Biology:Notch signaling pathway
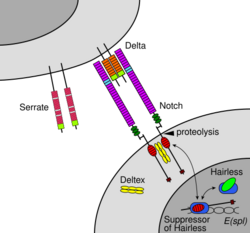

The Notch signaling pathway is a highly conserved cell signaling system present in most animals.[1] Mammals possess four different notch receptors, referred to as NOTCH1, NOTCH2, NOTCH3, and NOTCH4.[2] The notch receptor is a single-pass transmembrane receptor protein. It is a hetero-oligomer composed of a large extracellular portion, which associates in a calcium-dependent, non-covalent interaction with a smaller piece of the notch protein composed of a short extracellular region, a single transmembrane-pass, and a small intracellular region.[3]
Notch signaling promotes proliferative signaling during neurogenesis, and its activity is inhibited by Numb to promote neural differentiation. It plays a major role in the regulation of embryonic development.
Notch signaling is dysregulated in many cancers, and faulty notch signaling is implicated in many diseases, including T-cell acute lymphoblastic leukemia (T-ALL),[4] cerebral autosomal-dominant arteriopathy with sub-cortical infarcts and leukoencephalopathy (CADASIL), multiple sclerosis, Tetralogy of Fallot, and Alagille syndrome. Inhibition of notch signaling inhibits the proliferation of T-cell acute lymphoblastic leukemia in both cultured cells and a mouse model.[5][6]
Discovery
In 1914, John S. Dexter noticed the appearance of a notch in the wings of the fruit fly Drosophila ampelophila. The alleles of the gene were identified in 1917 by American evolutionary biologist Thomas Hunt Morgan.[7][8] Its molecular analysis and sequencing was independently undertaken in the 1980s by Spyros Artavanis-Tsakonas and Michael W. Young.[9][10] Alleles of the two C. elegans Notch genes were identified based on developmental phenotypes: lin-12[11] and glp-1.[12][13] The cloning and partial sequence of lin-12 was reported at the same time as Drosophila Notch by Iva Greenwald.[14]
Mechanism
The Notch protein spans the cell membrane, with part of it inside and part outside. Ligand proteins binding to the extracellular domain induce proteolytic cleavage and release of the intracellular domain, which enters the cell nucleus to modify gene expression.[15]
The cleavage model was first proposed in 1993 based on work done with Drosophila Notch and C. elegans lin-12,[16][17] informed by the first oncogenic mutation affecting a human Notch gene.[18] Compelling evidence for this model was provided in 1998 by in vivo analysis in Drosophila by Gary Struhl[19] and in cell culture by Raphael Kopan.[20] Although this model was initially disputed,[1] the evidence in favor of the model was irrefutable by 2001.[21][22]
The receptor is normally triggered via direct cell-to-cell contact, in which the transmembrane proteins of the cells in direct contact form the ligands that bind the notch receptor. The Notch binding allows groups of cells to organize themselves such that, if one cell expresses a given trait, this may be switched off in neighbouring cells by the intercellular notch signal. In this way, groups of cells influence one another to make large structures. Thus, lateral inhibition mechanisms are key to Notch signaling. lin-12 and Notch mediate binary cell fate decisions, and lateral inhibition involves feedback mechanisms to amplify initial differences.[21]
The Notch cascade consists of Notch and Notch ligands, as well as intracellular proteins transmitting the notch signal to the cell's nucleus. The Notch/Lin-12/Glp-1 receptor family[23] was found to be involved in the specification of cell fates during development in Drosophila and C. elegans.[24]
The intracellular domain of Notch forms a complex with CBF1 and Mastermind to activate transcription of target genes. The structure of the complex has been determined.[25][26]
Pathway
Maturation of the notch receptor involves cleavage at the prospective extracellular side during intracellular trafficking in the Golgi complex.[27] This results in a bipartite protein, composed of a large extracellular domain linked to the smaller transmembrane and intracellular domain. Binding of ligand promotes two proteolytic processing events; as a result of proteolysis, the intracellular domain is liberated and can enter the nucleus to engage other DNA-binding proteins and regulate gene expression.
Notch and most of its ligands are transmembrane proteins, so the cells expressing the ligands typically must be adjacent to the notch expressing cell for signaling to occur. The notch ligands are also single-pass transmembrane proteins and are members of the DSL (Delta/Serrate/LAG-2) family of proteins. In Drosophila melanogaster (the fruit fly), there are two ligands named Delta and Serrate. In mammals, the corresponding names are Delta-like and Jagged. In mammals there are multiple Delta-like and Jagged ligands, as well as possibly a variety of other ligands, such as F3/contactin.[28]
The notch extracellular domain is composed primarily of small cystine-rich motifs called EGF-like repeats.[29]
Notch 1, for example, has 36 of these repeats. Each EGF-like repeat is composed of approximately 40 amino acids, and its structure is defined largely by six conserved cysteine residues that form three conserved disulfide bonds. Each EGF-like repeat can be modified by O-linked glycans at specific sites.[30] An O-glucose sugar may be added between the first and second conserved cysteines, and an O-fucose may be added between the second and third conserved cysteines. These sugars are added by an as-yet-unidentified O-glucosyltransferase (except for Rumi), and GDP-fucose Protein O-fucosyltransferase 1 (POFUT1), respectively. The addition of O-fucose by POFUT1 is absolutely necessary for notch function, and, without the enzyme to add O-fucose, all notch proteins fail to function properly. As yet, the manner by which the glycosylation of notch affects function is not completely understood.
The O-glucose on notch can be further elongated to a trisaccharide with the addition of two xylose sugars by xylosyltransferases, and the O-fucose can be elongated to a tetrasaccharide by the ordered addition of an N-acetylglucosamine (GlcNAc) sugar by an N-Acetylglucosaminyltransferase called Fringe, the addition of a galactose by a galactosyltransferase, and the addition of a sialic acid by a sialyltransferase.[31]
To add another level of complexity, in mammals there are three Fringe GlcNAc-transferases, named lunatic fringe, manic fringe, and radical fringe. These enzymes are responsible for something called a "fringe effect" on notch signaling.[32] If Fringe adds a GlcNAc to the O-fucose sugar then the subsequent addition of a galactose and sialic acid will occur. In the presence of this tetrasaccharide, notch signals strongly when it interacts with the Delta ligand, but has markedly inhibited signaling when interacting with the Jagged ligand.[33] The means by which this addition of sugar inhibits signaling through one ligand, and potentiates signaling through another is not clearly understood.
