Biology:Plant evolutionary developmental biology
Evolutionary developmental biology (evo-devo) is the study of developmental programs and patterns from an evolutionary perspective.[1] It seeks to understand the various influences shaping the form and nature of life on the planet. Evo-devo arose as a separate branch of science rather recently. An early sign of this occurred in 1999.[2]
Most of the synthesis in evo-devo has been in the field of animal evolution, one reason being the presence of model systems like Drosophila melanogaster, C. elegans, zebrafish and Xenopus laevis. However, since 1980, a wealth of information on plant morphology, coupled with modern molecular techniques has helped shed light on the conserved and unique developmental patterns in the plant kingdom also.[3][4]
Historical perspective
Before 1900
The origin of the term "morphology" is generally attributed to Johann Wolfgang von Goethe (1749–1832). He was of the opinion that there is an underlying fundamental organisation (Bauplan) in the diversity of flowering plants. In his book The Metamorphosis of Plants, he proposed that the Bauplan enabled us to predict the forms of plants that had not yet been discovered.[5] Goethe was the first to make the perceptive suggestion that flowers consist of modified leaves. He also entertained different complementary interpretations.[6][7]
In the middle centuries, several basic foundations of our current understanding of plant morphology were laid down. Nehemiah Grew, Marcello Malpighi, Robert Hooke, Antonie van Leeuwenhoek, Wilhelm von Nageli were just some of the people who helped build knowledge on plant morphology at various levels of organisation. It was the taxonomical classification of Carl Linnaeus in the eighteenth century though, that generated a firm base for the knowledge to stand on and expand.[8] The introduction of the concept of Darwinism in contemporary scientific discourse also had had an effect on the thinking on plant forms and their evolution.
Wilhelm Hofmeister, one of the most brilliant botanists of his times, was the one to diverge away from the idealist way of pursuing botany. Over the course of his life, he brought an interdisciplinary outlook into botanical thinking. He came up with biophysical explanations on phenomena like phototaxis and geotaxis, and also discovered the alternation of generations in the plant life cycle.[5]
1900 to the present
The past century witnessed a rapid progress in the study of plant anatomy. The focus shifted from the population level to more reductionist levels. While the first half of the century saw expansion in developmental knowledge at the tissue and the organ level, in the latter half, especially since the 1990s, there has also been a strong impetus on gaining molecular information.
Edward Charles Jeffrey was one of the early evo-devo researchers of the 20th century. He performed a comparative analyses of the vasculatures of living and fossil gymnosperms and came to the conclusion that the storage parenchyma has been derived from tracheids.[9] His research[10] focussed primarily on plant anatomy in the context of phylogeny. This tradition of evolutionary analyses of plant architectures was further advanced by Katherine Esau, best known for her book The Plant Anatomy. Her work focussed on the origin and development of various tissues in different plants. Working with Vernon Cheadle,[11] she also explained the evolutionary specialization of the phloem tissue with respect to its function.
In 1959 Walter Zimmermann published a revised edition of Die Phylogenie der Planzen.[12] This very comprehensive work, which has not been translated into English, has no equal in the literature. It presents plant evolution as the evolution of plant development (hologeny). In this sense it is plant evolutionary developmental biology (plant evo-devo). According to Zimmermann, diversity in plant evolution occurs though various developmental processes. Three very basic processes are heterochrony (changes in the timing of developmental processes), heterotopy (changes in the relative positioning of processes), and heteromorphy (changes in form processes).[13]
In the meantime, by the beginning of the latter half of the 1900s, Arabidopsis thaliana had begun to be used in some developmental studies. The first collection of Arabidopsis thaliana mutants were made around 1945.[14] However it formally became established as a model organism only in 1998.[15]
The recent spurt in information on various plant-related processes has largely been a result of the revolution in molecular biology. Powerful techniques like mutagenesis and complementation were made possible in Arabidopsis thaliana via generation of T-DNA containing mutant lines, recombinant plasmids, techniques like transposon tagging etc. Availability of complete physical and genetic maps,[16] RNAi vectors, and rapid transformation protocols are some of the technologies that have significantly altered the scope of the field.[15] Recently, there has also been a massive increase in the genome and EST sequences[17] of various non-model species, which, coupled with the bioinformatics tools existing today, generate opportunities in the field of plant evo-devo research.
