Biology:Zingibain
| Zingibain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
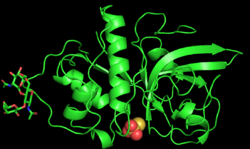 PYMOL generated 3D structure of zingibain monomer | |||||||||
| Identifiers | |||||||||
| EC number | 3.4.22.67 | ||||||||
| CAS number | 246044-91-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Zingibain, zingipain, or ginger protease (EC 3.4.22.67) is a cysteine protease enzyme found in ginger (Zingiber officinale) rhizomes.[1][2][3] It catalyses the preferential cleavage of peptides with a proline residue at the P2 position. It has two distinct forms, ginger protease I (GP-I) and ginger protease II (GP-II).[4]
As a member of the papain-like protease family of cysteine proteases, zingibain shares several structural and functional similarities with more well-studied enzymes such as papain, bromelain, and actinidin. These peptidases contain an active cysteine residue in their centers that catalyzes the hydrolytic cleavage of peptide bonds. Zingibain is noted for its activity as a proteinase and a collagenase.[1]
It was first isolated, purified, and reported in 1973 by Ichikawa et al. at Japan Women's University.[5] Recently, zingibain was found to exist as two isozymes, GP-I and GP-II, which were isolated by chromatography, with molecular weights of approximately 22,500 Da.[5]
Mechanism
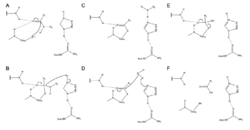
Zingibain utilizes a catalytic triad of Cys, His, and Asn residues in its active site in order to cleave peptide bonds hydrolytically. The presence of Asn175 stabilizes the imidazole ring of His, ensuring it is positioned optimally to catalyze hydrolysis.
The mechanism begins with a proton transfer from Cys25 to His159.[6] The sulfide anion then attacks the amino acid's alpha carbon, displacing the amine group, which attaches to His159.[6] The alpha carbon on the stabilized amino acid is then attacked by a water molecule, which displaces the sulfide of Cys25 to convert the amino acid to a carboxylic acid, which is released from the enzyme active site.[6]
The experimental introduction of dithiothreitol, a known thiol group protector, improves proteolytic activity, providing further verification of the importance of the central cysteine residue to enzymatic activity.[7]
Zingibain exhibits maximum turnover rate at 60 °C and rapidly denatures at 70 °C.[8] Proteolysis is largely unhampered during cooking with ginger. Optimal temperature ranges of papain and ficin are elevated relative to zingibain, whereas bromelain operates at a slightly lower range.[7]
Maximum proteolytic activity of zingibain occurs at pH of 6.0, although the enzyme is still active in pH ranges from 4.5 to 6.0 (optimal pH for meat marinades).[7]
GP-II, the more acidic of the two isozymes, exhibits a pI of 4.82, and GP-I exhibits pI values at 5.05 or 5.16.[1] These multiple pI values lend support to a theory that GP-I may be a mixture of two proteins.
Structure
Zingibain was first purified and characterized with X-ray crystallography in 2000 by researchers at Boston University.[1]
The enzyme is 221 amino acids long and glycosylated with 2 N-linked oligosaccharide chains at Asn96 and Asn154.[3] The polypeptide chain of zingibain folds into two polar domains of roughly equal size, divided by a central neutral cleft.[3] The first domain contains alpha helices, and the second has antiparallel beta sheets.[3] This separation of polar and non-polar regions facilitates protein-protein interactions between the enzyme and a large range of substrates.[3]
The active site of zingibain, located in the central cleft, is 5.5 Å deep and 9.5 Å long.[3] The active site contains the catalytic triad of Cys25, His159, and Asn175, which both cooperatively enable acid/base catalysis.
Zingibain exhibits binding specificity to peptide substrates with proline in the P2 position.[1] The S2 subsite of zingibain contains the amino acid chain Trp67-Met68-Asn69-Thr133-Ala157, which makes the site too compact to accommodate larger hydrophobic aromatic substrate residues favored by other enzymes in the papain family.[1] Proline, however, is stabilized by multiple non-covalent interactions with this region.
