and nematodes. Different Wolbachia can alter the reproductive abilities of its host by a variety of means, including cytoplasmic incompatibility, parthenogenesis, feminization and embryonic male killing.[86]
Mitochondrial male sterility: In many flowering plants, the mitochondria can cause hermaphrodite individuals to be unable to father offspring, effectively turning them into exclusive females. This is a form of mother's curse. It is an evolutionarily adaptive strategy for mitochondria as mitochondria are inherited exclusively from mother to offspring.[87] The first published case of mitochondrial male sterility among metazoans was reported in 2022 in the hermaphroditic snail Physa acuta.[88]
Paternal genome elimination
In some species of insects, springtails and mites, male offspring lose their paternal genome (in whole or in part) during development or in the germline. Males can either be diploid, diploid with missing sex chromosome, functionally haploid or truly haploid, depending on the mechanism of elimination.[89][87]
Monogeny
In some species of Hymenoptera (ants, bees and wasps), flies and crustaceans, all offspring of a particular individual female are either exclusively male or exclusively female.[87] The underlying mechanisms are diverse and include maternally controlled paternal genome elimination and Mendelian inherited maternal sex-determining factors.[90]
Evolution
Sex determination systems may have evolved from mating type, which is a feature of microorganisms.
Chromosomal sex determination may have evolved early in the history of eukaryotes.[91] But in plants it has been suggested to have evolved recently.[92]
The accepted hypothesis of XY and ZW sex chromosome evolution in amniotes is that they evolved at the same time, in two different branches.[93][94]
No genes are shared between the avian ZW and mammal XY chromosomes[31] and the chicken Z chromosome is similar to the human autosomal chromosome 9, rather than X or Y. This suggests not that the ZW and XY sex-determination systems share an origin but that the sex chromosomes are derived from autosomal chromosomes of the common ancestor of birds and mammals. In the platypus, a monotreme, the X1 chromosome shares homology with therian mammals, while the X5 chromosome contains an avian sex-determination gene, further suggesting an evolutionary link.[95]
However, there is some evidence to suggest that there could have been transitions between ZW and XY, such as in Xiphophorus maculatus, which have both ZW and XY systems in the same population, despite the fact that ZW and XY have different gene locations.[96][97] A recent theoretical model raises the possibility of both transitions between the XY/XX and ZZ/ZW system and environmental sex determination[98] The platypus' genes also back up the possible evolutionary link between XY and ZW, because they have the DMRT1 gene possessed by birds on their X chromosomes.[99] Regardless, XY and ZW follow a similar route. All sex chromosomes started out as an original autosome of an original amniote that relied upon temperature to determine the sex of offspring. After the mammals separated, the reptile branch further split into Lepidosauria and Archosauromorpha. These two groups both evolved the ZW system separately, as evidenced by the existence of different sex chromosomal locations.[94] In mammals, one of the autosome pair, now Y, mutated its SOX3 gene into the SRY gene, causing that chromosome to designate sex.[94][99][100] After this mutation, the SRY-containing chromosome inverted and was no longer completely homologous with its partner. The regions of the X and Y chromosomes that are still homologous to one another are known as the pseudoautosomal region.[101] Once it inverted, the Y chromosome became unable to remedy deleterious mutations, and thus degenerated.[94] There is some concern that the Y chromosome will shrink further and stop functioning in ten million years: but the Y chromosome has been strictly conserved after its initial rapid gene loss.[102][103]
There are some vertebrate species, such as the medaka fish, that evolved sex chromosomes separately; their Y chromosome never inverted and can still swap genes with the X. These species' sex chromosomes are relatively primitive and unspecialized. Because the Y does not have male-specific genes and can interact with the X, XY and YY females can be formed as well as XX males.[21] Non-inverted Y chromosomes with long histories are found in pythons and emus, each system being more than 120 million years old, suggesting that inversions are not necessarily an eventuality.[87] XO sex determination can evolve from XY sex determination with about 2 million years.[clarification needed][104]
See also
- Testis-determining factor
- Maternal influence on sex determination
- Sequential hermaphroditism
- Sex determination and differentiation (human)
References
- ↑ Schnebly, Risa Aria (2021). "Sex Determination in Humans". The Embryo Project Encyclopedia. https://embryo.asu.edu/pages/sex-determination-humans. Retrieved 2022-07-06.
- ↑ Encyclopedia of Animal Cognition and Behavior. Cham: Springer International Publishing. 2018. pp. 1–2. doi:10.1007/978-3-319-47829-6_329-1. ISBN 978-3-319-47829-6.
