Biology:Genetic history of North Africa
The genetic history of North Africa encompasses the genetic history of the people of North Africa. The most important source of gene flow to North Africa was from the Middle East, although the Sahara desert to the south and the Mediterranean Sea to the North were also important barriers to gene flow from sub-Saharan Africa and parts of Europe in prehistory. However, North Africa is connected to Western Asia via the Isthmus of Suez and the Sinai peninsula, while at the Straits of Gibraltar, North Africa and Europe are separated by only 15 km (9 mi), similarly Malta, Sicily, Canary Islands, and Crete are close to the coasts of North Africa.
North Africa is a genetically heterogenous and diverse region, and is characterized by its diverse ethnic groups, the main ones being Arabs, Berbers and Copts (in Egypt).[1] North African populations show a complex and heterogeneous genetic structure that has been described as an amalgam of at least four different ancestral components from the Middle East, sub-Saharan Africa, Europe and indigenous North Africans.[2] Although North Africa has experienced gene flows from the surrounding regions, it has also experienced long periods of genetic isolation.[3] Some genetic studies have been criticised for their interpretation and categorisation of African genetic data.[4][5][6][7][8][9][10]
Current scientific debate is concerned with determining the relative contributions of different periods of gene flow to the current gene pool of North Africans. Anatomically modern humans are known to have been present in North Africa during the Middle Paleolithic (300,000 years ago), as attested by the by Jebel Irhoud 1.[11] Without morphological discontinuity, the Aterian was succeeded by the Iberomaurusian industry, whose lithic assemblages bore close relations with the Cro-Magnon cultures of Europe and Western Asia, rather than to the contemporary cultures of sub-Saharan Africa or the Horn of Africa.[12] The Iberomaurusian industry was succeeded by the heavily West Asian influenced Capsian industry (8000 BCE to 2700 BCE) in the eastern part of North Africa (Egypt, Libya, Tunisia, eastern Algeria, Malta).
After migrating to North Africa in the 1st millennium BC, Phoenician settlers from the cities of Tyre and Sidon established over 300 coastal colonies throughout the region and built a powerful empire that controlled most of the region from the 8th century BC until the middle of the 2nd century BC.[13] A recent study has found that nearly 40% of modern Maghrebi males carry the paternal marker E-M81, which is thought to have expanded from the Levant into North Africa.[14]
In the 7th century A.D., the region was conquered by Muslim Umayyad Arabs from the Arabian Peninsula. Under the relatively brief Arab-Umayyad conquest and the later arrival of nomadic Bedouin, Levantine Arabs and Arabized peoples from the Near East in Western Asia and the arrival of some Sephardi Jews and Iberian Muslims fleeing the Spanish Catholic Reconquista of Iberia, a partial population mix or fusion have taken place and have resulted in some genetic diversity among some North Africans.[15]
A recent study from 2017 suggested that the Arab migrations to the Maghreb was mainly a demographic process that heavily implied gene flow and remodeled the genetic structure of the Maghreb, rather than a mere cultural replacement as claimed by older studies.[16] Another study found out that the majority of J-M267 (Eu10) chromosomes in the Maghreb are due to the recent gene flow caused by the Arab migrations to the Maghreb in the first millennium CE as both southern Qahtanite and northern Adnanite Arabs added to the heterogenous Maghrebi ethnic melting pot. The Eu10 chromosome pool in the Maghreb is derived not only from early Neolithic dispersions but to a much greater extent from recent expansions of Arab tribes from the Arabian Peninsula.[17]
Y-DNA of ancient North African samples
| Name | Country | Culture | Date
(Before Present) |