Gérard Cusset provided a detailed in-depth analysis of the history of plant morphology, including plant development and evolution, from its beginnings to the end of the 20th century.[18] Rolf Sattler discussed fundamental principles of plant morphology[19][20] and plant evo-devo.[21][22][23] Rolf Rutishauser surveyed the past and future of plant evo-devo with regard to continuum and process morphology.[24]
Organisms, databases and tools
The most important model systems in plant development have been arabidopsis and maize. Maize has traditionally been the favorite of plant geneticists, while extensive resources in almost every area of plant physiology and development are available for Arabidopsis thaliana. Apart from these, rice, Antirrhinum majus, Brassica, and tomato are also being used in a variety of studies. The genomes of Arabidopsis thaliana and rice have been completely sequenced, while the others are in process.[25] It must be emphasized here that the information from these "model" organisms form the basis of our developmental knowledge. While Brassica has been used primarily because of its convenient location in the phylogenetic tree in the mustard family, Antirrhinum majus is a convenient system for studying leaf architecture. Rice has been traditionally used for studying responses to hormones like abscissic acid and gibberelin as well as responses to stress. However, recently, not just the domesticated rice strain, but also the wild strains have been studied for their underlying genetic architectures.[26]
Some people have objected against extending the results of model organisms to the plant world. One argument is that the effect of gene knockouts in lab conditions wouldn't truly reflect even the same plant's response in the natural world. Also, these supposedly crucial genes might not be responsible for the evolutionary origin of that character. For these reasons, a comparative study of plant traits has been proposed as the way to go now.[27]
Since the past few years, researchers have indeed begun looking at non-model, "non-conventional" organisms using modern genetic tools. One example of this is the Floral Genome Project, which envisages to study the evolution of the current patterns in the genetic architecture of the flower through comparative genetic analyses, with a focus on EST sequences.[28] Like the FGP, there are several such ongoing projects that aim to find out conserved and diverse patterns in evolution of the plant shape. Expressed sequence tag (EST) sequences of quite a few non-model plants like sugarcane, apple, barley, cycas, coffee, to name a few, are available freely online.[29] The Cycad Genomics Project,[30] for example, aims to understand the differences in structure and function of genes between gymnosperms and angiosperms through sampling in the order Cycadales. In the process, it intends to make available information for the study of evolution of seeds, cones and evolution of life cycle patterns. Presently the most important sequenced genomes from an evo-devo point of view include those of A. thaliana (a flowering plant), poplar (a woody plant), Physcomitrella patens (a bryophyte), Maize (extensive genetic information), and Chlamydomonas reinhardtii (a green alga). The impact of such a vast amount of information on understanding common underlying developmental mechanisms can easily be realised.
Apart from EST and genome sequences, several other tools like PCR, yeast two-hybrid system, microarrays, RNA Interference, SAGE, QTL mapping etc. permit the rapid study of plant developmental patterns. Recently, cross-species hybridization has begun to be employed on microarray chips, to study the conservation and divergence in mRNA expression patterns between closely related species.[31] Techniques for analyzing this kind of data have also progressed over the past decade. We now have better models for molecular evolution, more refined analysis algorithms and better computing power as a result of advances in computer sciences.
Evolution of plant morphology
Overview of plant evolution
Evidence suggests that an algal scum formed on the land 1,200 million years ago, but it was not until the Ordovician period, around 500 million years ago, that land plants appeared. These began to diversify in the late Silurian period, around 420 million years ago, and the fruits of their diversification are displayed in remarkable detail in an early Devonian fossil assemblage known as the Rhynie chert. This chert preserved early plants in cellular detail, petrified in volcanic springs. By the middle of the Devonian period most of the features recognised in plants today are present, including roots and leaves. By the late Devonian, plants had reached a degree of sophistication that allowed them to form forests of tall trees. Evolutionary innovation continued after the Devonian period. Most plant groups were relatively unscathed by the Permo-Triassic extinction event, although the structures of communities changed. This may have set the scene for the evolution of flowering plants in the Triassic (~200 million years ago), which exploded the Cretaceous and Tertiary. The latest major group of plants to evolve were the grasses, which became important in the mid Tertiary, from around 40 million years ago. The grasses, as well as many other groups, evolved new mechanisms of metabolism to survive the low CO
2 and warm, dry conditions of the tropics over the last 10 million years. Although animals and plants evolved their bodyplan independently, they both express a developmental constraint during mid-embryogenesis that limits their morphological diversification.[32][33][34][35][36]
Meristems
The meristem architectures differ between angiosperms, gymnosperms and pteridophytes. The gymnosperm vegetative meristem lacks organization into distinct tunica and corpus layers. They possess large cells called central mother cells. In angiosperms, the outermost layer of cells divides anticlinally to generate the new cells, while in gymnosperms, the plane of division in the meristem differs for different cells. However, the apical cells do contain organelles like large vacuoles and starch grains, like the angiosperm meristematic cells.
Pteridophytes, like fern, on the other hand, do not possess a multicellular apical meristem. They possess a tetrahedral apical cell, which goes on to form the plant body. Any somatic mutation in this cell can lead to hereditary transmission of that mutation.[37] The earliest meristem-like organization is seen in an algal organism from group Charales that has a single dividing cell at the tip, much like the pteridophytes, yet simpler. One can thus see a clear pattern in evolution of the meristematic tissue, from pteridophytes to angiosperms: Pteridophytes, with a single meristematic cell; gymnosperms with a multicellular, but less defined organization; and finally, angiosperms, with the highest degree of organization.
Evolution of plant transcriptional regulation
Transcription factors and transcriptional regulatory networks play key roles in plant development and stress responses, as well as their evolution. During plant landing, many novel transcription factor families emerged and are preferentially wired into the networks of multicellular development, reproduction, and organ development, contributing to more complex morphogenesis of land plants.[38]
Evolution of leaves
Origins of the leaf
Leaves are the primary photosynthetic organs of a plant. Based on their structure, they are classified into two types - microphylls, that lack complex venation patterns and megaphylls, that are large and with a complex venation. It has been proposed that these structures arose independently.[39] Megaphylls, according to the telome theory, have evolved from plants that showed a three-dimensional branching architecture, through three transformations: planation, which involved formation of a planar architecture, webbing, or formation of the outgrowths between the planar branches and fusion, where these webbed outgrowths fused to form a proper leaf lamina. Studies have revealed that these three steps happened multiple times in the evolution of today's leaves.[40]
Contrary to the telome theory, developmental studies of compound leaves have shown that, unlike simple leaves, compound leaves branch in three dimensions.[41][42] Consequently, they appear partially homologous with shoots as postulated by Agnes Arber in her partial-shoot theory of the leaf.[43] They appear to be part of a continuum between morphological categories, especially those of leaf and shoot.[44][45] Molecular genetics confirmed these conclusions (see below).