The enzyme structure is stabilized by hydrogen bonds, as well as crosslinking sulfide bonds between three pairs of cysteine residues (Cys22-Cys63, Cys56-Cys95, and Cys153-Cys200), analogous to many other papains.[1]
While the enzyme exists as a monomer in solution, crystallized zingibain forms tetramers, or dimers of dimers, linked by glycosylation chains on each subunit. Zingibain exhibits complex-type N-linked oligosaccharide chains at two residues.[1] Chains are between 5-13 glycosyl units long, and composed of N-acetylglucosamine, fucose, mannose, and xylose. Zingibain sugar sequences are almost identical to oligosaccharides seen in lectins from Japanese pagoda tree seeds, laccase a from sycamore cells, and S-glycoproteins from Brassica campestris.[1]
Biological significance
Within ginger rhizomes, ginger protease participates in multiple functional roles for maintenance and upkeep of plant cells.
Zingibain, like most cysteine proteases, is synthesized as a 40-50 kDa proprotein within cytoplasmic polysomes bound to cell membranes.[9] Within the endoplasmic reticulum, these elongated chains are tagged with a KDEL ER retention signal and placed into large KDEL vesicles that move from the ER to protein storage vacuoles in cell walls.[9]
Zingibain likely participates in protein storage (within seeds or plant tissue), but predominantly degrades and mobilizes storage proteins. It can also respond to abiotic and biotic stresses, such as heat shock, cold temperatures, and dehydration, to eliminate any resulting misfolded or denatured proteins.[9]
Uses
Meat tenderizer
Like papain from papayas and bromelain from pineapples, it is used as a meat tenderizer.[10][11]
When added to cooking meat, usually within raw or dried ginger, zingibain has been shown to increase the tenderness of meat.[8][12] Meat tenderization occurs due to zingibain's rapid proteolysis of major muscle proteins within meat, especially actomyosin and Type I collagen, which is found in muscle joints.[8]
While other papain enzymes, including papain, ficin, and bromelain, are more commonly used to tenderize meat, zingibain shows similar or elevated proteolytic activity.[11] In fact, zingibain is the only catalogued plant protease with collagenolytic activity. Zingibain may be a preferable meat tenderizer to papain due to the resulting texture of meat produced. While papain can hydrolyze actomyosin, it also breaks down other major tissue proteins, that lead to a mushy meat texture.[11] The specificity of zingibain's binding ensures predominant hydrolyzation of actomyosin and Type I collagen.
Zingibain is also used to flavor sausages and baked products.[8]
Rennet substitute
For over the past 100 years, ginger protease has traditionally been used to curdle milk to create ginger milk curd, a gel-like Cantonese dish made from hot milk and ginger juice. The milk clotting ability and specificity of ginger protease to proteolysis of κ-casein make the enzyme a potential vegetable rennet substitute for cheese production.
Milk coagulation is traditionally accomplished by coagulating enzymes extracted from sources such as rennet. In rennet, three chymosin isozymes hydrolyze κ-casein, a major protein fraction within milk, between Phe105 and Met106. Hydrophilic sub-regions of κ-casein are cleaved off, leaving behind largely hydrophobic aggregate. The enzymes thus destabilize κ-casein micelles and encourage clumping of hydrophobic protein residues, causing milk to curdle.
Major industrial drawbacks of rennet include its limited supply and high cost, its inaccessibility to vegetarians and practicing members of certain religious groups, and recent European national bans on utilization of recombinant calf rennet.[13] Fungal proteases are largely unsuitable as rennet substitutes, and enzymes from many plant extracts have been shown to produce low yields, poor textures, and bitter flavors of cheese.[13]
Commercial drawbacks
However, crude ginger protease extracted from ginger extract is unstable, with a half-life of about 2 days at 5 °C, making it problematic for commercial applications.[14] While the enzyme's half-life does not impede its efficacy during cooking, this low storage stability requires improvement for commercialization.