Y-DNA Haplogroup | Genetic Profile | Study |
|---|---|---|---|---|---|---|
| TAF009 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | |
| TAF010 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | Loosdrecht et al. 2018[18] |
| TAF011 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | Loosdrecht et al. 2018[18] |
| TAF012 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | Loosdrecht et al. 2018[18] |
| TAF013 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | Loosdrecht et al. 2018[18] |
| TAF015 | Morocco | Iberomaurusian | 15,100-13,900 | E-M78 | Afro-Levantine | Loosdrecht et al. 2018[18] |
| KTG004 | Morocco | Cardial | 7,400-6,900 | G2 | European | Simoes et al. 2023[19] |
| KTG006 | Morocco | Cardial | 7,400-6,900 | G2 | European | Simoes et al. 2023[19] |
| IAM.4 | Morocco | Cardial | 7,300-6,700 | E-L19 | Levantine | Fregel et al. 2018[20] |
| IAM.5 | Morocco | Cardial | 7,300-6,700 | E-L19 | Levantine | Fregel et al. 2018[20] |
| SKH002 | Morocco | Proto-Beaker | 6,700-6,100 | T | Levantine | Simoes et al. 2023[19] |
| SKH005 | Morocco | Proto-Beaker | 6,700-6,100 | T | Levantine | Simoes et al. 2023[19] |
| KEB.6 | Morocco | Proto-Beaker | 5,700-5,600 | T | European | Fregel et al. 2018[20] |
| KEB.7 | Morocco | Proto-Beaker | 5,700-5,600 | CT | European | Fregel et al. 2018[20] |
| Nakht-Ankh | Egypt | Egyptian | 4,000 | H | Unknown | Drosou et al. 2018[21] |
| Tuthmoses 1 | Egypt | Egyptian | 3,450 | J1 | Unknown | Gad et al. 2021[22] |
| Yuya | Egypt | Egyptian | 3,390 | G2 | Unknown | Gad et al. 2021[22] |
| Amenhotep III | Egypt | Egyptian | 3,370 | R1b | Unknown | Gad et al. 2021[22] |
| Akhenaten | Egypt | Egyptian | 3,350 | R1b | Unknown | Gad et al. 2021[22] |
| Tutankhamun | Egypt | Egyptian | 3,340 | R1b | Unknown | Gad et al. 2021[22] |
| Ramses III | Egypt | Egyptian | 3,200 | E-V38 | Unknown | Gad et al. 2021[22] |
| Pentawer | Egypt | Egyptian | 3,200 | E-V38 | Unknown | Gad et al. 2021[22] |
| JK2134 | Egypt | Egyptian | 2,798-2,591 | J1 | Levantine | Schuenemann et al. 2017[23] |
| JK2911 | Egypt | Egyptian | 2,791-2,582 | J2 | Levantine | Schuenemann et al. 2017[23] |
| R11793 | Tunisia | Phoenician | 2,711-2,355 | J2 | European | Moots et al. 2023[24] |
| R11746 | Tunisia | Phoenician | 2,683-2,361 | R1b | European | Moots et al. 2023[24] |
| R11751 | Tunisia | Phoenician | 2,678-2,359 | J2 | European | Moots et al. 2023[24] |
| R11753 | Tunisia | Phoenician | 2,606-2,355 | J2 | European | Moots et al. 2023[24] |
| JK2888 | Egypt | Egyptian | 2,119-2,024 | E-V22 | Levantine | Schuenemann et al. 2017[23] |
| R10766 | Algeria | Berber | 1,887-1,746 | G2 | Berber | Antonio et al. 2022[25] |
Y-chromosome
Previous works with Y-chromosome markers have shown that North Africa is highly heterogenous, the Y lineages found in the region are: A, B, E-V38, E-M78, E-M81, E-M123, G, F, H, I1, I2, J1, J2 and R1b. E-M81 reaches an average frequency of 40% across the region.[26] J-M267 is the second most-frequent haplogroup, accounting for around 30% of North Africans and assumed to have spread out of the Caucasus, Turkey and Iran into North Africa.[27]
Haplogroup E is thought to have emerged in the Paleolithic Levant, and would have later dispersed into North Africa.[28] Common subclades include E1b1b1a, E1b1b1b and E1b1b1*. E1b1b1b is distributed along a west-to-east cline with frequencies that can reach as high as 80 per cent among Hassani Arabs and 1% among Siwa Berbers. E1b1b1a has been observed at low to moderate frequencies among Berber populations with significantly higher frequencies observed in Northeast Africa relative to Northwest Africa.[29][30][31] Loosdrecht et al. 2018 demonstrated that E1b1b is most likely indigenous to Levant and migrated from Levant to North Africa during the Paleolithic.[3]