Once the notch extracellular domain interacts with a ligand, an ADAM-family metalloprotease called ADAM10, cleaves the notch protein just outside the membrane.[34] This releases the extracellular portion of notch (NECD), which continues to interact with the ligand. The ligand plus the notch extracellular domain is then endocytosed by the ligand-expressing cell. There may be signaling effects in the ligand-expressing cell after endocytosis; this part of notch signaling is a topic of active research. After this first cleavage, an enzyme called γ-secretase (which is implicated in Alzheimer's disease) cleaves the remaining part of the notch protein just inside the inner leaflet of the cell membrane of the notch-expressing cell. This releases the intracellular domain of the notch protein (NICD), which then moves to the nucleus, where it can regulate gene expression by activating the transcription factor CSL. It was originally thought that these CSL proteins suppressed Notch target transcription. However, further research showed that, when the intracellular domain binds to the complex, it switches from a repressor to an activator of transcription.[35] Other proteins also participate in the intracellular portion of the notch signaling cascade.[36]
Ligand interactions

Notch signaling is initiated when Notch receptors on the cell surface engage ligands presented in trans on opposing cells. Despite the expansive size of the Notch extracellular domain, it has been demonstrated that EGF domains 11 and 12 are the critical determinants for interactions with Delta.[37] Additional studies have implicated regions outside of Notch EGF11-12 in ligand binding. For example, Notch EGF domain 8 plays a role in selective recognition of Serrate/Jagged[38] and EGF domains 6-15 are required for maximal signaling upon ligand stimulation.[39] A crystal structure of the interacting regions of Notch1 and Delta-like 4 (Dll4) provided a molecular-level visualization of Notch-ligand interactions, and revealed that the N-terminal MNNL (or C2) and DSL domains of ligands bind to Notch EGF domains 12 and 11, respectively.[40] The Notch1-Dll4 structure also illuminated a direct role for Notch O-linked fucose and glucose moieties in ligand recognition, and rationalized a structural mechanism for the glycan-mediated tuning of Notch signaling.[40]
Synthetic Notch signaling
It is possible to engineer synthetic Notch receptors by replacing the extracellular receptor and intracellular transcriptional domains with other domains of choice. This allows researchers to select which ligands are detected, and which genes are upregulated in response. Using this technology, cells can report or change their behavior in response to contact with user-specified signals, facilitating new avenues of both basic and applied research into cell-cell signaling.[41] Notably, this system allows multiple synthetic pathways to be engineered into a cell in parallel.[42][43]
Function
The Notch signaling pathway is important for cell-cell communication, which involves gene regulation mechanisms that control multiple cell differentiation processes during embryonic and adult life. Notch signaling also has a role in the following processes:
- neuronal function and development[44][45][46][47]
- stabilization of arterial endothelial fate and angiogenesis[48]
- regulation of crucial cell communication events between endocardium and myocardium during both the formation of the valve primordial and ventricular development and differentiation[49]
- cardiac valve homeostasis, as well as implications in other human disorders involving the cardiovascular system[50]
- timely cell lineage specification of both endocrine and exocrine pancreas[51]
- influencing of binary fate decisions of cells that must choose between the secretory and absorptive lineages in the gut[52]
- expansion of the hematopoietic stem cell compartment during bone development and participation in commitment to the osteoblastic lineage, suggesting a potential therapeutic role for notch in bone regeneration and osteoporosis[53]
- expansion of the hemogenic endothelial cells along with signaling axis involving Hedgehog signaling and Scl[54]
- T cell lineage commitment from common lymphoid precursor [55]
- regulation of cell-fate decision in mammary glands at several distinct development stages[56]
- possibly some non-nuclear mechanisms, such as control of the actin cytoskeleton through the tyrosine kinase Abl[28]
- Regulation of the mitotic/meiotic decision in the C. elegans germline[12]
- development of alveoli in the lung.[57]
It has also been found that Rex1 has inhibitory effects on the expression of notch in mesenchymal stem cells, preventing differentiation.[58]
Role in embryogenesis
The Notch signaling pathway plays an important role in cell-cell communication, and further regulates embryonic development.
Embryo polarity
Notch signaling is required in the regulation of polarity. For example, mutation experiments have shown that loss of Notch signaling causes abnormal anterior-posterior polarity in somites.[59] Also, Notch signaling is required during left-right asymmetry determination in vertebrates.[60]
Early studies in the nematode model organism C. elegans indicate that Notch signaling has a major role in the induction of mesoderm and cell fate determination.[12] As mentioned previously, C. elegans has two genes that encode for partially functionally redundant Notch homologs, glp-1 and lin-12.[61] During C. elegans, GLP-1, the C. elegans Notch homolog, interacts with APX-1, the C. elegans Delta homolog. This signaling between particular blastomeres induces differentiation of cell fates and establishes the dorsal-ventral axis.[62]
Role in somitogenesis
Notch signaling is central to somitogenesis. In 1995, Notch1 was shown to be important for coordinating the segmentation of somites in mice.[63] Further studies identified the role of Notch signaling in the segmentation clock. These studies hypothesized that the primary function of Notch signaling does not act on an individual cell, but coordinates cell clocks and keep them synchronized. This hypothesis explained the role of Notch signaling in the development of segmentation and has been supported by experiments in mice and zebrafish.[64][65][66] Experiments with Delta1 mutant mice that show abnormal somitogenesis with loss of anterior/posterior polarity suggest that Notch signaling is also necessary for the maintenance of somite borders.[63]
During somitogenesis, a molecular oscillator in paraxial mesoderm cells dictates the precise rate of somite formation. A clock and wavefront model has been proposed in order to spatially determine the location and boundaries between somites. This process is highly regulated as somites must have the correct size and spacing in order to avoid malformations within the axial skeleton that may potentially lead to spondylocostal dysostosis. Several key components of the Notch signaling pathway help coordinate key steps in this process. In mice, mutations in Notch1, Dll1 or Dll3, Lfng, or Hes7 result in abnormal somite formation. Similarly, in humans, the following mutations have been seen to lead to development of spondylocostal dysostosis: DLL3, LFNG, or HES7.[67]
Role in epidermal differentiation
Notch signaling is known to occur inside ciliated, differentiating cells found in the first epidermal layers during early skin development.[68] Furthermore, it has found that presenilin-2 works in conjunction with ARF4 to regulate Notch signaling during this development.[69] However, it remains to be determined whether gamma-secretase has a direct or indirect role in modulating Notch signaling.
Role in central nervous system development and function

Early findings on Notch signaling in central nervous system (CNS) development were performed mainly in Drosophila with mutagenesis experiments. For example, the finding that an embryonic lethal phenotype in Drosophila was associated with Notch dysfunction[70] indicated that Notch mutations can lead to the failure of neural and Epidermal cell segregation in early Drosophila embryos. In the past decade, advances in mutation and knockout techniques allowed research on the Notch signaling pathway in mammalian models, especially rodents.
The Notch signaling pathway was found to be critical mainly for neural progenitor cell (NPC) maintenance and self-renewal. In recent years, other functions of the Notch pathway have also been found, including glial cell specification,[71][72] neurites development,[73] as well as learning and memory.[74]
Neuron cell differentiation
The Notch pathway is essential for maintaining NPCs in the developing brain. Activation of the pathway is sufficient to maintain NPCs in a proliferating state, whereas loss-of-function mutations in the critical components of the pathway cause precocious neuronal differentiation and NPC depletion.[45] Modulators of the Notch signal, e.g., the Numb protein are able to antagonize Notch effects, resulting in the halting of cell cycle and the differentiation of NPCs.[75][76] Conversely, the fibroblast growth factor pathway promotes Notch signaling to keep stem cells of the cerebral cortex in the proliferative state, amounting to a mechanism regulating cortical surface area growth and, potentially, gyrification.[77][78] In this way, Notch signaling controls NPC self-renewal as well as cell fate specification.
A non-canonical branch of the Notch signaling pathway that involves the phosphorylation of STAT3 on the serine residue at amino acid position 727 and subsequent Hes3 expression increase (STAT3-Ser/Hes3 Signaling Axis) has been shown to regulate the number of NPCs in culture and in the adult rodent brain.[79]
In adult rodents and in cell culture, Notch3 promotes neuronal differentiation, having a role opposite to Notch1/2.[80] This indicates that individual Notch receptors can have divergent functions, depending on cellular context.
Neurite development
In vitro studies show that Notch can influence neurite development.[73] In vivo, deletion of the Notch signaling modulator, Numb, disrupts neuronal maturation in the developing cerebellum,[81] whereas deletion of Numb disrupts axonal arborization in sensory ganglia.[82] Although the mechanism underlying this phenomenon is not clear, together these findings suggest Notch signaling might be crucial in neuronal maturation.
Gliogenesis
In gliogenesis, Notch appears to have an instructive role that can directly promote the differentiation of many glial cell subtypes.[71][72] For example, activation of Notch signaling in the retina favors the generation of Muller glia cells at the expense of neurons, whereas reduced Notch signaling induces production of ganglion cells, causing a reduction in the number of Muller glia.[45]
Adult brain function
Apart from its role in development, evidence shows that Notch signaling is also involved in neuronal apoptosis, neurite retraction, and neurodegeneration of ischemic stroke in the brain[83] In addition to developmental functions, Notch proteins and ligands are expressed in cells of the adult nervous system,[84] suggesting a role in CNS plasticity throughout life. Adult mice heterozygous for mutations in either Notch1 or Cbf1 have deficits in spatial learning and memory.[74] Similar results are seen in experiments with presenilins1 and 2, which mediate the Notch intramembranous cleavage. To be specific, conditional deletion of presenilins at 3 weeks after birth in excitatory neurons causes learning and memory deficits, neuronal dysfunction, and gradual neurodegeneration.[85] Several gamma secretase inhibitors that underwent human clinical trials in Alzheimer's disease and MCI patients resulted in statistically significant worsening of cognition relative to controls, which is thought to be due to its incidental effect on Notch signalling.[86]
Role in cardiovascular development
The Notch signaling pathway is a critical component of cardiovascular formation and morphogenesis in both development and disease. It is required for the selection of endothelial tip and stalk cells during sprouting angiogenesis.[87]
Cardiac development
Notch signal pathway plays a crucial role in at least three cardiac development processes: Atrioventricular canal development, myocardial development, and cardiac outflow tract (OFT) development.[88]
Atrioventricular (AV) canal development
- AV boundary formation
- Notch signaling can regulate the atrioventricular boundary formation between the AV canal and the chamber myocardium.