Commercial attempts to stabilize the enzyme for large-scale production have investigated potential methods to inactivate the free sulfhydryl group within the enzyme's active site. Mechanistic possibilities include oxidizing the sulfhydryl, exchanging it with disulfide bridges, forming quinone-thiol adducts, or binding the sulfhydryl to a heavy metal ion.[14] 0.2% sodium ascorbate was found to stabilize zingibain for up to 14 days at 5 °C, whereas comparable concentrations of EDTA and CaCl2 had minimal impact on stability.[14]
Zingibain has been observed to deactivate itself through autolysis, which can be pre-empted by reacting the active sulfhydryl group with cystine or PCMB.[14]
Acetone powders are a viable commercial method of stabilization of zingibain. After hydrophobic plant polyphenols are removed from crude ginger, acetone powder is introduced at low temperatures in order to dehydrate the root pulp.[14] The enzyme is stabilized due to reduced water activity, lower concentrations of plant pigments, and more rigid 3D structures at lower temperatures.[14]
See also
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 "Amino-acid sequence and glycan structures of cysteine proteases with proline specificity from ginger rhizome Zingiber officinale". Eur. J. Biochem. 267 (5): 1516–26. 2000. doi:10.1046/j.1432-1327.2000.01152.x. PMID 10691991.
- ↑ "Purification of ginger proteases by DEAE-Sepharose and isoelectric focusing". Biochimica et Biophysica Acta (BBA) - General Subjects 1243 (2): 181–4. 1995. doi:10.1016/0304-4165(94)00145-n. PMID 7873561.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 "The 2.1 A structure of a cysteine protease with proline specificity from ginger rhizome, Zingiber officinale". Biochemistry 38 (36): 11624–33. 1999. doi:10.1021/bi990651b. PMID 10512617.
- ↑ "Purification, characterization, and milk coagulating properties of ginger proteases". J. Dairy Sci. 94 (5): 2259–69. 2011. doi:10.3168/jds.2010-4024. PMID 21524515.
- ↑ 5.0 5.1 喜美代, 道; 初世, 佐々; 芳江, 市川 (1973). "ショウガたん白分解酵素の分離精製" (in ja). 栄養と食糧 26 (6): 377–383. doi:10.4327/jsnfs1949.26.377. ISSN 1883-8863.
- ↑ 6.0 6.1 6.2 "Modes of inhibition of cysteine proteases". Acta Biochimica Polonica 51 (4): 861–73. 2004. PMID 15625558.
- ↑ 7.0 7.1 7.2 "Ginger Rhizome: A New Source of Proteolytic Enzyme". J. Food Sci. 38 (4): 652–655. 1973. doi:10.1111/j.1365-2621.1973.tb02836.x.
- ↑ 8.0 8.1 8.2 8.3 "Tenderization of Meat with Ginger Rhizome Protease". J. Food Sci. 51 (6): 1558–1559. 1986. doi:10.1111/j.1365-2621.1986.tb13860.x.
- ↑ 9.0 9.1 9.2 "Multifunctional role of plant cysteine proteinases". Acta Biochimica Polonica 51 (3): 609–24. 2004. doi:10.18388/abp.2004_3547. PMID 15448724.
- ↑ "Characterisation of commercial papain, bromelain, actinidin and zingibain protease preparations and their activities toward meat proteins". Food Chem. 134 (1): 95–105. 2012. doi:10.1016/j.foodchem.2012.02.071.
- ↑ 11.0 11.1 11.2 "Plant collagenase: unique collagenolytic activity of cysteine proteases from ginger". Biochimica et Biophysica Acta (BBA) - General Subjects 1770 (12): 1627–35. 2007. doi:10.1016/j.bbagen.2007.08.003. PMID 17920199.
- ↑ "Effect of Proteolytic Enzymes and Ginger Extract on Tenderization of M. pectoralis profundus from Holstein Steer". Korean Journal for Food Science of Animal Resources 38 (1): 143–151. 2018. doi:10.5851/kosfa.2018.38.1.143. PMID 29725232.
- ↑ 13.0 13.1 "Ginger rhizome as a potential source of milk coagulating cysteine protease". Phytochemistry 72 (6): 458–64. 2011. doi:10.1016/j.phytochem.2010.12.002. PMID 21353685. Bibcode: 2011PChem..72..458H.
- ↑ 14.0 14.1 14.2 14.3 14.4 14.5 "Stabilization and Partial Purification of a Protease from Ginger Rhizome (Zingiber offinale Roscoe)". J. Food Sci. 70 (3): C231–C234. 2005. doi:10.1111/j.1365-2621.2005.tb07130.x.
External links
- Zingipain at the US National Library of Medicine Medical Subject Headings (MeSH)
 |