Haplogroup J-M267 is another very common haplogroup in the Maghreb, being the second most-frequent haplogroup after haplogroup E.[17] Haplogroup R1 has also been observed at moderate frequencies. A thorough study by Arredi et al. (2004), which analyzed populations from Algeria, concludes that the North African pattern of Y-chromosomal variation (including both J1 and E1b1b main haplogroups) is largely of Middle Eastern origin, which suggests that their introduction in this part of the world was the result of a migration of recent Semitic pastoralists from the Middle East,[29] although later papers have suggested that this date could have been as long as ten thousand years ago, with the transition from the Oranian to the Capsian culture in North Africa.[32] Another study concluded that the J-M267 chromosome pool in the Maghreb is derived not only from early Neolithic dispersions but to a much greater extent from recent expansions of Arab tribes from the Arabian Peninsula during the Arab migrations to the Maghreb.[17]
Keita (2008) examined a published Y-chromosome dataset on Afro-Asiatic populations and found that a key lineage E-M35/E-M78, sub-clade of haplogroup E, was shared between the populations in the locale of original Egyptian and Libyan speakers and modern Cushitic speakers from the Horn. These lineages are present in Egyptians, Berbers, Cushitic speakers from the Horn of Africa, and Semitic speakers in the Near-East. He noted that variants are also found in the Aegean and Balkans, but the origin of the M35 subclade mutation was in Egypt or Libya, and its clades were dominant in a core portion of Afro-Asiatic speaking populations which included Cushitic, Egyptian and Berber groups, in contrast Semitic speakers showed a decline in frequency going west to east in the Levantine-Syria region. Keita identified high frequencies of M35 (>50%) among Omotic populations, but stated that this derived from a small, published sample of 12. Keita also wrote that the PN2 mutation was shared by M35 and M2 lineages and this paternal clade originated from East Africa. He concluded that "the genetic data give population profiles that clearly indicate males of African origin, as opposed to being of Asian or European descent" but acknowledged that the biodiversity does not indicate any specific set of skin colors or facial features as populations were subject to microevolutionary pressures.[33]
E1b1b1b (Haplogroup E-M81); formerly E3b1b, E3b2
E1b1b1b (E-M81) is the most common Y chromosome haplogroup in North Africa, dominated by its sub-clade E-M183. It might have originated in the Levant around 5,600 years ago.[34][14] The parent clade, E1b1b, has been found among Prehistoric Levantines and North-Africans.[28][35] This haplogroup has been found at high levels in Medieval Arabs from Al-Andalus and among Canary islands, with lower but significant amounts also in Portugal, Southern Spain, Southern Italy and the Provence region of France .[36] As a result of European colonization, E-M81 is found in parts of Latin America,[37] among Hispanic in USA.[38] This sub-clade of E1b1b1 has also been observed at 40 per cent in the comarca Pasiegos from Cantabria.[28]
In smaller numbers, E-M81 men can be found in Malta, Northern Sudan, Cyprus and among Sephardic Jews.
There are two recognized sub-clades, although one is much more common than the other.
- Sub clades of E1b1b1b (E-M81):
- E1b1b1b1 (E-M107). Underhill et al. (2000) found one example among Malian Arabs.
- E1b1b1b2 (E-M165). Underhill et al. (2000) found one example among Israeli Arabs.
- E1b1b1b2b (E-M183). Most individuals belong to this subclade.[39]
The general parent Y-chromosome Haplogroup E1b1b (formerly known as E3b), which might have originated in the Levant and is by far the most common clade in North and Northeast Africa and found in some populations in Europe, particularly in the Mediterranean and South Eastern Europe. E1b1b reaches Greece, Malta and the Balkan region in Europe but is not as high there as it is among North African populations.[29] Outside of North and Northeast Africa, E1b1b's two most prevalent clades are E1b1b1a (E-M78, formerly E3b1a) and E1b1b1b (E-M81, formerly E3b1b).