Studies have revealed that both loss- and gain-of-function of the Notch pathway results in defects in AV canal development.[88] In addition, the Notch target genes HEY1 and HEY2 are involved in restricting the expression of two critical developmental regulator proteins, BMP2 and Tbx2, to the AV canal.[89][90]
- AV epithelial-mesenchymal transition (EMT)
- Notch signaling is also important for the process of AV EMT, which is required for AV canal maturation. After the AV canal boundary formation, a subset of endocardial cells lining the AV canal are activated by signals emanating from the myocardium and by interendocardial signaling pathways to undergo EMT.[88] Notch1 deficiency results in defective induction of EMT. Very few migrating cells are seen and these lack mesenchymal morphology.[91] Notch may regulate this process by activating matrix metalloproteinase2 (MMP2) expression, or by inhibiting vascular endothelial (VE)-cadherin expression in the AV canal endocardium[92] while suppressing the VEGF pathway via VEGFR2.[93] In RBPJk/CBF1-targeted mutants, the heart valve development is severely disrupted, presumably because of defective endocardial maturation and signaling.[91]
Ventricular development
Some studies in Xenopus[94] and in mouse embryonic stem cells[95] indicate that cardiomyogenic commitment and differentiation require Notch signaling inhibition. Active Notch signaling is required in the ventricular endocardium for proper trabeculae development subsequent to myocardial specification by regulating BMP10, NRG1, and EphrinB2 expression.[49] Notch signaling sustains immature cardiomyocyte proliferation in mammals [96][97][98] and zebrafish.[99] A regulatory correspondence likely exists between Notch signaling and Wnt signaling, whereby upregulated Wnt expression downregulates Notch signaling, and a subsequent inhibition of ventricular cardiomyocyte proliferation results. This proliferative arrest can be rescued using Wnt inhibitors.[100]
The downstream effector of Notch signaling, HEY2, was also demonstrated to be important in regulating ventricular development by its expression in the interventricular septum and the endocardial cells of the cardiac cushions.[101] Cardiomyocyte and smooth muscle cell-specific deletion of HEY2 results in impaired cardiac contractility, malformed right ventricle, and ventricular septal defects.[102]
Ventricular outflow tract development
During development of the aortic arch and the aortic arch arteries, the Notch receptors, ligands, and target genes display a unique expression pattern.[103] When the Notch pathway was blocked, the induction of vascular smooth muscle cell marker expression failed to occur, suggesting that Notch is involved in the differentiation of cardiac neural crest cells into vascular cells during outflow tract development.
Angiogenesis
Endothelial cells use the Notch signaling pathway to coordinate cellular behaviors during the blood vessel sprouting that occurs sprouting angiogenesis.[104][105][106][107]
Activation of Notch takes place primarily in "connector" cells and cells that line patent stable blood vessels through direct interaction with the Notch ligand, Delta-like ligand 4 (Dll4), which is expressed in the endothelial tip cells.[108] VEGF signaling, which is an important factor for migration and proliferation of endothelial cells,[109] can be downregulated in cells with activated Notch signaling by lowering the levels of Vegf receptor transcript.[110] Zebrafish embryos lacking Notch signaling exhibit ectopic and persistent expression of the zebrafish ortholog of VEGF3, flt4, within all endothelial cells, while Notch activation completely represses its expression.[111]
Notch signaling may be used to control the sprouting pattern of blood vessels during angiogenesis. When cells within a patent vessel are exposed to VEGF signaling, only a restricted number of them initiate the angiogenic process. Vegf is able to induce DLL4 expression. In turn, DLL4 expressing cells down-regulate Vegf receptors in neighboring cells through activation of Notch, thereby preventing their migration into the developing sprout. Likewise, during the sprouting process itself, the migratory behavior of connector cells must be limited to retain a patent connection to the original blood vessel.[108]
Role in endocrine development
During development, definitive endoderm and ectoderm differentiates into several gastrointestinal epithelial lineages, including endocrine cells. Many studies have indicated that Notch signaling has a major role in endocrine development.
Pancreatic development
The formation of the pancreas from endoderm begins in early development. The expression of elements of the Notch signaling pathway have been found in the developing pancreas, suggesting that Notch signaling is important in pancreatic development.[112][113] Evidence suggests Notch signaling regulates the progressive recruitment of endocrine cell types from a common precursor,[114] acting through two possible mechanisms. One is the "lateral inhibition", which specifies some cells for a primary fate but others for a secondary fate among cells that have the potential to adopt the same fate. Lateral inhibition is required for many types of cell fate determination. Here, it could explain the dispersed distribution of endocrine cells within pancreatic epithelium.[115] A second mechanism is "suppressive maintenance", which explains the role of Notch signaling in pancreas differentiation. Fibroblast growth factor10 is thought to be important in this activity, but the details are unclear.[116][117]
Intestinal development
The role of Notch signaling in the regulation of gut development has been indicated in several reports. Mutations in elements of the Notch signaling pathway affect the earliest intestinal cell fate decisions during zebrafish development.[118] Transcriptional analysis and gain of function experiments revealed that Notch signaling targets Hes1 in the intestine and regulates a binary cell fate decision between adsorptive and secretory cell fates.[118]
Bone development
Early in vitro studies have found the Notch signaling pathway functions as down-regulator in osteoclastogenesis and osteoblastogenesis.[119] Notch1 is expressed in the mesenchymal condensation area and subsequently in the hypertrophic chondrocytes during chondrogenesis.[120] Overexpression of Notch signaling inhibits bone morphogenetic protein2-induced osteoblast differentiation. Overall, Notch signaling has a major role in the commitment of mesenchymal cells to the osteoblastic lineage and provides a possible therapeutic approach to bone regeneration.[53]
Role in cell cycle control
Notch signaling is critical for cell fate identity and differentiation and regulates these processes in part by controlling cell cycle progression. Specifically, Notch has been shown to promote cell cycle progression at the G1/S transition in various systems.
Photoreceptor development
In Drosophila eye development, photoreceptors undergo two waves of differentiation, where five out of eight photoreceptors differentiate in the first wave (R8, R2, R5, R3, and R4), and the other three differentiate in the second wave (R1, R6, and R7).[121] Notch has been shown to promote the second mitotic wave in Drosophila eye development.[122] Specifically, it mediates the G1/S transition by promoting dE2F activation (Drosophila E2F), a member of the E2F transcription factor family, which regulates the expression of genes important for cell proliferation, specifically those involved in the G1/S transition.[122][123] Notch does this by inhibiting RBF1 (the Drosophila homolog of the tumor suppressor Rb), which represses dE2F.[122] Additionally, Notch is required for cyclin A activation, which accumulates during the G1/S transition and may be involved in S phase onset.[122]
Health and disease
The role of Notch signaling in cell cycle regulation also has implications in health and disease. For example, Notch has been found to promote the expression of cyclin D3 and Cdk4/6 in human T-cells, thereby promoting the phosphorylation of Rb and facilitating the G1/S transition, implicating its role in cancer as several gain-of-function mutations in NOTCH1 have been identified in human acute T-cell lymphoblastic leukemias and lymphomas.[124][125] Additionally, in ventricular cardiomyocytes, which stop dividing shortly after birth, NOTCH2 signaling activation promotes cell cycle reentry.[98] It induces the expression and nuclear translocation of cyclin D, which along with Cdk4/6 promotes the phosphorylation of Rb and causes cell cycle progression through the G1/S transition.[98] This suggests that Notch signaling might regulate ventricular growth as well as cardiomyocyte regeneration, though this is unclear.[98]
Migratory identity
In the zebrafish trunk neural crest (TNC), cells migrate collectively in single-file chains, with a cell "leader" at the front of the chain that instructs the directionality of the trailing "follower" cells.[126] Notch has been found to specify cell migratory identity in the trunk neural crest – specifically, high Notch specifies leaders while low Notch specifies followers.[127] Further, cell cycle progression required for migration is regulated by Notch such that leader cells with high Notch activity quickly undergo the G1/S transition while cells with low Notch activity remain in the G1 phase for longer and thus become followers.[127]
Role in cancer
Leukemia
Aberrant Notch signaling is a driver of T cell acute lymphoblastic leukemia (T-ALL)[128] and is mutated in at least 65% of all T-ALL cases.[129] Notch signaling can be activated by mutations in Notch itself, inactivating mutations in FBXW7 (a negative regulator of Notch1), or rarely by t(7;9)(q34;q34.3) translocation. In the context of T-ALL, Notch activity cooperates with additional oncogenic lesions such as c-MYC to activate anabolic pathways such as ribosome and protein biosynthesis thereby promoting leukemia cell growth.[130]
Urothelial bladder cancer
Loss of Notch activity is a driving event in urothelial cancer. A study identified inactivating mutations in components of the Notch pathway in over 40% of examined human bladder carcinomas. In mouse models, genetic inactivation of Notch signaling results in Erk1/2 phosphorylation leading to tumorigenesis in the urinary tract.[131] As not all NOTCH receptors are equally involved in the urothelial bladder cancer, 90% of samples in one study had some level of NOTCH3 expression, suggesting that NOTCH3 plays an important role in urothelial bladder cancer. A higher level of NOTCH3 expression was observed in high-grade tumors, and a higher level of positivity was associated with a higher mortality risk. NOTCH3 was identified as an independent predictor of poor outcome. Therefore, it is suggested that NOTCH3 could be used as a marker for urothelial bladder cancer-specific mortality risk. It was also shown that NOTCH3 expression could be a prognostic immunohistochemical marker for clinical follow-up of urothelial bladder cancer patients, contributing to a more individualized approach by selecting patients to undergo control cystoscopy after a shorter time interval.[132]
Liver cancer
In hepatocellular carcinoma, for instance, it was suggesting that AXIN1 mutations would provoke Notch signaling pathway activation, fostering the cancer development, but a recent study demonstrated that such an effect cannot be detected.[133] Thus the exact role of Notch signaling in the cancer process awaits further elucidation.