- ↑ The Biology of Reproduction. Cambridge University Press. 2019. pp. 116–117. ISBN 978-1-108-49985-9. https://books.google.com/books?id=AKGsDwAAQBAJ&q=Intersex%20. Retrieved 11 October 2020.
- ↑ "Rudolph Jacob Camerarius | Plant Physiology, Taxonomy & Genetics | Britannica" (in en). https://www.britannica.com/biography/Rudolph-Jacob-Camerarius.
- ↑ "Deutsches Textarchiv – Mendel, Gregor: Versuche über Pflanzen-Hybriden. In: Verhandlungen des Naturforschenden Vereines in Brünn 4 (1866), S. 3-47.". https://www.deutschestextarchiv.de/book/view/mendel_pflanzenhybriden_1866?p=14.
- ↑ MCCLUNG, C. E. (May 1902). "THE ACCESSORY CHROMOSOME—SEX DETERMINANT?". The Biological Bulletin 3 (1–2): 43–84. doi:10.2307/1535527. ISSN 0006-3185.
- ↑ Stevens, Nettie Maria (1905). Studies in spermatogenesis. Washington, D. C.: Carnegie Institution of Washington. doi:10.5962/bhl.title.56517.
- ↑ Ogilvie, Marilyn Bailey (February 2000). Stevens, Nettie Maria (1861-1912), cytologist. American National Biography Online. Oxford University Press. doi:10.1093/anb/9780198606697.article.1302015.
- ↑ The American Naturalist 51 (605). May 1917. doi:10.1086/an.1917.51.issue-605. ISSN 0003-0147.
- ↑ Bridges, Calvin B. (January 1922). "The Origin of Variations in Sexual and Sex-Limited Characters". The American Naturalist 56 (642): 51–63. doi:10.1086/279847. ISSN 0003-0147. Bibcode: 1922ANat...56...51B.
- ↑ Buchner, Paul (1928), "Ergebnisse der Symbioseforschung", Ergebnisse der Biologie (Berlin, Heidelberg: Springer Berlin Heidelberg): pp. 1–129, doi:10.1007/978-3-642-91066-1_1, ISBN 978-3-642-89210-3
- ↑ Ashman, Tia-Lynn; Bachtrog, Doris; Blackmon, Heath; Goldberg, Emma E.; Hahn, Matthew W.; Kirkpatrick, Mark; Kitano, Jun; Mank, Judith E. et al. (2014-06-24). "Tree of Sex: A database of sexual systems". Scientific Data 1. doi:10.1038/sdata.2014.15. ISSN 2052-4463. PMID 25977773.
- ↑ Hayashi, Shun; Abe, Takuya; Igawa, Takeshi; Katsura, Yukako; Kazama, Yusuke; Nozawa, Masafumi (2024-07-31). "Sex chromosome cycle as a mechanism of stable sex determination". The Journal of Biochemistry 176 (2): 81–95. doi:10.1093/jb/mvae045. ISSN 0021-924X. PMID 38982631. PMC 11289310. https://academic.oup.com/jb/article/176/2/81/7709515.
- ↑ 14.0 14.1 14.2 14.3 14.4 "Genetic Mechanisms of Sex Determination". Nature Education 1 (1). 2008. http://www.nature.com/scitable/topicpage/genetic-mechanisms-of-sex-determination-314. Retrieved 8 December 2011.
- ↑ "DAX-1, an 'antitestis' gene". Cellular and Molecular Life Sciences 55 (6–7): 857–863. June 1999. doi:10.1007/PL00013201. PMID 10412368.
- ↑ 16.0 16.1 Chandra, H. S. (25 April 1999). "Another way of looking at the enigma of sex determination in Ellobius lutescens". Current Science 76 (8): 1072.
- ↑ "A SOX9 duplication and familial 46,XX developmental testicular disorder". The New England Journal of Medicine 364 (1): 91–93. January 2011. doi:10.1056/NEJMc1010311. PMID 21208124.
- ↑ "Autosomal XX sex reversal caused by duplication of SOX9". American Journal of Medical Genetics 87 (4): 349–353. December 1999. doi:10.1002/(SICI)1096-8628(19991203)87:4<349::AID-AJMG13>3.0.CO;2-N. PMID 10588843.
- ↑ "Somatic sex reprogramming of adult ovaries to testes by FOXL2 ablation". Cell 139 (6): 1130–1142. December 2009. doi:10.1016/j.cell.2009.11.021. PMID 20005806.
- ↑ "RNA binding protein sex-lethal (Sxl) and control of Drosophila sex determination and dosage compensation". Microbiology and Molecular Biology Reviews 67 (3): 343–59, table of contents. September 2003. doi:10.1128/MMBR.67.3.343-359.2003. PMID 12966139.