A study from Semino (published 2004) showed that Y-chromosome haplotype E1b1b1b (E-M81), is specific to North African populations and almost absent in Europe, Western Asia and sub-Saharan Africa, except Iberia (Spain , Portugal, Gibraltar, Andorra) and Sicily.[29] Another 2004 study showed that E1b1b1b is found present, albeit at low levels throughout Southern Europe (ranging from 1.5 per cent in Northern Italians, 2.2 per cent in Central Italians, 1.6 per cent in Southern Spaniards, 3.5 per cent in the French, 4 per cent in the Northern Portuguese, 12.2 per cent in the Southern Portuguese and 41.2 per cent in the genetic isolate of the Pasiegos from Cantabria in Italy).[28]
The findings of this latter study contradict a more thorough analysis Y-chromosome analysis of the Iberian peninsula according to which haplogroup E1b1b1b surpasses frequencies of 10 per cent in Southern Spain and southern Portugal. The study points only to a very limited influence from Northern Africa and West Asia in Iberia, both in historic and prehistoric times.[36][29][40]
A wide-ranging study (published 2007) using 6,501 unrelated Y-chromosome samples from 81 populations found that: "Considering both these E-M78 sub-haplogroups (E-V12, E-V22, E-V65) and the E-M81 haplogroup, the contribution of Northern African lineages to the entire male gene pool of Iberia (barring Pasiegos), continental Italy and Sicily can be estimated as 5.6 percent, 3.6 percent and 6.6 percent, respectively."[41] It has also been argued that the European distribution of E-M78 and its sub-clades is compatible with the Neolithic demic diffusion of agriculture, but also possibly partly from at least, the Mesolithic. For example, Battaglia et al. (2008) estimated that E-M78 (called E1b1b1a1 in that paper) has been in Europe longer than 10,000 years. In support of this theory, human remains excavated in a Spanish funeral cave dating from approximately 7,000 years ago were shown to be in this haplogroup.[42] More recently, two E-M78 have been found in the Neolithic Sopot and Lengyel cultures from the same period,[43][44] which seems supported by the most recent studies (including autosomal research).
A very recent study about Sicily by Gaetano et al. 2008 found that "The Hg E3b1b-M81, widely diffused in northwestern African populations, is estimated to contribute to the Sicilian gene pool at a rate of 6 percent."[45]
According to the most recent and thorough study about Iberia by Adams et al. 2008 that analysed 1,140 unrelated Y-chromosome samples in Iberia, a limited contribution of northern African lineages to the entire male gene pool of Iberia was found : "mean North African admixture is just 10.6 percent, with wide geographical variation, ranging from zero in Gascony to 21.7 percent in Northwest Castile".[46]
J-M267 (Haplogroup J1)
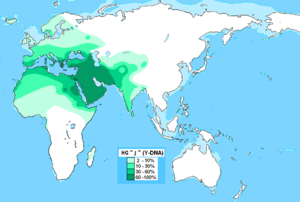
J-M267 (J1) is the second most common Y chromosome haplogroup in North Africa. It originated in the Middle East, and its highest frequency of 30%–62.5% has been observed in Muslim Arab populations in the Middle East.[17] A study found out that the majority of J1 (Eu10) chromosomes in the Maghreb are due to the recent gene flow caused by the Arab migrations to the Maghreb in the first millennium CE. The J-M267 chromosome pool in the Maghreb is derived not only from early Neolithic dispersions but to a much greater extent from recent expansions of Arab tribes from the Arabian Peninsula, during which both southern Qahtanite and northern Adnanite Arabs added to the heterogenous Maghrebi ethnic melting pot.[17] A study from 2017 suggested that these Arab migrations were a demographic process that heavily implied gene flow and remodeled the genetic structure of the Maghreb, rather than a mere cultural replacement as claimed by older studies.[1]
Recent genome-wide analysis of North Africans found substantial shared ancestry with the Middle East, and to a lesser extent sub-Saharan Africa and Europe. The recent gene flow caused by the Arab migrations to the Maghreb increased genetic similarities between North Africans and Middle Easterners.[47] Haplogroup J1-M267 accounts for around 30% of North Africans and has spread from the Arabian Peninsula, second after E1b1b1b which accounts for 45% of North Africans. A study from 2021 has shown that the highest frequency of the Middle Eastern component ever observed in North Africa so far was observed in the Arabs of Wesletia in Tunisia, who had a Middle Eastern component frequency of 71.8%.[48] According to a study from 2004, Haplogroup J1 had a frequency of 35% in Algerians and 34.2% in Tunisians.[49]