Role in Long-lived bat
The negative regulatory region (NRR) of human NOTCH3 maintains the receptor in an autoinhibited state by shielding the S2 cleavage site from ADAM proteases until ligand binding occurs. This region contains three Lin12–Notch repeats (LNRs) and a heterodimerization domain that together prevent premature activation. Comparative sequence analyses show that the long-lived bat Myotis brandtii lacks two short segments within LNR-A and LNR-C. One deleted motif in LNR-A, which normally helps occlude the S2 site, may increase its accessibility to ADAM10. Molecular dynamics simulations indicate that this deletion alone does not cause constitutive signaling but may facilitate a more readily cleavable S2 site and alter how the NRR responds to ligand-induced conformational changes. Although ligand-independent Notch activity is linked to several cancers—most notably involving NOTCH1—the functional consequences of altered NOTCH3 regulation remain comparatively understudied. Bats, including M. brandtii, are exceptional for their longevity relative to body size, yet the contribution of modified intercellular signaling pathways such as Notch to this trait is only beginning to be explored[134].
Notch inhibitors
The involvement of Notch signaling in many cancers has led to investigation of notch inhibitors (especially gamma-secretase inhibitors) as cancer treatments which are in different phases of clinical trials.[2][135] As of 2013[update] at least 7 notch inhibitors were in clinical trials.[136] MK-0752 has given promising results in an early clinical trial for breast cancer.[137] Preclinical studies showed beneficial effects of gamma-secretase inhibitors in endometriosis,[138] a disease characterised by increased expression of notch pathway constituents.[139][140] Several notch inhibitors, including the gamma-secretase inhibitor LY3056480, are being studied for their potential ability to regenerate hair cells in the cochlea, which could lead to treatments for hearing loss and tinnitus.[141][142]
Mathematical modeling
Mathematical modeling in Notch-Delta signaling has become a pivotal tool in understanding pattern formation driven by cell-cell interactions, particularly in the context of lateral-inhibition mechanisms. The Collier model,[143] a cornerstone in this field, employs a system of coupled ordinary differential equations to describe the feedback loop between adjacent cells. The model is defined by the equations:
where and represent the levels of Notch and Delta activity in cell , respectively. Functions and are typically Hill functions, reflecting the regulatory dynamics of the signaling process. The term denotes the average level of Delta activity in the cells adjacent to cell , integrating juxtacrine signaling effects.
Recent extensions of this model incorporate long-range signaling, acknowledging the role of cell protrusions like filopodia (cytonemes) that reach non-neighboring cells.[144][145][146][147] One extended model, often referred to as the -Collier model,[144] introduces a weighting parameter to balance juxtacrine and long-range signaling. The interaction term is modified to include these protrusions, creating a more complex, non-local signaling network. This model is instrumental in exploring pattern formation robustness and biological pattern refinement, considering the stochastic nature of filopodia dynamics and intrinsic noise. The application of mathematical modeling in Notch-Delta signaling has been particularly illuminating in understanding the patterning of sensory organ precursors (SOPs) in the Drosophila's notum and wing margin.[148][149]
The mathematical modeling of Notch-Delta signaling thus provides significant insights into lateral inhibition mechanisms and pattern formation in biological systems. It enhances the understanding of cell-cell interaction variations leading to diverse tissue structures, contributing to developmental biology and offering potential therapeutic pathways in diseases related to Notch-Delta dysregulation.
See also
- Alagille syndrome
- Netpath – A curated resource of signal transduction pathways in humans
References
- ↑ 1.0 1.1 "Notch signaling: cell fate control and signal integration in development". Science 284 (5415): 770–776. April 1999. doi:10.1126/science.284.5415.770. PMID 10221902. Bibcode: 1999Sci...284..770A.
- ↑ 2.0 2.1 "Notch Antagonists: Potential Modulators of Cancer and Inflammatory Diseases". Journal of Medicinal Chemistry 59 (17): 7719–7737. September 2016. doi:10.1021/acs.jmedchem.5b01516. PMID 27045975. https://figshare.com/articles/journal_contribution/7928825.
- ↑ "A novel proteolytic cleavage involved in Notch signaling: the role of the disintegrin-metalloprotease TACE". Molecular Cell 5 (2): 207–216. February 2000. doi:10.1016/S1097-2765(00)80417-7. PMID 10882063.
- ↑ "The Notch1/c-Myc pathway in T cell leukemia". Cell Cycle 6 (8): 927–930. April 2007. doi:10.4161/cc.6.8.4134. PMID 17404512.
- ↑ "Direct inhibition of the NOTCH transcription factor complex". Nature 462 (7270): 182–188. November 2009. doi:10.1038/nature08543. PMID 19907488. Bibcode: 2009Natur.462..182M.
- ↑ "Chemical biology: A Notch above other inhibitors". Nature 462 (7270): 171–173. November 2009. doi:10.1038/462171a. PMID 19907487. Bibcode: 2009Natur.462..171A.
- ↑ "The theory of the gene". The American Naturalist 51 (609): 513–544. 1917. doi:10.1086/279629. Bibcode: 1917ANat...51..513M. https://zenodo.org/record/1431357.
- ↑ The theory of the gene (revised ed.). Yale University Press. 1928. pp. 77–81. ISBN 978-0-8240-1384-4. http://onlinebooks.library.upenn.edu/webbin/book/lookupid?key=olbp24097.
- ↑ "Nucleotide sequence from the neurogenic locus notch implies a gene product that shares homology with proteins containing EGF-like repeats". Cell 43 (3 Pt 2): 567–581. December 1985. doi:10.1016/0092-8674(85)90229-6. PMID 3935325.
- ↑ "Sequence of the notch locus of Drosophila melanogaster: relationship of the encoded protein to mammalian clotting and growth factors". Molecular and Cellular Biology 6 (9): 3094–3108. September 1986. doi:10.1128/mcb.6.9.3094. PMID 3097517.
- ↑ "The lin-12 locus specifies cell fates in Caenorhabditis elegans". Cell 34 (2): 435–444. September 1983. doi:10.1016/0092-8674(83)90377-x. PMID 6616618.
- ↑ 12.0 12.1 12.2 "glp-1 is required in the germ line for regulation of the decision between mitosis and meiosis in C. elegans". Cell 51 (4): 589–599. November 1987. doi:10.1016/0092-8674(87)90128-0. PMID 3677168. Bibcode: 1987Cell...51..589A.
- ↑ "The glp-1 locus and cellular interactions in early C. elegans embryos". Cell 51 (4): 601–611. November 1987. doi:10.1016/0092-8674(87)90129-2. PMID 3677169.
- ↑ "lin-12, a nematode homeotic gene, is homologous to a set of mammalian proteins that includes epidermal growth factor". Cell 43 (3 Pt 2): 583–590. December 1985. doi:10.1016/0092-8674(85)90230-2. PMID 3000611.