- ↑ 21.0 21.1 21.2 21.3 21.4 "A comparative view on sex determination in medaka". Mechanisms of Development 121 (7–8): 639–645. July 2004. doi:10.1016/j.mod.2004.03.001. PMID 15210173.
- ↑ Gruetzner, Frank; Ashley, Terry; Rowell, David M.; Marshall Graves, Jennifer A. (2006-04-01). "How did the platypus get its sex chromosome chain? A comparison of meiotic multiples and sex chromosomes in plants and animals". Chromosoma 115 (2): 75–88. doi:10.1007/s00412-005-0034-4. ISSN 1432-0886. PMID 16344965.
- ↑ Grützner, Frank; Rens, Willem; Tsend-Ayush, Enkhjargal; El-Mogharbel, Nisrine; O'Brien, Patricia C. M.; Jones, Russell C.; Ferguson-Smith, Malcolm A.; Marshall Graves, Jennifer A. (December 2004). "In the platypus a meiotic chain of ten sex chromosomes shares genes with the bird Z and mammal X chromosomes". Nature 432 (7019): 913–917. doi:10.1038/nature03021. ISSN 1476-4687. PMID 15502814. Bibcode: 2004Natur.432..913G. https://www.nature.com/articles/nature03021.
- ↑ "Genome analysis of the platypus reveals unique signatures of evolution". Nature 453 (7192): 175–183. May 2008. doi:10.1038/nature06936. PMID 18464734. Bibcode: 2008Natur.453..175W.
- ↑ "How did the platypus get its sex chromosome chain? A comparison of meiotic multiples and sex chromosomes in plants and animals". Chromosoma 115 (2): 75–88. April 2006. doi:10.1007/s00412-005-0034-4. PMID 16344965.
- ↑ "Additional copies of CBX2 in the genomes of males of mammals lacking SRY, the Amami spiny rat (Tokudaia osimensis) and the Tokunoshima spiny rat (Tokudaia tokunoshimensis)". Chromosome Research 19 (5): 635–644. July 2011. doi:10.1007/s10577-011-9223-6. PMID 21656076.
- ↑ 27.0 27.1 27.2 27.3 Sex wars: genes, bacteria, and biased sex ratios. Princeton University Press. 2003. p. 250. ISBN 978-0-691-00981-0. https://books.google.com/books?id=vDHOYPQ2mmYC&q=zo,+zww,+zzww+lepidoptera. Retrieved 4 November 2011.
- ↑ "tra-2 encodes a membrane protein and may mediate cell communication in the Caenorhabditis elegans sex determination pathway". Molecular Biology of the Cell 3 (4): 461–473. April 1992. doi:10.1091/mbc.3.4.461. PMID 1498366.
- ↑ "The avian Z-linked gene DMRT1 is required for male sex determination in the chicken". Nature 461 (7261): 267–271. September 2009. doi:10.1038/nature08298. PMID 19710650. Bibcode: 2009Natur.461..267S.
- ↑ "A single female-specific piRNA is the primary determiner of sex in the silkworm". Nature 509 (7502): 633–636. May 2014. doi:10.1038/nature13315. PMID 24828047. Bibcode: 2014Natur.509..633K.
- ↑ 31.0 31.1 "A new look at the evolution of avian sex chromosomes". Cytogenetic and Genome Research 117 (1–4): 103–109. 2007. doi:10.1159/000103170. PMID 17675850.
- ↑ "In the platypus a meiotic chain of ten sex chromosomes shares genes with the bird Z and mammal X chromosomes". Nature 432 (7019): 913–917. December 2004. doi:10.1038/nature03021. PMID 15502814. Bibcode: 2004Natur.432..913G.
- ↑ "Virgin births for giant lizards". BBC News. 20 December 2006. https://news.bbc.co.uk/2/hi/science/nature/6196225.stm.
- ↑ "Evolution of the Y Chromosome". http://www.learner.org/channel/courses/biology/textbook/gender/gender_4.html.
- ↑ "Sex chromosomes and sex determination in Lepidoptera". Sexual Development 1 (6): 332–346. 2007. doi:10.1159/000111765. PMID 18391545.
- ↑ "Genetic Mechanisms of Sex Determination - Learn Science at Scitable". http://www.nature.com/scitable/topicpage/genetic-mechanisms-of-sex-determination-314.
- ↑ Handbuch Der Zoologie / Handbook of Zoology. Walter de Gruyter. 1925. ISBN 978-3-11-016210-3. https://books.google.com/books?id=5w8FgSGuH34C&q=ZO+sex-determination+system+moth&pg=PA461.