Mitochondrial DNA
Individuals receive mtDNA only from their mothers. According to Macaulay et al. 1999, "one-third (33%) of Mozabite Berber mtDNAs have a Near Eastern ancestry, probably having arrived in North Africa less than 50,000 years ago, and one-eighth (12.5%) have an origin in sub-Saharan Africa. Europe appears to be the source of many of the remaining sequences, with the rest having arisen either in Europe or in the Near East".[50] Maca-Meyer et al. 2003 analyze the "autochthonous North African lineage U6" in mtDNA, and conclude that:
- The most probable origin of the proto-U6 lineage was the Near East. Around 30,000 years ago it spread to North Africa where it represents a signature of regional continuity. Subgroup U6a reflects the first North African expansion from the Maghreb returning to the east in Paleolithic times. Derivative clade U6a1 signals a posterior movement from Western Asia back to the Maghreb and North Africa. This migration coincides with a possible Afroasiatic linguistic expansion.
A genetic study by Fadhlaoui-Zid et al. 2004[51] argues concerning certain exclusively North African haplotypes that "expansion of this group of lineages took place around 10,500 years ago in North Africa, and spread to neighbouring population", and apparently that a specific Northwestern African haplotype, U6, probably originated in the Near East 30,000 years ago accounts for 28 per cent in Mozabites, 18 per cent in Kabyles, but only accounts for 6-8 per cent in the southern Moroccan Berbers. Rando et al. 1998 "detected female-mediated gene flow from sub-Saharan Africa to NW Africa" amounting to as much as 21.5 per cent of the mtDNA sequences in a sample of NW African populations;[44] the amount varied from 82 per cent in Tuaregs to less than 3 per cent in Riffians in north of Morocco. This north-south gradient in the sub-Saharan contribution to the gene pool is supported by Esteban et al.[52]
Nevertheless, individual Berber communities display a considerably high mtDNA heterogeneity among them. The Berbers of Jerba Island, located in South Eastern Tunisia, display an 87 per cent West Eurasian contribution with no U6 haplotypes,[53] while the Kesra of Tunisia, for example, display a much higher proportion of typical sub-Saharan mtDNA haplotypes (49 per cent),[54] as compared to the Zriba (8 per cent). According to the article, "The North African patchy mtDNA landscape has no parallel in other regions of the world and increasing the number of sampled populations has not been accompanied by any substantial increase in our understanding of its phylogeography. Available data up to now rely on sampling small, scattered populations, although they are carefully characterized in terms of their ethnic, linguistic, and historical backgrounds. It is therefore doubtful that this picture truly represents the complex historical demography of the region rather than being just the result of the type of samplings performed so far."
A 2005 study discovered a close mitochondrial link between Berbers and the Uralic speaking Saami of northern Scandinavia and the sub-Arctic, and argues that Southwestern Europe and North Africa was the source of late-glacial expansions of hunter-gatherers that repopulated Northern Europe after a retreat south during the Last Glacial Maximum, and reveals a direct maternal link between those European hunter-gatherer populations and the Berbers.[55] With regard to Mozabite Berbers, one-third (33%) of Mozabite Berber mtDNAs have a Near Eastern ancestry, probably having arrived in North Africa ~50,000 years ago, and one-eighth (12.5%) have an origin in sub-Saharan Africa. Europe appears to be the source of many of the remaining sequences, with the rest (54.5%) having arisen either in Europe or in the Near East."[50]
According to the most recent and thorough study on Berber mtDNA from Coudray et al. 2008, which analysed 614 individuals from 10 different regions (Morocco (Asni, Bouhria, Figuig, Souss), Algeria (Mozabites), Tunisia (Chenini-Douiret, Sened, Matmata, Jerba) and Egypt (Siwa)),[56] the results may be summarized as follows:
- Total West Eurasian lineages (H, HV, R0, J, M, T, U, K, N1, N2, X) : 80 per cent
- Total African lineages (L0, L1, L2, L3, L4, L5) : 20 per cent
The Berber mitochondrial pool is characterized by an overall high frequency of Western Eurasian haplogroups, a markedly lower frequency of sub-Saharan L lineages, and a significant (but differential) presence of North African haplogroups U6 and M1.