It has been proposed that the before the evolution of leaves, plants had the photosynthetic apparatus on the stems. Today's megaphyll leaves probably became commonplace some 360 mya, about 40 my after the simple leafless plants had colonized the land in the early Devonian period. This spread has been linked to the fall in the atmospheric carbon dioxide concentrations in the late Paleozoic era associated with a rise in density of stomata on leaf surface. This must have allowed for better transpiration rates and gas exchange. Large leaves with less stomata would have heated up in the sun's rays, but an increased stomatal density allowed for a better-cooled leaf, thus making its spread feasible.[46][47]
Factors influencing leaf architectures
Various physical and physiological forces like light intensity, humidity, temperature, wind speeds etc. are thought to have influenced evolution of leaf shape and size. It is observed that high trees rarely have large leaves, owing to the obstruction they generate for winds. This obstruction can eventually lead to the tearing of leaves, if they are large. Similarly, trees that grow in temperate or taiga regions have pointed leaves, presumably to prevent nucleation of ice onto the leaf surface and reduce water loss due to transpiration. Herbivory, not only by large mammals, but also small insects has been implicated as a driving force in leaf evolution, an example being plants of the genus Aciphylla, that are commonly found in New Zealand. The now-extinct moas (birds) fed upon these plants, and the spines on the leaves probably discouraged the moas from feeding on them. Other members of Aciphylla that did not co-exist with the moas were spineless.[48]
Genetic evidences for leaf evolution
At the genetic level, developmental studies have shown that repression of the KNOX genes is required for initiation of the leaf primordium. This is brought about by ARP genes, which encode transcription factors. Genes of this type have been found in many plants studied till now, and the mechanism i.e. repression of KNOX genes in leaf primordia, seems to be quite conserved. Expression of KNOX genes in leaves produces complex leaves. It is speculated that the ARP function arose quite early in vascular plant evolution, because members of the primitive group lycophytes also have a functionally similar gene [49] Other players that have a conserved role in defining leaf primordia are the phytohormone auxin, gibberelin and cytokinin.
One feature of a plant is its phyllotaxy. The arrangement of leaves on the plant body is such that the plant can maximally harvest light under the given constraints, and hence, one might expect the trait to be genetically robust. However, it may not be so. In maize, a mutation in only one gene called abphyl (abnormal phyllotaxy) was enough to change the phyllotaxy of the leaves. It implies that sometimes, mutational tweaking of a single locus on the genome is enough to generate diversity. The abphyl gene was later on shown to encode a cytokinin response regulator protein.[50]
Once the leaf primordial cells are established from the SAM cells, the new axes for leaf growth are defined, one important (and more studied) among them being the abaxial-adaxial (lower-upper surface) axis. The genes involved in defining this, and the other axes seem to be more or less conserved among higher plants. Proteins of the HD-ZIPIII family have been implicated in defining the adaxial identity. These proteins deviate some cells in the leaf primordium from the default abaxial state, and make them adaxial. It is believed that in early plants with leaves, the leaves just had one type of surface - the abaxial one. This is the underside of today's leaves. The definition of the adaxial identity occurred some 200 million years after the abaxial identity was established.[27] One can thus imagine the early leaves as an intermediate stage in evolution of today's leaves, having just arisen from spiny stem-like outgrowths of their leafless ancestors, covered with stomata all over, and not optimized as much for light harvesting.
How the infinite variety of plant leaves is generated is a subject of intense research. Some common themes have emerged. One of the most significant is the involvement of KNOX genes in generating compound leaves, as in tomato (see above). But this again is not universal. For example, pea uses a different mechanism for doing the same thing.[51][52] Mutations in genes affecting leaf curvature can also change leaf form, by changing the leaf from flat, to a crinkly shape,[53] like the shape of cabbage leaves. There also exist different morphogen gradients in a developing leaf which define the leaf's axis. Changes in these morphogen gradients may also affect the leaf form. Another very important class of regulators of leaf development are the microRNAs, whose role in this process has just begun to be documented. The coming years should see a rapid development in comparative studies on leaf development, with many EST sequences involved in the process coming online.
Molecular genetics has also shed light on the relation between radial symmetry (characteristic of stems) and dorsiventral symmetry (typical for leaves). James (2009) stated that "it is now widely accepted that... radiality [characteristic of most shoots] and dorsiventrality [characteristic of leaves] are but extremes of a continuous spectrum. In fact, it is simply the timing of the KNOX gene expression!"[54] In fact there is evidence for this continuum already at the beginning of land plant evolution.[55] Furthermore, studies in molecular genetics confirmed that compound leaves are intermediate between simple leaves and shoots, that is, they are partially homologous with simple leaves and shoots, since "it is now generally accepted that compound leaves express both leaf and shoot properties”.[56] This conclusion was reached by several authors on purely morphological grounds.[41][42]
Evolution of flowers
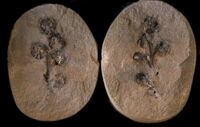
Flower-like structures first appear in the fossil records some ~130 mya, in the Cretaceous era.[57]
The flowering plants have long been assumed to have evolved from within the gymnosperms; according to the traditional morphological view, they are closely allied to the gnetales. However, recent molecular evidence is at odds to this hypothesis,[58][59] and further suggests that gnetales are more closely related to some gymnosperm groups than angiosperms,[60] and that gymnosperms form a distinct clade to the angiosperms,.[58][59][60] Molecular clock analysis predicts the divergence of flowering plants (anthophytes) and gymnosperms to ~300 mya[61]
The main function of a flower is reproduction, which, before the evolution of the flower and angiosperms, was the job of microsporophylls and megasporophylls. A flower can be considered a powerful evolutionary innovation, because its presence allowed the plant world to access new means and mechanisms for reproduction.