- ↑ "Are all sex chromosomes created equal?". Trends in Genetics 27 (9): 350–357. September 2011. doi:10.1016/j.tig.2011.05.005. PMID 21962970.
- ↑ Renner, S. S.; Heinrichs, J.; Sousa, A. (2017). "The sex chromosomes of bryophytes: Recent insights, open questions, and reinvestigations of Frullania dilatata and Plagiochila asplenioides.". Journal of Systematics and Evolution 55 (4): 333–339. doi:10.1111/jse.12266. Bibcode: 2017JSyEv..55..333R.
- ↑ Idnurm, Alexander; Hood, Michael E.; Johannesson, Hanna; Giraud, Tatiana (2015-12-01). "Contrasted patterns in mating-type chromosomes in fungi: Hotspots versus coldspots of recombination". Fungal Biology Reviews. Special Issue: Fungal sex and mushrooms – A credit to Lorna Casselton 29 (3): 220–229. doi:10.1016/j.fbr.2015.06.001. ISSN 1749-4613. PMID 26688691. Bibcode: 2015FunBR..29..220I.
- ↑ "The gene csd is the primary signal for sexual development in the honeybee and encodes an SR-type protein". Cell 114 (4): 419–429. August 2003. doi:10.1016/S0092-8674(03)00606-8. PMID 12941271.
- ↑ "Duplication and concerted evolution in a master sex determiner under balancing selection". Proceedings. Biological Sciences 280 (1758). May 2013. doi:10.1098/rspb.2012.2968. PMID 23466984.
- ↑ "The doublesex gene integrates multi-locus complementary sex determination signals in the Japanese ant, Vollenhovia emeryi". Insect Biochemistry and Molecular Biology 94: 42–49. March 2018. doi:10.1016/j.ibmb.2018.01.006. PMID 29408414. Bibcode: 2018IBMB...94...42M.
- ↑ "Single locus complementary sex determination in Hymenoptera: an "unintelligent" design?". Frontiers in Zoology 3 (1). January 2006. doi:10.1186/1742-9994-3-1. PMID 16393347.
- ↑ Moore, Emily C.; Roberts, Reade B. (June 2013). "Polygenic sex determination". Current Biology 23 (12): R510–R512. doi:10.1016/j.cub.2013.04.004. PMID 23787041. Bibcode: 2013CBio...23.R510M. https://linkinghub.elsevier.com/retrieve/pii/S0960982213004120.
- ↑ Liew, Woei Chang; Bartfai, Richard; Lim, Zijie; Sreenivasan, Rajini; Siegfried, Kellee R.; Orban, Laszlo (2012-04-10). Alsina, Berta. ed. "Polygenic Sex Determination System in Zebrafish". PLOS ONE 7 (4). doi:10.1371/journal.pone.0034397. ISSN 1932-6203. PMID 22506019. Bibcode: 2012PLoSO...734397L.
- ↑ Liew, Woei Chang; Orbán, László (March 2014). "Zebrafish sex: a complicated affair". Briefings in Functional Genomics 13 (2): 172–187. doi:10.1093/bfgp/elt041. ISSN 2041-2649. PMID 24148942.
- ↑ Pan, Qiaowei; Darras, Hugo; Keller, Laurent (2024-05-31). "LncRNA gene ANTSR coordinates complementary sex determination in the Argentine ant". Science Advances 10 (22). doi:10.1126/sciadv.adp1532. ISSN 2375-2548. PMID 38820161.
- ↑ Yu, Chuanxin; Hodapp, Dean; Moog, Safira; Dupont, Simon; Darrouzet, Eric; Keller Valsecchi, Claudia Isabelle; Colgan, Thomas Joseph; Pan, Qiaowei et al. (2026-01-13). "Deep evolutionary conservation of a sex-determining locus without sequence homology". Proceedings of the National Academy of Sciences 123 (2). doi:10.1073/pnas.2522417123. PMID 41490485. PMC 12799146. https://www.pnas.org/doi/10.1073/pnas.2522417123.
- ↑ 50.0 50.1 Wright, Alison E.; Dean, Rebecca; Zimmer, Fabian; Mank, Judith E. (2016-07-04). "How to make a sex chromosome". Nature Communications 7 (1). doi:10.1038/ncomms12087. ISSN 2041-1723. PMID 27373494. Bibcode: 2016NatCo...712087W.