There is a degree of dispute about when and how the minority sub-Saharan L haplogroups entered the North African gene pool. Some papers suggest that the distribution of the main L haplogroups in North Africa was mainly due to the Islamic era trans-Saharan slave trade, as espoused by Harich et .al in a study conducted in 2010.[57] However, also in September 2010, a study of Berber mtDNA by Frigi et al. concluded that some of the L haplogroups were much older and introduced by an ancient African gene flow around 20,000 years ago.[58]
Genetic studies on Iberian populations also show that North African mitochondrial DNA sequences (haplogroup U6) and sub-Saharan sequences (Haplogroup L), although present at only low levels, are still at higher levels than those generally observed elsewhere in Europe, though very likely, most of the L mtDNA that has been found in minor amounts in Iberia, is actually pre-neolithic in origin, as it was demonstrated by María Cerezo et al., (Reconstructing ancient mitochondrial DNA links between Africa and Europe) and U6 too, which also have a very old presence in Iberia, since Iberia has a great diversity in lineages from this haplogroup, it was already found in some local hunter-gatherer remains and its local geographic distribution is not compatible, in many cases, with Moor occupation area.[59][60][61] Haplogroup U6 have also been detected in Sicily and Southern Italy at much lower frequencies.[62] It happens also to be a characteristic genetic marker of the Saami populations of Northern Scandinavia.[55]
It is difficult to ascertain that U6's presence is the consequence of Islam's expansion into Europe during the Middle Ages, particularly because it is more frequent in the west of the Iberian Peninsula rather than in the east. In smaller numbers it is also attested in the British Isles, again in its northern and western borders. It may be a trace of a prehistoric Neolithic/Megalithic/Mesolithic or even Upper Paleolithic expansion along the Atlantic coasts from North Africa or Iberian Peninsula, perhaps in conjunction with seaborne trade, although an alternative, but less likely explanation, would attribute this distribution in Northern Britain to the Roman period. One subclade of U6 is particularly common among Canarian Spaniards as a result of native Guanche (proto-Berber) ancestry.[citation needed]
Autosomal DNA
On 13 January 2012, an exhaustive genetic study of North Africa's human populations was published in PLoS Genetics and was undertaken jointly by researchers in the Evolutionary Biology Institute (CSIC-UPF) and Stanford University, among other institutions.[63]
The study highlights the complex genetic makeup of North Africa. This genetic composition shows a significant local component that became more distinct around 12,000 years ago, possibly influenced by migrations, population expansions, or other demographic events. According to David Comas, coordinator of the study and researcher at the Institute for Evolutionary Biology (CSIC-UPF), "some of the questions we wanted to answer were whether today's inhabitants are direct descendants of the populations with the oldest archaeological remains in the region, dating back fifty thousand years, or whether they are descendants of the Neolithic populations in the Middle East, which introduced agriculture to the region around nine thousand years ago. We also wondered if there had been any genetic exchange between the North African populations and the neighbouring regions and if so, when these took place".[64]
To explore these questions, the research team analyzed nearly 800,000 genetic markers across the entire genomes of 125 North African individuals from seven representative populations. This data was then juxtaposed with information from neighboring populations.[64]
The findings reveal a distinct native genetic component in North Africans, setting them apart from sub-Saharan Africans and aligning them more closely with West-Eurasians, primarily Middle Easterners and Europeans. Though the study emphasizes a dominant genetic lineage in contemporary North Africans tracing back to around 12,000 years ago, it doesn't dismiss the likelihood of genetic continuity from ancient human groups present in North Africa over 60,000 years ago. The data suggests that while ancient human groups indeed inhabited the region, the majority of the modern identifiable genetic makeup stems from more recent periods. The unique North African (Maghrebi) genetic signature is distinct from ancestries found in the populations of sub-Saharan Africa. Modern North African populations were observed to share genetic markers in varying degrees with all the neighbouring regions (Southern Europe, West Asia, sub-Saharan Africa), probably as a result of more recent migrations.[64]