Origins of the flower
It seems that on the level of the organ, the leaf may be the ancestor of the flower, or at least some floral organs. When we mutate some crucial genes involved in flower development, we end up with a cluster of leaf-like structures. Thus, sometime in history, the developmental program leading to formation of a leaf must have been altered to generate a flower. There probably also exists an overall robust framework within which the floral diversity has been generated. An example of that is a gene called LEAFY (LFY), which is involved in flower development in Arabidopsis thaliana. The homologs of this gene are found in angiosperms as diverse as tomato, snapdragon, pea, maize and even gymnosperms. Expression of Arabidopsis thaliana LFY in distant plants like poplar and citrus also results in flower-production in these plants. The LFY gene regulates the expression of some gene belonging to the MADS-box family. These genes, in turn, act as direct controllers of flower development.
Evolution of the MADS-box family
The members of the MADS-box family of transcription factors play a very important and evolutionarily conserved role in flower development. According to the ABC model of flower development, three zones - A, B and C - are generated within the developing flower primordium, by the action of some transcription factors, that are members of the MADS-box family. Among these, the functions of the B and C domain genes have been evolutionarily more conserved than the A domain gene. Many of these genes have arisen through gene duplications of ancestral members of this family. Quite a few of them show redundant functions.
The evolution of the MADS-box family has been extensively studied. These genes are present even in pteridophytes, but the spread and diversity is many times higher in angiosperms.[62] There appears to be quite a bit of pattern into how this family has evolved. Consider the evolution of the C-region gene AGAMOUS (AG). It is expressed in today's flowers in the stamens, and the carpel, which are reproductive organs. It's ancestor in gymnosperms also has the same expression pattern. Here, it is expressed in the strobili, an organ that produces pollens or ovules.[63] Similarly, the B-genes' (AP3 and PI) ancestors are expressed only in the male organs in gymnosperms. Their descendants in the modern angiosperms also are expressed only in the stamens, the male reproductive organ. Thus, the same, then-existing components were used by the plants in a novel manner to generate the first flower. This is a recurring pattern in evolution.
Factors influencing floral diversity
How is the enormous diversity in the shape, color and sizes of flowers established? There is enormous variation in the developmental program in different plants. For example, monocots possess structures like lodicules and palea, that were believed to be analogous to the dicot petals and carpels respectively. It turns out that this is true, and the variation is due to slight changes in the MADS-box genes and their expression pattern in the monocots. Another example is that of the toad-flax, Linaria vulgaris, which has two kinds of flower symmetries: radial and bilateral. These symmetries are due to changes in copy number, timing, and location of expression in CYCLOIDEA, which is related to TCP1 in Arabidopsis.[57][64]
Arabidopsis thaliana has a gene called AGAMOUS that plays an important role in defining how many petals and sepals and other organs are generated. Mutations in this gene give rise to the floral meristem obtaining an indeterminate fate, and many floral organs keep on getting produced. We have flowers like roses, carnations and morning glory, for example, that have very dense floral organs. These flowers have been selected by horticulturists since long for increased number of petals. Researchers have found that the morphology of these flowers is because of strong mutations in the AGAMOUS homolog in these plants, which leads to them making a large number of petals and sepals.[65] Several studies on diverse plants like petunia, tomato, impatiens, maize etc. have suggested that the enormous diversity of flowers is a result of small changes in genes controlling their development.[66]
Some of these changes also cause changes in expression patterns of the developmental genes, resulting in different phenotypes. The Floral Genome Project looked at the EST data from various tissues of many flowering plants. The researchers confirmed that the ABC Model of flower development is not conserved across all angiosperms. Sometimes expression domains change, as in the case of many monocots, and also in some basal angiosperms like Amborella. Different models of flower development like the fading boundaries model, or the overlapping-boundaries model which propose non-rigid domains of expression, may explain these architectures.[67] There is a possibility that from the basal to the modern angiosperms, the domains of floral architecture have gotten more and more fixed through evolution.
Flowering time
Another floral feature that has been a subject of natural selection is flowering time. Some plants flower early in their life cycle, others require a period of vernalization before flowering. This decision is based on factors like temperature, light intensity, presence of pollinators and other environmental signals. In Arabidopsis thaliana it is known that genes like CONSTANS (CO), FRIGIDA, Flowering Locus C (FLC) and FLOWERING LOCUS T (FT) integrate the environmental signals and initiate the flower development pathway. Allelic variation in these loci have been associated with flowering time variations between plants. For example, Arabidopsis thaliana ecotypes that grow in the cold temperate regions require prolonged vernalization before they flower, while the tropical varieties and common lab strains, do not. Much of this variation is due to mutations in the FLC and FRIGIDA genes, rendering them non-functional.[68]
Many genes in the flowering time pathway are conserved across all plants studied to date. However, this does not mean that the mechanism of action is similarly conserved. For example, the monocot rice accelerates its flowering in short-day conditions, while Arabidopsis thaliana, a eudicot, responds to long-day conditions. In both plants, the proteins CO and FT are present but in Arabidopsis thaliana CO enhances FT production, while in rice the CO homolog represses FT production, resulting in completely opposite downstream effects.[69]
Theories of flower evolution
There are many theories that propose how flowers evolved. Some of them are described below.