- ↑ Stöck, Matthias; Horn, Agnès; Grossen, Christine; Lindtke, Dorothea; Sermier, Roberto; Betto-Colliard, Caroline; Dufresnes, Christophe; Bonjour, Emmanuel et al. (2011-05-17). Rice, William R.. ed. "Ever-Young Sex Chromosomes in European Tree Frogs". PLOS Biology 9 (5). doi:10.1371/journal.pbio.1001062. ISSN 1545-7885. PMID 21629756.
- ↑ Furman, Benjamin L S; Metzger, David C H; Darolti, Iulia; Wright, Alison E; Sandkam, Benjamin A; Almeida, Pedro; Shu, Jacelyn J; Mank, Judith E (2020-06-01). Fraser, Bonnie. ed. "Sex Chromosome Evolution: So Many Exceptions to the Rules". Genome Biology and Evolution 12 (6): 750–763. doi:10.1093/gbe/evaa081. ISSN 1759-6653. PMID 32315410. PMC 7268786. https://academic.oup.com/gbe/article/12/6/750/5823304.
- ↑ Martin, Jon; Lee, B. T. O. (September 1984). "A phylogenetic study of sex determiner location in a group of Australasian Chironomus species (Diptera, Chironomidae)". Chromosoma 90 (3): 190–197. doi:10.1007/BF00292396. ISSN 0009-5915. https://link.springer.com/10.1007/BF00292396.
- ↑ Martin, Jon; Kuvangkadilok, Chaliow; Peart, Dianne H.; Lee, Barry T. O. (June 1980). "Multiple sex determining regions in a group of related Chironomus species (Diptera:Chironomidae)". Heredity 44 (3): 367–382. doi:10.1038/hdy.1980.34. ISSN 1365-2540. Bibcode: 1980Hered..44..367M. https://www.nature.com/articles/hdy198034.
- ↑ Green, David M. (1988-09-01). "Cytogenetics of the endemic New Zealand frog, Leiopelma hochstetteri: extraordinary supernumerary chromosome variation and a unique sex-chromosome system". Chromosoma 97 (1): 55–70. doi:10.1007/BF00331795. ISSN 1432-0886.
- ↑ Uno, Yoshinobu; Nishida, Chizuko; Oshima, Yuki; Yokoyama, Satoshi; Miura, Ikuo; Matsuda, Yoichi; Nakamura, Masahisa (June 2008). "Comparative chromosome mapping of sex-linked genes and identification of sex chromosomal rearrangements in the Japanese wrinkled frog (Rana rugosa, Ranidae) with ZW and XY sex chromosome systems". Chromosome Research 16 (4): 637–647. doi:10.1007/s10577-008-1217-7. ISSN 0967-3849. PMID 18484182. https://link.springer.com/10.1007/s10577-008-1217-7.
- ↑ 57.0 57.1 Graves, Jennifer A. Marshall (2008-12-01). "Weird Animal Genomes and the Evolution of Vertebrate Sex and Sex Chromosomes". Annual Review of Genetics 42 (1): 565–586. doi:10.1146/annurev.genet.42.110807.091714. ISSN 0066-4197. PMID 18983263.
- ↑ Nguyen, Dung Ho My; Panthum, Thitipong; Ponjarat, Jatupong; Laopichienpong, Nararat; Kraichak, Ekaphan; Singchat, Worapong; Ahmad, Syed Farhan; Muangmai, Narongrit et al. (2021-01-05). "An Investigation of ZZ/ZW and XX/XY Sex Determination Systems in North African Catfish (Clarias gariepinus, Burchell, 1822)". Frontiers in Genetics 11. doi:10.3389/fgene.2020.562856. ISSN 1664-8021. PMID 33584785.
- ↑ Sember, Alexandr; Nguyen, Petr; Perez, Manolo F.; Altmanová, Marie; Ráb, Petr; Cioffi, Marcelo de Bello (2021-09-13). "Multiple sex chromosomes in teleost fishes from a cytogenetic perspective: state of the art and future challenges". Philosophical Transactions of the Royal Society B: Biological Sciences 376 (1833). doi:10.1098/rstb.2020.0098. ISSN 0962-8436. PMID 34304595.
- ↑ Traut, W.; Sahara, K.; Marec, F. (2008-01-18). "Sex Chromosomes and Sex Determination in Lepidoptera". Sexual Development 1 (6): 332–346. doi:10.1159/000111765. ISSN 1661-5425. PMID 18391545.
- ↑ Kallman, Klaus D. (1984), Turner, Bruce J., ed., "A New Look at Sex Determination in Poeciliid Fishes", Evolutionary Genetics of Fishes (Boston, MA: Springer US): pp. 95–171, doi:10.1007/978-1-4684-4652-4_3, ISBN 978-1-4684-4652-4
- ↑ "A comparative view on sex determination in medaka". Mechanisms of Development 121 (7–8): 639–645. July 2004. doi:10.1016/j.mod.2004.03.001. PMID 15210173.