- ↑ "p300 acts as a transcriptional coactivator for mammalian Notch-1". Molecular and Cellular Biology 21 (22): 7761–7774. November 2001. doi:10.1128/MCB.21.22.7761-7774.2001. PMID 11604511.
- ↑ "Antineurogenic phenotypes induced by truncated Notch proteins indicate a role in signal transduction and may point to a novel function for Notch in nuclei". Genes & Development 7 (10): 1949–1965. October 1993. doi:10.1101/gad.7.10.1949. PMID 8406001.
- ↑ "Intrinsic activity of the Lin-12 and Notch intracellular domains in vivo". Cell 74 (2): 331–345. July 1993. doi:10.1016/0092-8674(93)90424-O. PMID 8343960.
- ↑ "TAN-1, the human homolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms". Cell 66 (4): 649–661. August 1991. doi:10.1016/0092-8674(91)90111-B. PMID 1831692.
- ↑ "Nuclear access and action of notch in vivo". Cell 93 (4): 649–660. May 1998. doi:10.1016/S0092-8674(00)81193-9. PMID 9604939.
- ↑ "Notch-1 signalling requires ligand-induced proteolytic release of intracellular domain". Nature 393 (6683): 382–386. May 1998. doi:10.1038/30756. PMID 9620803. Bibcode: 1998Natur.393..382S.
- ↑ 21.0 21.1 "Notch and the awesome power of genetics". Genetics 191 (3): 655–669. July 2012. doi:10.1534/genetics.112.141812. PMID 22785620.
- ↑ "Presenilin-mediated transmembrane cleavage is required for Notch signal transduction in Drosophila". Proceedings of the National Academy of Sciences of the United States of America 98 (1): 229–234. January 2001. doi:10.1073/pnas.98.1.229. PMID 11134525. Bibcode: 2001PNAS...98..229S.
- ↑ "Notch signaling". Science 268 (5208): 225–232. April 1995. doi:10.1126/science.7716513. PMID 7716513. Bibcode: 1995Sci...268..225A.
- ↑ "The C. elegans spe-9 gene encodes a sperm transmembrane protein that contains EGF-like repeats and is required for fertilization". Cell 93 (1): 71–79. April 1998. doi:10.1016/S0092-8674(00)81147-2. PMID 9546393.
- ↑ "Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes". Cell 124 (5): 973–983. March 2006. doi:10.1016/j.cell.2005.12.037. PMID 16530044.
- ↑ "Crystal structure of the CSL-Notch-Mastermind ternary complex bound to DNA". Cell 124 (5): 985–996. March 2006. doi:10.1016/j.cell.2006.01.035. PMID 16530045.
- ↑ "The notch signalling regulator fringe acts in the Golgi apparatus and requires the glycosyltransferase signature motif DXD". Current Biology 10 (14): 813–820. July 2000. doi:10.1016/S0960-9822(00)00578-9. PMID 10899003. Bibcode: 2000CBio...10..813M.
- ↑ 28.0 28.1 "Notch signaling: control of cell communication and cell fate". Development 131 (5): 965–973. March 2004. doi:10.1242/dev.01074. PMID 14973298.
- ↑ "Fucosylation in prokaryotes and eukaryotes". Glycobiology 16 (12): 158R–184R. December 2006. doi:10.1093/glycob/cwl040. PMID 16973733.
- ↑ "O-glycosylation of EGF repeats: identification and initial characterization of a UDP-glucose: protein O-glucosyltransferase". Glycobiology 12 (11): 763–770. November 2002. doi:10.1093/glycob/cwf085. PMID 12460944.
- ↑ "Roles of O-Fucose Glycans in Notch Signaling Revealed by Mutant Mice". Functional Glycomics. Methods in Enzymology. 417. 2006. pp. 127–136. doi:10.1016/S0076-6879(06)17010-X. ISBN 978-0-12-182822-6.
- ↑ "The glycosyltransferase Fringe promotes Delta-Notch signaling between neurons and glia, and is required for subtype-specific glial gene expression". Development 134 (3): 591–600. February 2007. doi:10.1242/dev.02754. PMID 17215308.
- ↑ "The Notch ligands, Jagged and Delta, are sequentially processed by alpha-secretase and presenilin/gamma-secretase and release signaling fragments". The Journal of Biological Chemistry 278 (36): 34427–34437. September 2003. doi:10.1074/jbc.M302659200. PMID 12826675.
- ↑ "Metalloprotease ADAM10 is required for Notch1 site 2 cleavage". The Journal of Biological Chemistry 284 (45): 31018–31027. November 2009. doi:10.1074/jbc.M109.006775. PMID 19726682.
- ↑ "Drosophila Patterning: Delta-Notch Interactions". Encyclopedia of Life Sciences. 2006. doi:10.1038/npg.els.0004194. ISBN 0-470-01617-5.
- ↑ "Fine-tuning of the intracellular canonical Notch signaling pathway". Cell Cycle 11 (2): 264–276. January 2012. doi:10.4161/cc.11.2.18995. PMID 22223095.
- ↑ "Specific EGF repeats of Notch mediate interactions with Delta and Serrate: implications for Notch as a multifunctional receptor". Cell 67 (4): 687–699. November 1991. doi:10.1016/0092-8674(91)90064-6. PMID 1657403.
- ↑ "Specific EGF repeats of Notch mediate interactions with Delta and Serrate: implications for Notch as a multifunctional receptor". Cell 67 (4): 687–699. November 1991. doi:10.1016/0092-8674(91)90064-6. PMID 1657403. Bibcode: 2012Sci...338.1229Y.
- ↑ "Intrinsic selectivity of Notch 1 for Delta-like 4 over Delta-like 1". Journal of Biological Chemistry. 2013.
- ↑ 40.0 40.1 "Structural biology. Structural basis for Notch1 engagement of Delta-like 4". Science 347 (6224): 847–853. February 2015. doi:10.1126/science.1261093. PMID 25700513. Bibcode: 2015Sci...347..847L.
- ↑ "Protein binders and their applications in developmental biology". Development 145 (2). January 2018. doi:10.1242/dev.148874. PMID 29374062.
- ↑ "Combinatorial Antigen Targeting: Ideal T-Cell Sensing and Anti-Tumor Response". Trends in Molecular Medicine 22 (4): 271–273. April 2016. doi:10.1016/j.molmed.2016.02.009. PMID 26971630.
- ↑ "Chimeric antigen receptors: driving immunology towards synthetic biology". Current Opinion in Immunology 41: 68–76. August 2016. doi:10.1016/j.coi.2016.06.004. PMID 27372731.
- ↑ "The role of notch in promoting glial and neural stem cell fates". Annual Review of Neuroscience 25 (1): 471–490. 2002. doi:10.1146/annurev.neuro.25.030702.130823. PMID 12052917.
- ↑ 45.0 45.1 45.2 "Notch signaling in development and cancer". Endocrine Reviews 28 (3): 339–363. May 2007. doi:10.1210/er.2006-0046. PMID 17409286.
- ↑ "Notch and EGFR pathway interaction regulates neural stem cell number and self-renewal". Nature 467 (7313): 323–327. September 2010. doi:10.1038/nature09347. PMID 20844536. Bibcode: 2010Natur.467..323A.
- ↑ "Notch pathway molecules are essential for the maintenance, but not the generation, of mammalian neural stem cells". Genes & Development 16 (7): 846–858. April 2002. doi:10.1101/gad.975202. PMID 11937492.
- ↑ "Regulation of Notch1 and Dll4 by vascular endothelial growth factor in arterial endothelial cells: implications for modulating arteriogenesis and angiogenesis". Molecular and Cellular Biology 23 (1): 14–25. January 2003. doi:10.1128/MCB.23.1.14-25.2003. PMID 12482957.
- ↑ 49.0 49.1 "Notch signaling is essential for ventricular chamber development". Developmental Cell 12 (3): 415–429. March 2007. doi:10.1016/j.devcel.2006.12.011. PMID 17336907.
- ↑ The notch signaling pathway in cardiac development and tissue homeostasis
- ↑ "Notch signaling controls multiple steps of pancreatic differentiation". Proceedings of the National Academy of Sciences of the United States of America 100 (25): 14920–14925. December 2003. doi:10.1073/pnas.2436557100. PMID 14657333. Bibcode: 2003PNAS..10014920M.