The Anthophyte Theory was based on the observation that a gymnospermic family Gnetaceae has a flower-like ovule. It has partially developed vessels as found in the angiosperms, and the megasporangium is covered by three envelopes, like the ovary structure of angiosperm flowers. However, many other lines of evidence show that gnetophytes are not related to angiosperms.[70]
The Mostly Male Theory has a more genetic basis. Proponents of this theory point out that the gymnosperms have two very similar copies of the gene LFY while angiosperms only have one. Molecular clock analysis has shown that the other LFY paralog was lost in angiosperms around the same time as flower fossils become abundant, suggesting that this event might have led to floral evolution.[71] According to this theory, loss of one of the LFY paralog led to flowers that were more male, with the ovules being expressed ectopically. These ovules initially performed the function of attracting pollinators, but sometime later, may have been integrated into the core flower.
Evolution of secondary metabolism

Plant secondary metabolites are low molecular weight compounds, sometimes with complex structures that have no essential role in primary metabolism. They function in processes such as anti-herbivory, pollinator attraction, communication between plants, allelopathy, maintenance of symbiotic associations with soil flora and enhancing the rate of fertilization. Secondary metabolites have great structural and functional diversity and many thousands of enzymes may be involved in their synthesis, coded for by as much as 15–25% of the genome.[72] Many plant secondary metabolites such as the colour and flavor components of saffron and the chemotherapeutic drug taxol are of culinary and medical significance to humans and are therefore of commercial importance. In plants they seem to have diversified using mechanisms such as gene duplications, evolution of novel genes and the development of novel biosynthetic pathways. Studies have shown that diversity in some of these compounds may be positively selected for.[citation needed] Cyanogenic glycosides may have been proposed to have evolved multiple times in different plant lineages, and there are several other instances of convergent evolution. For example, the enzymes for synthesis of limonene – a terpene – are more similar between angiosperms and gymnosperms than to their own terpene synthesis enzymes. This suggests independent evolution of the limonene biosynthetic pathway in these two lineages.[73]
Mechanisms and players in evolution
While environmental factors are significantly responsible for evolutionary change, they act merely as agents for natural selection. Some of the changes develop through interactions with pathogens. Change is inherently brought about via phenomena at the genetic level – mutations, chromosomal rearrangements and epigenetic changes. While the general types of mutations hold true across the living world, in plants, some other mechanisms have been implicated as highly significant.
Polyploidy is a very common feature in plants. It is believed that at least half plants are or have been polyploids. Polyploidy leads to genome doubling, thus generating functional redundancy in most genes. The duplicated genes may attain new function, either by changes in expression pattern or changes in activity. Polyploidy and gene duplication are believed to be among the most powerful forces in evolution of plant form. It is not known though, why genome doubling is such a frequent process in plants. One possible reason is the production of large amounts of secondary metabolites in plant cells. Some of them might interfere in the normal process of chromosomal segregation, leading to polypoidy.
In recent times, plants have been shown to possess significant microRNA families, which are conserved across many plant lineages. In comparison to animals, while the number of plant miRNA families is less, the size of each family is much larger. The miRNA genes are also much more spread out in the genome than those in animals, where they are found clustered. It has been proposed that these miRNA families have expanded by duplications of chromosomal regions.[74] Many miRNA genes involved in regulation of plant development have been found to be quite conserved between plants studied.
Domestication of plants such as maize, rice, barley, wheat etc. has also been a significant driving force in their evolution. Some studies[clarification needed] have looked at the origins of the maize plant and found that maize is a domesticated derivative of a wild plant from Mexico called teosinte. Teosinte belongs to the genus Zea, just as maize, but bears very small inflorescence, 5–10 hard cobs, and a highly branched and spread-out stem.

Crosses between a particular teosinte variety and maize yield fertile offspring that are intermediate in phenotype between maize and teosinte. QTL analysis has also revealed some loci that when mutated in maize yield a teosinte-like stem or teosinte-like cobs. Molecular clock analysis of these genes estimates their origins to some 9000 years ago, well in accordance with other records of maize domestication. It is believed that a small group of farmers must have selected some maize-like natural mutant of teosinte some 9000 years ago in Mexico, and subjected it to continuous selection to yield the maize plant as known today.[75]
Another case is that of cauliflower. The edible cauliflower is a domesticated version of the wild plant Brassica oleracea, which does not possess the dense undifferentiated inflorescence, called the curd, that cauliflower possesses.
| Wikispecies has information related to Brassicaceae |
Cauliflower possesses a single mutation in a gene called CAL, controlling meristem differentiation into inflorescence. This causes the cells at the floral meristem to gain an undifferentiated identity, and instead of growing into a flower, they grow into a lump of undifferentiated cells.[76] This mutation has been selected through domestication at least since the Greek empire.
See also
- Plant morphology
- Comparative phylogenetics
- Plant evolution
- Evolutionary history of plants
References
- ↑ Hall B (2000). "Evo-Devo or Devo-Evo - Does it matter?". Evolution and Development 2 (4): 177–178. doi:10.1046/j.1525-142x.2000.00003e.x. PMID 11252559.
- ↑ Goodman C, Coughlin B (2000). "The evolution of evo-devo biology". Proc. Natl. Acad. Sci. USA 97 (9): 4424–4425. doi:10.1073/pnas.97.9.4424. PMID 10781035.
- ↑ Vergara-Silva, F (2003). "Plants and the conceptual articulation of evolutionary developmental biology". Biology and Philosophy 18 (2): 249–284. doi:10.1023/A:1023936102602.