- ↑ Schmid, M.; Steinlein, C.; Feichtinger, W. (March 1992). "Chromosome banding in amphibia: XVII. First demonstration of multiple sex chromosomes in amphibians: Eleutherodactylus maussi (Anura, Leptodactylidae)". Chromosoma 101 (5–6): 284–292. doi:10.1007/BF00346007. ISSN 0009-5915. PMID 1576881. https://link.springer.com/10.1007/BF00346007.
- ↑ Schmid, M.; Feichtinger, W.; Steinlein, C.; Haaf, T.; Schartl, M.; Visbal García, R.; Manzanilla Pupo, J.; Fernández Badillo, A. (2003-08-14). "Chromosome banding in Amphibia: XXVI. Coexistence of homomorphic XY sex chromosomes and a derived Y-autosome translocation in Eleutherodactylus maussi (Anura, Leptodactylidae)". Cytogenetics and Cell Genetics 99 (1–4): 330–343. doi:10.1159/000071612. ISSN 0301-0171. PMID 12900583.
- ↑ 65.0 65.1 Zhu, Zexian; Younas, Lubna; Zhou, Qi (2024-07-18). "Evolution and regulation of animal sex chromosomes". Nature Reviews Genetics 26 (1): 59–74. doi:10.1038/s41576-024-00757-3. ISSN 1471-0064. PMID 39026082. https://www.nature.com/articles/s41576-024-00757-3.
- ↑ Lahn, Bruce T.; Page, David C. (1999-10-29). "Four Evolutionary Strata on the Human X Chromosome". Science 286 (5441): 964–967. doi:10.1126/science.286.5441.964. ISSN 0036-8075. PMID 10542153. https://www.science.org/doi/10.1126/science.286.5441.964.
- ↑ Lemaitre, Claire; Braga, Marilia D. V.; Gautier, Christian; Sagot, Marie-France; Tannier, Eric; Marais, Gabriel A. B. (2009-01-01). "Footprints of Inversions at Present and Past Pseudoautosomal Boundaries in Human Sex Chromosomes". Genome Biology and Evolution 1: 56–66. doi:10.1093/gbe/evp006. ISSN 1759-6653. PMID 20333177.
- ↑ Zhou, Yang; Zhan, Xiaoyu; Jin, Jiazheng; Zhou, Long; Bergman, Juraj; Li, Xuemei; Rousselle, Marjolaine Marie C.; Belles, Meritxell Riera et al. (July 2023). "Eighty million years of rapid evolution of the primate Y chromosome". Nature Ecology & Evolution 7 (7): 1114–1130. doi:10.1038/s41559-022-01974-x. ISSN 2397-334X. PMID 37268856. Bibcode: 2023NatEE...7.1114Z. https://www.nature.com/articles/s41559-022-01974-x.
- ↑ 69.0 69.1 Solari, Alberto J. (1994). Sex chromosomes and sex determination in vertebrates. Boca Raton: CRC Press. ISBN 978-0-8493-4571-5.
- ↑ Zhou, Qi; Wang, Jun; Huang, Ling; Nie, Wenhui; Wang, Jinhuan; Liu, Yan; Zhao, Xiangyi; Yang, Fengtang et al. (2008-06-14). "Neo-sex chromosomes in the black muntjac recapitulate incipient evolution of mammalian sex chromosomes". Genome Biology 9 (6): R98. doi:10.1186/gb-2008-9-6-r98. ISSN 1474-760X. PMID 18554412.
- ↑ Pala, I.; Naurin, S.; Stervander, M.; Hasselquist, D.; Bensch, S.; Hansson, B. (March 2012). "Evidence of a neo-sex chromosome in birds". Heredity 108 (3): 264–272. doi:10.1038/hdy.2011.70. ISSN 1365-2540. PMID 21897438. Bibcode: 2012Hered.108..264P.
- ↑ Vicoso, Beatriz (December 2019). "Molecular and evolutionary dynamics of animal sex-chromosome turnover". Nature Ecology & Evolution 3 (12): 1632–1641. doi:10.1038/s41559-019-1050-8. ISSN 2397-334X. PMID 31768022. Bibcode: 2019NatEE...3.1632V. https://www.nature.com/articles/s41559-019-1050-8.
- ↑ Palmer, Daniela H.; Rogers, Thea F.; Dean, Rebecca; Wright, Alison E. (November 2019). "How to identify sex chromosomes and their turnover". Molecular Ecology 28 (21): 4709–4724. doi:10.1111/mec.15245. ISSN 0962-1083. PMID 31538682. Bibcode: 2019MolEc..28.4709P.