- ↑ "Expression of notch receptors and ligands in the adult gut". The Journal of Histochemistry and Cytochemistry 52 (4): 509–516. April 2004. doi:10.1177/002215540405200409. PMID 15034002.
- ↑ 53.0 53.1 "Critical regulation of bone morphogenetic protein-induced osteoblastic differentiation by Delta1/Jagged1-activated Notch1 signaling". The Journal of Biological Chemistry 280 (16): 15842–15848. April 2005. doi:10.1074/jbc.M412891200. PMID 15695512.
- ↑ "Signaling axis involving Hedgehog, Notch, and Scl promotes the embryonic endothelial-to-hematopoietic transition". Proceedings of the National Academy of Sciences of the United States of America 110 (2): E141–E150. January 2013. doi:10.1073/pnas.1214361110. PMID 23236128. Bibcode: 2013PNAS..110E.141K.
- ↑ "Notch signaling in CD4 and CD8 T cell development". Current Opinion in Immunology 20 (2): 197–202. April 2008. doi:10.1016/j.coi.2008.03.004. PMID 18434124.
- ↑ "Role of Notch signaling in cell-fate determination of human mammary stem/progenitor cells". Breast Cancer Research 6 (6): R605–R615. 2004. doi:10.1186/bcr920. PMID 15535842.
- ↑ "Notch signaling promotes airway mucous metaplasia and inhibits alveolar development". Development 136 (10): 1751–1759. May 2009. doi:10.1242/dev.029249. PMID 19369400.
- ↑ "REX-1 expression and p38 MAPK activation status can determine proliferation/differentiation fates in human mesenchymal stem cells". PLOS ONE 5 (5). May 2010. doi:10.1371/journal.pone.0010493. PMID 20463961. Bibcode: 2010PLoSO...510493B.
- ↑ "Noncyclic Notch activity in the presomitic mesoderm demonstrates uncoupling of somite compartmentalization and boundary formation". Genes & Development 22 (16): 2166–2171. August 2008. doi:10.1101/gad.480408. PMID 18708576.
- ↑ "Left-right asymmetry in embryonic development: a comprehensive review". Mechanisms of Development 122 (1): 3–25. January 2005. doi:10.1016/j.mod.2004.08.006. PMID 15582774.
- ↑ "Two homologous regulatory genes, lin-12 and glp-1, have overlapping functions". Development 112 (1): 231–240. May 1991. doi:10.1242/dev.112.1.231. PMID 1769331.
- ↑ Gilbert SF (2016). Developmental biology (11th ed.). Sinauer. p. 272. ISBN 978-1-60535-470-5.
- ↑ 63.0 63.1 "Notch1 is required for the coordinate segmentation of somites". Development 121 (5): 1533–1545. May 1995. doi:10.1242/dev.121.5.1533. PMID 7789282.
- ↑ "Maintenance of somite borders in mice requires the Delta homologue DII1". Nature 386 (6626): 717–721. April 1997. doi:10.1038/386717a0. PMID 9109488. Bibcode: 1997Natur.386..717D.
- ↑ "Mutations affecting somite formation and patterning in the zebrafish, Danio rerio". Development 123: 153–164. December 1996. doi:10.1242/dev.123.1.153. PMID 9007237.
- ↑ "Analysis of Notch function in presomitic mesoderm suggests a gamma-secretase-independent role for presenilins in somite differentiation". Developmental Cell 8 (5): 677–688. May 2005. doi:10.1016/j.devcel.2005.02.019. PMID 15866159.
- ↑ "The many roles of Notch signaling during vertebrate somitogenesis". Seminars in Cell & Developmental Biology 49: 68–75. January 2016. doi:10.1016/j.semcdb.2014.11.010. PMID 25483003.
- ↑ "Stimulation of human epidermal differentiation by delta-notch signalling at the boundaries of stem-cell clusters". Current Biology 10 (9): 491–500. May 2000. doi:10.1016/s0960-9822(00)00451-6. PMID 10801437. Bibcode: 2000CBio...10..491L.
- ↑ "A Presenilin-2-ARF4 trafficking axis modulates Notch signaling during epidermal differentiation". The Journal of Cell Biology 214 (1): 89–101. July 2016. doi:10.1083/jcb.201508082. PMID 27354375.
- ↑ "Chromosomal Deficiencies and the Embryonic Development of Drosophila Melanogaster". Proceedings of the National Academy of Sciences of the United States of America 23 (3): 133–137. March 1937. doi:10.1073/pnas.23.3.133. PMID 16588136. Bibcode: 1937PNAS...23..133P.
- ↑ 71.0 71.1 "rax, Hes1, and notch1 promote the formation of Müller glia by postnatal retinal progenitor cells". Neuron 26 (2): 383–394. May 2000. doi:10.1016/S0896-6273(00)81171-X. PMID 10839357.
- ↑ 72.0 72.1 "An instructive function for Notch in promoting gliogenesis in the zebrafish retina". Development 128 (7): 1099–1107. April 2001. doi:10.1242/dev.128.7.1099. PMID 11245575.
- ↑ 73.0 73.1 "Nuclear Notch1 signaling and the regulation of dendritic development". Nature Neuroscience 3 (1): 30–40. January 2000. doi:10.1038/71104. PMID 10607392.
- ↑ 74.0 74.1 "Learning and memory deficits in Notch mutant mice". Current Biology 13 (15): 1348–1354. August 2003. doi:10.1016/S0960-9822(03)00492-5. PMID 12906797. Bibcode: 2003CBio...13.1348C.
- ↑ "Differential expression of mammalian Numb, Numblike and Notch1 suggests distinct roles during mouse cortical neurogenesis". Development 124 (10): 1887–1897. May 1997. doi:10.1242/dev.124.10.1887. PMID 9169836.
- ↑ "Inactivation of Numb and Numblike in embryonic dorsal forebrain impairs neurogenesis and disrupts cortical morphogenesis". Neuron 40 (6): 1105–1118. December 2003. doi:10.1016/S0896-6273(03)00755-4. PMID 14687546.
- ↑ "FGF signaling expands embryonic cortical surface area by regulating Notch-dependent neurogenesis". The Journal of Neuroscience 31 (43): 15604–15617. October 2011. doi:10.1523/JNEUROSCI.4439-11.2011. PMID 22031906.
- ↑ "Cortical gyrification induced by fibroblast growth factor 2 in the mouse brain". The Journal of Neuroscience 33 (26): 10802–10814. June 2013. doi:10.1523/JNEUROSCI.3621-12.2013. PMID 23804101.
- ↑ "Notch signalling regulates stem cell numbers in vitro and in vivo". Nature 442 (7104): 823–826. August 2006. doi:10.1038/nature04940. PMID 16799564. Bibcode: 2006Natur.442..823A. https://zenodo.org/record/1233295.
- ↑ "Notch3 is necessary for neuronal differentiation and maturation in the adult spinal cord". Journal of Cellular and Molecular Medicine 18 (10): 2103–2116. October 2014. doi:10.1111/jcmm.12362. PMID 25164209.
- ↑ "Murine numb regulates granule cell maturation in the cerebellum". Developmental Biology 266 (1): 161–177. February 2004. doi:10.1016/j.ydbio.2003.10.017. PMID 14729486.
- ↑ "Targeted deletion of numb and numblike in sensory neurons reveals their essential functions in axon arborization". Genes & Development 19 (1): 138–151. January 2005. doi:10.1101/gad.1246005. PMID 15598981.
- ↑ "Alzheimer's disease and Notch signaling". Biochemical and Biophysical Research Communications 390 (4): 1093–1097. December 2009. doi:10.1016/j.bbrc.2009.10.093. PMID 19853579. Bibcode: 2009BBRC..390.1093W.
- ↑ "Requirement of Notch in adulthood for neurological function and longevity". NeuroReport 12 (15): 3321–3325. October 2001. doi:10.1097/00001756-200110290-00035. PMID 11711879.
- ↑ "Loss of presenilin function causes impairments of memory and synaptic plasticity followed by age-dependent neurodegeneration". Neuron 42 (1): 23–36. April 2004. doi:10.1016/S0896-6273(04)00182-5. PMID 15066262.
- ↑ "Lessons from a failed γ-secretase Alzheimer trial". Cell 159 (4): 721–726. November 2014. doi:10.1016/j.cell.2014.10.016. PMID 25417150.
- ↑ "Ligand-Dependent Notch Signaling in Vascular Formation". Notch Signaling in Embryology and Cancer. Advances in Experimental Medicine and Biology. 727. 2012. pp. 210–222. doi:10.1007/978-1-4614-0899-4_16. ISBN 978-1-4614-0898-7.