- ↑ Minelli, A. 2018. Plant Evolutionary Biology. The Evolvablity of the Phenotype. New York: Cambridge University Press.
- ↑ 5.0 5.1 Kaplan D (2001). "The Science of Plant Morphology: Definition, History and Role in Modern Biology". Am. J. Bot. (Botanical Society of America) 88 (10): 1711–1741. doi:10.2307/3558347. PMID 21669604.
- ↑ Cusset G (1982). "The Conceptual Bases of Plant Morphology." In: Sattler R (1982). Axioms and Principles of Plant Construction. The Hague: Martinus Nijoff/ Dr W. Junk Publishers, p.27 (also published in Acta Biotheretica 31A)
- ↑ Sattler, R. (2018). "Philosophy of Plant Morphology". Elemente der Naturwissenschaft 108: 55–79.(for an expanded version of this article see [1])
- ↑ http://www.cas.muohio.edu/~meicenrd/ANATOMY/Ch0_History/history.html Plant Morphology: Timeline
- ↑ Jeffrey CE (1925). "The Origin of Parenchyma in Geological Time". Proc. Natl. Acad. Sci. USA 11 (1): 106–110. doi:10.1073/pnas.11.1.106. PMID 16586955. Bibcode: 1925PNAS...11..106J.
- ↑ Jeffrey CE (1910). Collected papers. pp. 317–. https://books.google.com/books?id=RIouAAAAIAAJ&pg=PA317.
- ↑ Plant Science Bulletin
- ↑ Zimmermann, W. 1959. Die Phylogenie der Pflanzen. Revised 2nd edition. Stuttgart: Gustav Fischer Verlag.
- ↑ Sattler, R.; Rutishauser, R. (2023). "Fundamentals of plant morphology and plant evo-devo (evolutionary developmental biology)". Plants 12 (1): 118–131. doi:10.3390/plants12010118. PMID 36616247.
- ↑ [2] TAIR: About Arabidopsis
- ↑ 15.0 15.1 Fink G (1998). "Anatomy of a Revolution". Genetics 149 (2): 473–477. doi:10.1093/genetics/149.2.473. PMID 9611166.
- ↑ The Arabidopsis Information Resource
- ↑ NCBI dbEST Statistics
- ↑ Cusset G (1982). "The Conceptual Bases of Plant Morphology." In: Sattler R (1982). Axioms and Principles of Plant Construction. The Hague: Martinus Nijoff/ Dr W. Junk Publishers, pp. 8–86 (also published in Acta Biotheretica 31A)
- ↑ Sattler, R. "Plant Morphology. Fundamental Issues".[3]
- ↑ Sattler, R. (2018). "Philosophy of Plant Morphology". Elemente der Naturwissenschaft 108: 55–79.(for an expanded version of this article see [4])
- ↑ Sattler, R.; Rutishauser, R. (2023). "Fundamentals of plant morphology and plant evo-devo (evolutionary developmental biology)". Plants 12 (1): 118–131. doi:10.3390/plants12010118. PMID 36616247.
- ↑ Sattler, R. "Plant evo-devo of the gynoecium".[5]
- ↑ Sattler, R. "Plant evo-devo and the morphological research by Rolf Sattler and collaboraters". [6]
- ↑ Rutishauser, R. (2020). "EvoDevo: Past and future of continuum and process plant morphology". Philosophies 5 (4): 166–168. doi:10.3390/philosophies5040041. PMID 5040041.
- ↑ NCBI Plant Genomes Central
- ↑ "Phylogeny of rice genomes with emphasis on origins of allotetraploid species". Proc. Natl. Acad. Sci. USA 96 (25): 14400–14405. 1999. doi:10.1073/pnas.96.25.14400. PMID 10588717. Bibcode: 1999PNAS...9614400G.
- ↑ 27.0 27.1 Cronk Q. (2001). "Plant evolution and development in a post-genomic context". Nature Reviews Genetics 2 (8): 607–619. doi:10.1038/35084556. PMID 11483985.
- ↑ The Floral Genome Project Home
- ↑ PlantGDB
- ↑ Cycad Genomics Project home
- ↑ Lai Z; Gross, BL; Zou, Y; Andrews, J; Rieseberg, LH (2006). "Microarray analysis reveals differential gene expression in hybrid sunflower species". Molecular Ecology 15 (5): 1213–1227. doi:10.1111/j.1365-294X.2006.02775.x. PMID 16626449.
- ↑ Drost, Hajk-Georg; Janitza, Philipp; Grosse, Ivo; Quint, Marcel (2017). "Cross-kingdom comparison of the developmental hourglass". Current Opinion in Genetics & Development 45: 69–75. doi:10.1016/j.gde.2017.03.003. PMID 28347942.
- ↑ Irie, Naoki; Kuratani, Shigeru (2011-03-22). "Comparative transcriptome analysis reveals vertebrate phylotypic period during organogenesis" (in en). Nature Communications 2: 248. doi:10.1038/ncomms1248. ISSN 2041-1723. PMID 21427719. Bibcode: 2011NatCo...2..248I.
- ↑ Domazet-Lošo, Tomislav; Tautz, Diethard (2010-12-09). "A phylogenetically based transcriptome age index mirrors ontogenetic divergence patterns" (in en). Nature 468 (7325): 815–818. doi:10.1038/nature09632. ISSN 0028-0836. PMID 21150997. Bibcode: 2010Natur.468..815D.