- ↑ van Doorn, G Sander; Kirkpatrick, Mark (2010-10-01). "Transitions Between Male and Female Heterogamety Caused by Sex-Antagonistic Selection". Genetics 186 (2): 629–645. doi:10.1534/genetics.110.118596. ISSN 1943-2631. PMID 20628036.
- ↑ Perrin, Nicolas (December 2009). "Sex Reversal: A Fountain of Youth for Sex Chromosomes?". Evolution 63 (12): 3043–3049. doi:10.1111/j.1558-5646.2009.00837.x. PMID 19744117. https://academic.oup.com/evolut/article/63/12/3043/6881025.
- ↑ Nagler, J J; Bouma, J; Thorgaard, G H; Dauble, D D (January 2001). "High incidence of a male-specific genetic marker in phenotypic female chinook salmon from the Columbia River.". Environmental Health Perspectives 109 (1): 67–69. doi:10.1289/ehp.0110967. ISSN 0091-6765. PMID 11171527. Bibcode: 2001EnvHP.109...67N.
- ↑ Rodrigues, Nicolas; Studer, Tania; Dufresnes, Christophe; Perrin, Nicolas (2018-04-01). "Sex-Chromosome Recombination in Common Frogs Brings Water to the Fountain-of-Youth". Molecular Biology and Evolution 35 (4): 942–948. doi:10.1093/molbev/msy008. ISSN 0737-4038. PMID 29394416. https://academic.oup.com/mbe/article/35/4/942/4823133.
- ↑ "Temperature-dependent sex ratio in a bird". Biology Letters 1 (1): 31–33. March 2005. doi:10.1098/rsbl.2004.0247. PMID 17148121.
- ↑ 79.0 79.1 "Expression profiles of Dax1, Dmrt1, and Sox9 during temperature sex determination in gonads of the sea turtle Lepidochelys olivacea". General and Comparative Endocrinology 129 (1): 20–26. October 2002. doi:10.1016/s0016-6480(02)00511-7. PMID 12409092.
- ↑ "Sex Determination in Reptiles". The Quarterly Review of Biology 55 (1): 3–21. March 1980. doi:10.1086/411613.
- ↑ 81.0 81.1 "Nest-site philopatry and the evolution of temperature-dependent sex determination". Evolutionary Ecology Research 3: 779–794. 2001. http://www.public.iastate.edu/~fjanzen/pdf/01EvolEcolRes.pdf. Retrieved 7 December 2011.
- ↑ "Exploring the evolution of environmental sex determination, especially in reptiles". Journal of Evolutionary Biology 19 (6): 1775–1784. November 2006. doi:10.1111/j.1420-9101.2006.01138.x. PMID 17040374.
- ↑ Developmental biology (8th. ed.). Sunderland, Mass.: Sinauer Associates, Inc. Publishers. 2006. pp. 550–553. ISBN 978-0-87893-250-4. https://archive.org/details/developmentalbio00gilb_292.
- ↑ Casas, L., Saborido-Rey, F., Ryu, T., Michell, C., Ravasi, T., & Irigoien, X. (2016). Sex Change in Clownfish: Molecular Insights from Transcriptome Analysis. Scientific Reports, 6(1). https://doi.org/10.1038/srep35461
- ↑ "Parthenogenesis in Komodo dragons". Nature 444 (7122): 1021–1022. December 2006. doi:10.1038/4441021a. PMID 17183308. Bibcode: 2006Natur.444.1021W.
- ↑ Stevens, Lori; Giordano, Rosanna; Fialho, Roberto F. (November 2001). "Male-Killing, Nematode Infections, Bacteriophage Infection, and Virulence of Cytoplasmic Bacteria in the Genus Wolbachia" (in en). Annual Review of Ecology and Systematics 32 (1): 519–545. doi:10.1146/annurev.ecolsys.32.081501.114132. ISSN 0066-4162. Bibcode: 2001AnRES..32..519S. https://www.annualreviews.org/doi/10.1146/annurev.ecolsys.32.081501.114132.
- ↑ 87.0 87.1 87.2 87.3 "Sex determination: why so many ways of doing it?". PLOS Biology 12 (7). July 2014. doi:10.1371/journal.pbio.1001899. PMID 24983465.
- ↑ David, Patrice; Degletagne, Cyril; Saclier, Nathanaëlle; Jennan, Aurel; Jarne, Philippe; Plénet, Sandrine; Konecny, Lara; François, Clémentine et al. (May 2022). "Extreme mitochondrial DNA divergence underlies genetic conflict over sex determination" (in en). Current Biology 32 (10): 2325–2333.e6. doi:10.1016/j.cub.2022.04.014. PMID 35483362. Bibcode: 2022CBio...32E2325D. https://linkinghub.elsevier.com/retrieve/pii/S0960982222005838.