- ↑ 88.0 88.1 88.2 "Notch signaling in cardiac development". Circulation Research 102 (10): 1169–1181. May 2008. doi:10.1161/CIRCRESAHA.108.174318. PMID 18497317.
- ↑ "Developmental patterning of the cardiac atrioventricular canal by Notch and Hairy-related transcription factors". Development 133 (21): 4381–4390. November 2006. doi:10.1242/dev.02607. PMID 17021042.
- ↑ "Hesr1 and Hesr2 regulate atrioventricular boundary formation in the developing heart through the repression of Tbx2". Development 134 (4): 747–755. February 2007. doi:10.1242/dev.02777. PMID 17259303.
- ↑ 91.0 91.1 "Notch promotes epithelial-mesenchymal transition during cardiac development and oncogenic transformation". Genes & Development 18 (1): 99–115. January 2004. doi:10.1101/gad.276304. PMID 14701881.
- ↑ "VE-cadherin is not required for the formation of nascent blood vessels but acts to prevent their disassembly". Blood 105 (7): 2771–2776. April 2005. doi:10.1182/blood-2004-06-2244. PMID 15604224.
- ↑ "Notch activation results in phenotypic and functional changes consistent with endothelial-to-mesenchymal transformation". Circulation Research 94 (7): 910–917. April 2004. doi:10.1161/01.RES.0000124300.76171.C9. PMID 14988227.
- ↑ "Serrate and Notch specify cell fates in the heart field by suppressing cardiomyogenesis". Development 127 (17): 3865–3876. September 2000. doi:10.1242/dev.127.17.3865. PMID 10934030.
- ↑ "Induction of cardiogenesis in embryonic stem cells via downregulation of Notch1 signaling". Circulation Research 98 (12): 1471–1478. June 2006. doi:10.1161/01.RES.0000226497.52052.2a. PMID 16690879.
- ↑ "Control of the adaptive response of the heart to stress via the Notch1 receptor pathway". The Journal of Experimental Medicine 205 (13): 3173–3185. December 2008. doi:10.1084/jem.20081427. PMID 19064701.
- ↑ "Notch1 signaling stimulates proliferation of immature cardiomyocytes". The Journal of Cell Biology 183 (1): 117–128. October 2008. doi:10.1083/jcb.200806091. PMID 18824567.
- ↑ 98.0 98.1 98.2 98.3 "Notch activates cell cycle reentry and progression in quiescent cardiomyocytes". The Journal of Cell Biology 183 (1): 129–141. October 2008. doi:10.1083/jcb.200806104. PMID 18838555.
- ↑ "Notch signaling regulates cardiomyocyte proliferation during zebrafish heart regeneration". Proceedings of the National Academy of Sciences of the United States of America 111 (4): 1403–1408. January 2014. doi:10.1073/pnas.1311705111. PMID 24474765. Bibcode: 2014PNAS..111.1403Z.
- ↑ "An Evolving Role for Notch Signaling in Heart Regeneration of the Zebrafish Danio rerio". https://www.researchgate.net/publication/362704455.
- ↑ "Atrioventricular cushion transformation is mediated by ALK2 in the developing mouse heart". Developmental Biology 286 (1): 299–310. October 2005. doi:10.1016/j.ydbio.2005.07.035. PMID 16140292.
- ↑ "Essential roles of the bHLH transcription factor Hrt2 in repression of atrial gene expression and maintenance of postnatal cardiac function". Proceedings of the National Academy of Sciences of the United States of America 104 (19): 7975–7980. May 2007. doi:10.1073/pnas.0702447104. PMID 17468400. Bibcode: 2007PNAS..104.7975X.
- ↑ "An essential role for Notch in neural crest during cardiovascular development and smooth muscle differentiation". The Journal of Clinical Investigation 117 (2): 353–363. February 2007. doi:10.1172/JCI30070. PMID 17273555.
- ↑ "Dll4 signalling through Notch1 regulates formation of tip cells during angiogenesis". Nature 445 (7129): 776–780. February 2007. doi:10.1038/nature05571. PMID 17259973. Bibcode: 2007Natur.445..776H.
- ↑ "Endothelial signalling by the Notch ligand Delta-like 4 restricts angiogenesis". Development 134 (5): 839–844. March 2007. doi:10.1242/dev.003244. PMID 17251261.
- ↑ "Delta-like ligand 4 (Dll4) is induced by VEGF as a negative regulator of angiogenic sprouting". Proceedings of the National Academy of Sciences of the United States of America 104 (9): 3219–3224. February 2007. doi:10.1073/pnas.0611206104. PMID 17296940. Bibcode: 2007PNAS..104.3219L.
- ↑ "Notch signalling limits angiogenic cell behaviour in developing zebrafish arteries". Nature 445 (7129): 781–784. February 2007. doi:10.1038/nature05577. PMID 17259972. Bibcode: 2007Natur.445..781S.
- ↑ 108.0 108.1 "Notch signalling and the regulation of angiogenesis". Cell Adhesion & Migration 1 (2): 104–106. 2007. doi:10.4161/cam.1.2.4488. PMID 19329884.
- ↑ "Signaling transduction mechanisms mediating biological actions of the vascular endothelial growth factor family". Cardiovascular Research 49 (3): 568–581. February 2001. doi:10.1016/S0008-6363(00)00268-6. PMID 11166270.
- ↑ "Up-regulation of the Notch ligand Delta-like 4 inhibits VEGF-induced endothelial cell function". Blood 107 (3): 931–939. February 2006. doi:10.1182/blood-2005-03-1000. PMID 16219802.
- ↑ "Notch signaling is required for arterial-venous differentiation during embryonic vascular development". Development 128 (19): 3675–3683. October 2001. doi:10.1242/dev.128.19.3675. PMID 11585794.
- ↑ "Notch signalling controls pancreatic cell differentiation". Nature 400 (6747): 877–881. August 1999. doi:10.1038/23716. PMID 10476967. Bibcode: 1999Natur.400..877A.
- ↑ "Notch gene expression during pancreatic organogenesis". Mechanisms of Development 94 (1–2): 199–203. June 2000. doi:10.1016/S0925-4773(00)00317-8. PMID 10842072.
- ↑ "Formation of the digestive system in zebrafish. II. Pancreas morphogenesis". Developmental Biology 261 (1): 197–208. September 2003. doi:10.1016/S0012-1606(03)00308-7. PMID 12941629.
- ↑ "Control of endodermal endocrine development by Hes-1". Nature Genetics 24 (1): 36–44. January 2000. doi:10.1038/71657. PMID 10615124.
- ↑ "Gene regulatory factors in pancreatic development". Developmental Dynamics 229 (1): 176–200. January 2004. doi:10.1002/dvdy.10460. PMID 14699589.
- ↑ "FGF10 signaling maintains the pancreatic progenitor cell state revealing a novel role of Notch in organ development". Developmental Biology 264 (2): 323–338. December 2003. doi:10.1016/j.ydbio.2003.08.013. PMID 14651921.
- ↑ 118.0 118.1 "Delta-Notch signalling controls commitment to a secretory fate in the zebrafish intestine". Development 132 (5): 1093–1104. March 2005. doi:10.1242/dev.01644. PMID 15689380.
- ↑ "Regulation of osteoclast development by Notch signaling directed to osteoclast precursors and through stromal cells". Blood 101 (6): 2227–2234. March 2003. doi:10.1182/blood-2002-06-1740. PMID 12411305.
- ↑ "Suppression of differentiation and proliferation of early chondrogenic cells by Notch". Journal of Bone and Mineral Metabolism 21 (6): 344–352. 2003. doi:10.1007/s00774-003-0428-4. PMID 14586790.
- ↑ Treisman, Jessica E. (July 2013). "Retinal differentiation in Drosophila" (in en). WIREs Developmental Biology 2 (4): 545–557. doi:10.1002/wdev.100. ISSN 1759-7684. PMID 24014422.
- ↑ 122.0 122.1 122.2 122.3 Baonza, Antonio; Freeman, Matthew (2005-04-01). "Control of Cell Proliferation in the Drosophila Eye by Notch Signaling" (in English). Developmental Cell 8 (4): 529–539. doi:10.1016/j.devcel.2005.01.019. ISSN 1534-5807. PMID 15809035.