- ↑ Quint, Marcel; Drost, Hajk-Georg; Gabel, Alexander; Ullrich, Kristian Karsten; Bönn, Markus; Grosse, Ivo (2012-10-04). "A transcriptomic hourglass in plant embryogenesis" (in en). Nature 490 (7418): 98–101. doi:10.1038/nature11394. ISSN 0028-0836. PMID 22951968. Bibcode: 2012Natur.490...98Q.
- ↑ Drost, Hajk-Georg; Gabel, Alexander; Grosse, Ivo; Quint, Marcel (2015-05-01). "Evidence for Active Maintenance of Phylotranscriptomic Hourglass Patterns in Animal and Plant Embryogenesis" (in en). Molecular Biology and Evolution 32 (5): 1221–1231. doi:10.1093/molbev/msv012. ISSN 0737-4038. PMID 25631928.
- ↑ Klekowski E. (2003). "Plant clonality, mutation, diplontic selection and mutational meltdown". Biol. J. Linn. Soc. 79 (1): 61–67. doi:10.1046/j.1095-8312.2003.00183.x.
- ↑ Jin JP (July 2015). "An Arabidopsis transcriptional regulatory map reveals distinct functional and evolutionary features of novel transcription factors". Molecular Biology and Evolution 32 (7): 1767–1773. doi:10.1093/molbev/msv058. PMID 25750178.
- ↑ Crane and Kenrick; Kenrick, Paul (1997). "Diverted development of reproductive organs: A source of morphological innovation in land plants". Plant Systematics and Evolution 206 (1): 161–174. doi:10.1007/BF00987946.
- ↑ "Evolution of leaf developmental mechanisms". New Phytol. 167 (3): 693–710. 2005. doi:10.1111/j.1469-8137.2005.01466.x. PMID 16101907.
- ↑ 41.0 41.1 Rutishauser, R. and Sattler, R. 1997. Expression of shoot processes in leaf development of Polemonium caeruleum. Botanische Jahrbücher für Systematik 119: 563-582.
- ↑ 42.0 42.1 Lacroix C. (2003). "Shoot and compound leaf comparisons in eudicots: dynamic morphology as an alternative approach". Botanical Journal of the Linnean Society 143 (3): 219–230. doi:10.1046/j.1095-8339.2003.00222.x.
- ↑ Rutishauser, R.; Isler, B. (2001). "Developmental genetics and morphological evolution of flowering plants, especially bladderworts (Utricularia): fuzzy arberian morphology complements classical morphology". Annals of Botany 88: 1184. doi:10.1006/anbo.2001.1498. http://www.systbot.uzh.ch/static/personen/rolf_rutishauser_assets/Rut_Isl_AnnBot_2001.pdf.
- ↑ Sattler, R.; Jeune, B. (1992). "Multivariate analysis confirms the continuum view of plant form". Annals of Botany 69 (3): 249–262. doi:10.1093/oxfordjournals.aob.a088338.
- ↑ Jeune B., Sattler R. (1992). "Multivariate analysis in process morphology". Journal of Theoretical Biology 156: 147–167. doi:10.1016/s0022-5193(05)80670-8.
- ↑ "Evolution of leaf-form in land plants linked to atmospheric CO2 decline in the Late Palaeozoic era". Nature 410 (6826): 352–354. 2001. doi:10.1038/35066546. PMID 11268207. Bibcode: 2001Natur.410..352B. http://eprints.whiterose.ac.uk/59/1/osbornecp2.pdf.
- ↑ A perspective on the CO2 theory of early leaf evolution
- ↑ "Herbivory and the Evolution of Leaf Size and Shape". Philosophical Transactions of the Royal Society B 333 (1267): 265–272. 1991. doi:10.1098/rstb.1991.0076.
- ↑ Harrison C. J. (2005). "Independent recruitment of a conserved developmental mechanism during leaf evolution". Nature 434 (7032): 509–514. doi:10.1038/nature03410. PMID 15791256. Bibcode: 2005Natur.434..509H.
- ↑ Jackson D., Hake S. (1999). "Control of Phyllotaxy in Maize by the ABPHYL1 Gene". Development 126 (2): 315–323. doi:10.1242/dev.126.2.315. PMID 9847245.
- ↑ Tattersall et al. (2005). "The Mutant crispa Reveals Multiple Roles for PHANTASTICA in Pea Compound Leaf Development". Plant Cell 17 (4): 1046–1060. doi:10.1105/tpc.104.029447. PMID 15749758.
- ↑ Bharathan and Sinha; Sinha, NR (Dec 2001). "The Regulation of Compound Leaf Development". Plant Physiol. 127 (4): 1533–1538. doi:10.1104/pp.010867. PMID 11743098.
- ↑ "Genetic Control of Surface Curvature". Science 299 (5611): 1404–1407. 2003. doi:10.1126/science.1079354. PMID 12610308. http://rico-coen.jic.ac.uk/uploads/0/0f/Nath_Science.pdf.
- ↑ James, P. J. 2009. 'Tree and Leaf': A different angle. The Linnean 25, p. 17.
- ↑ Sattler, R. 1998. On the origin of symmetry, branching and phyllotaxis in land plants. In: R.V. Jean and D. Barabé (eds) Symmetry in Plants. World Scientific, Singapore, pp. 775-793.
- ↑ Eckardt N. A., Baum D. (2010). "The Podostemad puzzle: The evolution of unusual morphology in the Podostemonaceae". The Plant Cell 22 (7): 2131–2140. doi:10.1105/tpc.110.220711. PMID 20647343.
- ↑ 57.0 57.1 "Molecular evolution of flower development". Trends Ecol. Evol. 15 (4): 144–149. 2000. doi:10.1016/S0169-5347(99)01816-9. PMID 10717683.