- ↑ Herbette, Marion; Ross, Laura (2023). "Paternal genome elimination: patterns and mechanisms of drive and silencing" (in en). Current Opinion in Genetics & Development 81. doi:10.1016/j.gde.2023.102065. PMID 37413739.
- ↑ Baird, Robert B.; Mongue, Andrew J.; Ross, Laura (2023). "Why put all your eggs in one basket? Evolutionary perspectives on the origins of monogenic reproduction" (in en). Heredity 131 (2): 87–95. doi:10.1038/s41437-023-00632-7. ISSN 0018-067X. PMID 37328587. Bibcode: 2023Hered.131...87B.
- ↑ "Gamete competition, gamete limitation, and the evolution of the two sexes". Molecular Human Reproduction 20 (12): 1161–1168. December 2014. doi:10.1093/molehr/gau068. PMID 25323972.
- ↑ Cannabis: Evolution and Ethnobotany. Univ of California Press. 2016-06-28. p. 359. ISBN 978-0-520-29248-2. https://books.google.com/books?id=bs4hEAAAQBAJ&q=XY+sex+determination+in+plants&pg=PA359.
- ↑ "XY and ZW: is meiotic sex chromosome inactivation the rule in evolution?". PLOS Genetics 5 (5). May 2009. doi:10.1371/journal.pgen.1000493. PMID 19461890.
- ↑ 94.0 94.1 94.2 94.3 "Multiple independent origins of sex chromosomes in amniotes". Proceedings of the National Academy of Sciences of the United States of America 103 (48): 18031–18032. November 2006. doi:10.1073/pnas.0608879103. PMID 17116892. Bibcode: 2006PNAS..10318031V.
- ↑ "Bird-like sex chromosomes of platypus imply recent origin of mammal sex chromosomes". Genome Research 18 (6): 965–973. June 2008. doi:10.1101/gr.7101908. PMID 18463302.
- ↑ "Human Y chromosome, sex determination, and spermatogenesis- a feminist view". Biology of Reproduction 63 (3): 667–676. September 2000. doi:10.1095/biolreprod63.3.667b. PMID 10952906.
- ↑ "Relationships between vertebrate ZW and XY sex chromosome systems". Current Biology 16 (17): R736–R743. September 2006. doi:10.1016/j.cub.2006.08.021. PMID 16950100. Bibcode: 2006CBio...16.R736E.
- ↑ "Evolutionary transitions between mechanisms of sex determination in vertebrates". Biology Letters 7 (3): 443–448. June 2011. doi:10.1098/rsbl.2010.1126. PMID 21212104.
- ↑ 99.0 99.1 "Sex chromosome specialization and degeneration in mammals". Cell 124 (5): 901–914. March 2006. doi:10.1016/j.cell.2006.02.024. PMID 16530039.
- ↑ "The evolution of the sex chromosomes: Step by step" (Press release). University of Chicago Medical Center. 28 October 1999. Retrieved 23 October 2011.
- ↑ "The organization and evolution of the human Y chromosome". Genome Biology 4 (9): 226. 14 August 2003. doi:10.1186/gb-2003-4-9-226. PMID 12952526.
- ↑ "The degenerate Y chromosome--can conversion save it?". Reproduction, Fertility, and Development 16 (5): 527–534. 22 July 2004. doi:10.1071/RD03096. PMID 15367368.
- ↑ "Strict evolutionary conservation followed rapid gene loss on human and rhesus Y chromosomes". Nature 483 (7387): 82–86. February 2012. doi:10.1038/nature10843. PMID 22367542. Bibcode: 2012Natur.483...82H.
- ↑ Mutation-Driven Evolution. OUP Oxford. 2013-05-02. p. 168. ISBN 978-0-19-163781-0. https://books.google.com/books?id=sJtoAgAAQBAJ&q=sexual+systems+can+evolve&pg=PA168.
Further reading
- The Evolution of Sex Determination. Oxford University Press. 2014. ISBN 978-0-19-163140-5. https://books.google.com/books?id=d4cLBAAAQBAJ.
- Lahn, Bruce T. (1999-10-29). "Four Evolutionary Strata on the Human X Chromosome". Science 286 (5441): 964–967. doi:10.1126/science.286.5441.964. ISSN 0036-8075. PMID 10542153. https://www.science.org/doi/10.1126/science.286.5441.964.
 |