- ↑ Johnson, David G.; Schneider-Broussard, Robin (1998-04-27). "Role of E2F in cell cycle control and cancer". Frontiers in Bioscience (Landmark Edition) 3 (4): 447–458. doi:10.2741/A291. ISSN 2768-6701. PMID 9556498. https://www.imrpress.com/journal/FBL/3/4/10.2741/A291.
- ↑ Joshi, Ila; Minter, Lisa M.; Telfer, Janice; Demarest, Renée M.; Capobianco, Anthony J.; Aster, Jon C.; Sicinski, Piotr; Fauq, Abdul et al. (2009-02-19). "Notch signaling mediates G1/S cell-cycle progression in T cells via cyclin D3 and its dependent kinases" (in en). Blood 113 (8): 1689–1698. doi:10.1182/blood-2008-03-147967. ISSN 0006-4971. PMID 19001083. PMC 2647664. https://ashpublications.org/blood/article/113/8/1689/26066/Notch-signaling-mediates-G1S-cellcycle-progression.
- ↑ Aster, Jon C. (2005-11-01). "Deregulated NOTCH Signaling in Acute T-Cell Lymphoblastic Leukemia/Lymphoma: New Insights, Questions, and Opportunities" (in en). International Journal of Hematology 82 (4): 295–301. doi:10.1532/IJH97.05096. ISSN 1865-3774. PMID 16298817. https://link.springer.com/article/10.1532/IJH97.05096.
- ↑ Richardson, Jo; Gauert, Anton; Briones Montecinos, Luis; Fanlo, Lucía; Alhashem, Zainalabdeen Mohmammed; Assar, Rodrigo; Marti, Elisa; Kabla, Alexandre et al. (2016-05-31). "Leader Cells Define Directionality of Trunk, but Not Cranial, Neural Crest Cell Migration" (in English). Cell Reports 15 (9): 2076–2088. doi:10.1016/j.celrep.2016.04.067. ISSN 2211-1247. PMID 27210753.
- ↑ 127.0 127.1 Alhashem, Zain; Feldner-Busztin, Dylan; Revell, Christopher; Alvarez-Garcillan Portillo, Macarena; Camargo-Sosa, Karen; Richardson, Joanna; Rocha, Manuel; Gauert, Anton et al. (2022-04-19). Piotrowski, Tatjana; White, Richard M. eds. "Notch controls the cell cycle to define leader versus follower identities during collective cell migration". eLife 11. doi:10.7554/eLife.73550. ISSN 2050-084X. PMID 35438077.
- ↑ "NOTCH1 pathway activation is an early hallmark of SCL T leukemogenesis". Blood 110 (10): 3753–3762. November 2007. doi:10.1182/blood-2006-12-063644. PMID 17698635.
- ↑ "Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia". Science 306 (5694): 269–271. October 2004. doi:10.1126/science.1102160. PMID 15472075. Bibcode: 2004Sci...306..269W.
- ↑ "NOTCH1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth". Proceedings of the National Academy of Sciences of the United States of America 103 (48): 18261–18266. November 2006. doi:10.1073/pnas.0606108103. PMID 17114293. Bibcode: 2006PNAS..10318261P.
- ↑ "A new tumor suppressor role for the Notch pathway in bladder cancer". Nature Medicine 20 (10): 1199–1205. October 2014. doi:10.1038/nm.3678. PMID 25194568.
- ↑ "The association between NOTCH3 expression and the clinical outcome in the urothelial bladder cancer patients". Bosnian Journal of Basic Medical Sciences 22 (4): 523–530. July 2022. doi:10.17305/bjbms.2021.6767. PMID 35073251.
- ↑ "Unraveling the impact of AXIN1 mutations on HCC development: Insights from CRISPR/Cas9 repaired AXIN1-mutant liver cancer cell lines.". PLOS ONE 19 (6). June 2024. doi:10.1371/journal.pone.0304607. PMID 38848383. Bibcode: 2024PLoSO..1904607Z.
- ↑ Siddiqui, Khawar Sohail; Ertan, Haluk; Ren, Yi; Poljak, Anne; Jayasena, Tharusha; Sachdev, Perminder (2025-09-29). "Unveiling the evolutionary code of NOTCH3: mammalian bioinformatics sheds light on human pathogenicity". bioRxiv 10.1101/2025.09.28.678708.
- ↑ "Notch Inhibition as a Promising New Approach to Cancer Therapy". Notch Signaling in Embryology and Cancer. Advances in Experimental Medicine and Biology. 727. 2012. pp. 305–319. doi:10.1007/978-1-4614-0899-4_23. ISBN 978-1-4614-0898-7.
- ↑ "Notch inhibitors for cancer treatment". Pharmacology & Therapeutics 139 (2): 95–110. August 2013. doi:10.1016/j.pharmthera.2013.02.003. PMID 23458608.
- ↑ "Notch inhibitors could help overcome therapy resistance in ER-positive breast cancer". 2015. https://oncology-central.com/content/news/notch-inhibitors-could-help-overcome-therapy-resistance-in-er-positive-breast-cancer.
- ↑ "γ-Secretase inhibition affects viability, apoptosis, and the stem cell phenotype of endometriotic cells". Acta Obstetricia et Gynecologica Scandinavica 98 (12): 1565–1574. December 2019. doi:10.1111/aogs.13707. PMID 31424097.
- ↑ "The endometrial stem cell markers notch-1 and numb are associated with endometriosis". Reproductive Biomedicine Online 36 (3): 294–301. March 2018. doi:10.1016/j.rbmo.2017.11.010. PMID 29398419. https://zenodo.org/record/3695800.
- ↑ "Increased expression of the adult stem cell marker Musashi-1 in endometriosis and endometrial carcinoma". The Journal of Pathology 215 (3): 317–329. July 2008. doi:10.1002/path.2364. PMID 18473332.
- ↑ "Hair Cell Generation in Cochlear Culture Models Mediated by Novel γ-Secretase Inhibitors". Frontiers in Cell and Developmental Biology (Frontiers Media SA) 9. 13 August 2021. doi:10.3389/fcell.2021.710159. PMID 34485296.
- ↑ "Therapeutic Potential of Wnt and Notch Signaling and Epigenetic Regulation in Mammalian Sensory Hair Cell Regeneration". Molecular Therapy (Elsevier BV) 27 (5): 904–911. May 2019. doi:10.1016/j.ymthe.2019.03.017. PMID 30982678.
- ↑ "Pattern formation by lateral inhibition with feedback: a mathematical model of delta-notch intercellular signalling". Journal of Theoretical Biology 183 (4): 429–446. December 1996. doi:10.1006/jtbi.1996.0233. PMID 9015458. Bibcode: 1996JThBi.183..429C. https://ora.ox.ac.uk/objects/uuid:d18e2570-90fd-47d5-94ad-3a4df1c0fe4c.
- ↑ 144.0 144.1 "Coupling dynamics of 2D Notch-Delta signalling". Mathematical Biosciences 360. June 2023. doi:10.1016/j.mbs.2023.109012. PMID 37142213.
- ↑ "Pattern formation in discrete cell tissues under long range filopodia-based direct cell to cell contact". Mathematical Biosciences 273: 1–15. March 2016. doi:10.1016/j.mbs.2015.12.008. PMID 26748293.
- ↑ "Faculty Opinions recommendation of Dynamic filopodia transmit intermittent Delta-Notch signaling to drive pattern refinement during lateral inhibition.". 2010-07-27. doi:10.3410/f.4361976.4187082.
- ↑ "The importance of structured noise in the generation of self-organizing tissue patterns through contact-mediated cell-cell signalling". Journal of the Royal Society, Interface 8 (59): 787–798. June 2011. doi:10.1098/rsif.2010.0488. PMID 21084342.
- ↑ "Modeling the Notch Response". Molecular Mechanisms of Notch Signaling. Advances in Experimental Medicine and Biology. 1066. Cham: Springer International Publishing. 2018. pp. 79–98. doi:10.1007/978-3-319-89512-3_5. ISBN 978-3-319-89512-3.
- ↑ "Coordinated control of Notch/Delta signalling and cell cycle progression drives lateral inhibition-mediated tissue patterning". Development 143 (13): 2305–2310. July 2016. doi:10.1242/dev.134213. PMID 27226324.
External links
- Diagram: notch signaling pathway in Homo sapiens
- Diagram: Notch signaling in Drosophila
- Notch+Receptors at the US National Library of Medicine Medical Subject Headings (MeSH)
Template:Notch signaling pathway
 |