- ↑ 58.0 58.1 Chaw, S.M.; Parkinson, C.L.; Cheng, Y.; Vincent, T.M.; Palmer, J.D. (2000). "Seed plant phylogeny inferred from all three plant genomes: Monophyly of extant gymnosperms and origin of Gnetales from conifers". Proceedings of the National Academy of Sciences 97 (8): 4086–91. doi:10.1073/pnas.97.8.4086. PMID 10760277. Bibcode: 2000PNAS...97.4086C.
- ↑ 59.0 59.1 Soltis, D.E.; Soltis, P.S.; Zanis, M.J. (2002). "Phylogeny of seed plants based on evidence from eight genes". American Journal of Botany 89 (10): 1670–81. doi:10.3732/ajb.89.10.1670. PMID 21665594.
- ↑ 60.0 60.1 Bowe, L.M.; Coat, G.; Depamphilis, C.W. (2000). "Phylogeny of seed plants based on all three genomic compartments: Extant gymnosperms are monophyletic and Gnetales' closest relatives are conifers". Proceedings of the National Academy of Sciences 97 (8): 4092–7. doi:10.1073/pnas.97.8.4092. PMID 10760278. Bibcode: 2000PNAS...97.4092B.
- ↑ Nam, J.; Depamphilis, CW; Ma, H; Nei, M (2003). "Antiquity and Evolution of the MADS-Box Gene Family Controlling Flower Development in Plants". Mol. Biol. Evol. 20 (9): 1435–1447. doi:10.1093/molbev/msg152. PMID 12777513. http://mbe.oxfordjournals.org/cgi/content/full/20/9/1435.
- ↑ Medarg NG and Yanofsky M (March 2001). "Function and evolution of the plant MADS-box gene family". Nature Reviews Genetics 2 (3): 186–195. doi:10.1038/35056041. PMID 11256070.
- ↑ Jager et al. (2003). "MADS-Box Genes in Ginkgo biloba and the Evolution of the AGAMOUS Family". Mol. Biol. Evol. 20 (5): 842–854. doi:10.1093/molbev/msg089. PMID 12679535. http://mbe.oxfordjournals.org/cgi/content/full/20/5/842.
- ↑ Kramer, Elena M. (2019). "Plus ça change, plus c'est la même chose: The developmental evolution of flowers". Current Topics in Developmental Biology 131: 211–238. doi:10.1016/bs.ctdb.2018.11.015. ISBN 9780128098042. ISSN 1557-8933. PMID 30612618. https://pubmed.ncbi.nlm.nih.gov/30612618/.
- ↑ Kitahara K and Matsumoto S. (2000). "Rose MADS-box genes 'MASAKO C1 and D1' homologous to class C floral identity genes". Plant Science 151 (2): 121–134. doi:10.1016/S0168-9452(99)00206-X. PMID 10808068.
- ↑ "Multiple AGAMOUS Homologs from Cucumber and Petunia Differ in Their Ability to Induce Reproductive Organ Fate". Plant Cell 10 (2): 171–182. 1998. doi:10.1105/tpc.10.2.171. PMID 9490741.
- ↑ "The floral genome: an evolutionary history of gene duplication and shifting patterns of gene expression". Trends Plant Sci. 12 (8): 358–367. 2007. doi:10.1016/j.tplants.2007.06.012. PMID 17658290.
- ↑ Putterill (2004). "It's time to flower: the genetic control of flowering time". BioEssays 26 (4): 353–363. doi:10.1002/bies.20021. PMID 15057934. http://www3.interscience.wiley.com/cgi-bin/abstract/107640082/ABSTRACT.
- ↑ Blazquez et al. (2001). "Flowering on time: genes that regulate the floral transition". EMBO Reports 2 (12): 1078–1082. doi:10.1093/embo-reports/kve254. PMID 11743019.
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedCrepet2000 - ↑ "The Mostly Male Theory of Flower Evolutionary Origins: from Genes to Fossils". Syst. Bot. (American Society of Plant Taxonomists) 25 (2): 155–170. 2000. doi:10.2307/2666635.
- ↑ Pichersky E. and Gang D. (2000). "Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective". Trends Plant Sci. 5 (10): 439–445. doi:10.1016/S1360-1385(00)01741-6. PMID 11044721.
- ↑ "Plant terpenoid synthases: molecular and phylogenetic analysis". Proc. Natl. Acad. Sci. USA 95 (8): 4126–4133. 1998. doi:10.1073/pnas.95.8.4126. PMID 9539701.
- ↑ Li A and Mao L. (2007). "Evolution of plant microRNA gene families". Cell Research 17 (3): 212–218. doi:10.1038/sj.cr.7310113. PMID 17130846.
- ↑ Doebley J.F. (2004). "The genetics of maize evolution". Annu. Rev. Genet. 38: 37–59. doi:10.1146/annurev.genet.38.072902.092425. PMID 15568971. https://zenodo.org/record/895359.
- ↑ Purugannan et al. (2000). "Variation and Selection at the CAULIFLOWER Floral Homeotic Gene Accompanying the Evolution of Domesticated Brassica olerace". Genetics 155 (2): 855–862. doi:10.1093/genetics/155.2.855. PMID 10835404.
Further reading
- The Genetics of plant morphological evolution
- Plant evolution and development in a post-genomic context
- Evolution of leaf developmental mechanisms
- Developmental genetics and plant evolution ISBN:0-415-25791-3
 |