Hodgson et al. 2014 found a distinct non-African ancestry component among Northeastern Africans (dubbed "Ethio-Somali"), which split from other West-Eurasian ancestries, and is most closely related to the North African (Maghrebi), and Arabian ancestry components. Both would have entered Africa during a pre-agricultural period (between 12,000 to 23,000 years ago). This component is suggested to have been present in considerable amounts among the Proto-Afroasiatic-speaking peoples. The authors argue that the Ethio-Somali component and the Maghrebi component descended from a single ancestral lineage, which split from the Arabian lineage and migrated into Africa from the Middle East. That is, a common ancestral population migrated into Africa through Sinai and then split into two, with one branch continuing west across North Africa and the other heading south into the HOA.[65]
A 2015 study by Dobon et al. identified another ancestral autosomal component of West Eurasian origin that is common to many modern Afro-Asiatic-speaking populations in Northeast Africa. Known as the Coptic component, it peaks among Egyptian Copts, including those who settled in Sudan over the past two centuries. The Coptic component evolved out of a main North African and Middle Eastern ancestral component that is shared by other Egyptians and also found at high frequencies among other populations in Northern Africa. The scientists suggest that this points to a common origin for the general population of Egypt. They also associate the Coptic component with Ancient Egyptian ancestry, without the later minority Medieval Era Arabian and sub-Saharan African influence that is present among other Egyptians.[66]
According to a paper published in 2017, most of the genetic studies on North African populations agree with a limited correlation between genetics and geography, showing a high population heterogeneity in the region (without strong differences between Arabs and Berbers). Northern African populations have been described as a mosaic of North African (Taforalt), Middle Eastern, European (Early European Farmers) and sub-Saharan ancestries.[67][68]
Ancient North African samples such as the Paleolithic Taforalt, were found to be composed of two major components: a Holocene Levantine component, and from an indigenous Hadza/West African-like component. The Taforalt individuals show closest genetic affinity for ancient Epipaleolithic Natufian individuals, with slightly greater affinity for the Natufians than later Neolithic Levantines. A two-way admixture scenario using Levantine samples and modern West/East African samples as reference populations inferred that the Taforalt individuals bore 63.5% West-Eurasian Levantine-related and 36.5% sub-Saharan African-related ancestries, with no evidence for additional gene flow from the Epigravettian culture of Upper Paleolithic Europe. The Taforalt individuals also show evidence of limited Neanderthal ancestry.[69][70][71][72] A recent genetic study published in the "European Journal of Human Genetics" in Nature (2019) showed that Northern Africans are closely related to West Asians as well as Europeans. Northern Africans can be distinguished from West Africans and other African populations dwelling south of the Sahara.[73]
According to Lucas-Sánchez, Marcel et al. (2021) despite the geneflow from the Middle-East, Europe and sub-Saharan Africa, an autochthonous genetic component that dates back to pre-Holocene times is still present in North African groups. The analysis also showed as a whole no genetic pattern of differentiation between Tamazight (i.e. Berber) and Arabs.[74]
Ancient DNA
Unlike sub-Saharan Africans, North Africans have a similar level of Neanderthal DNA to South Europeans and West Asians, which is pre-Neolithic in origin, rather than via any later admixture with peoples from outside of North Africa during the historical period. It was found that modern North Africans derive mainly from a "back to Africa" population from Eurasia "from before 12,000 years ago (ya) (i.e., prior to the Neolithic migrations)" but more recent than 40,000 years ago which seems to "represent a genetic discontinuity with the earliest modern human settlers of North Africa (those with the Aterian industry).[75]
In 2013, Nature announced the publication of the first genetic study utilizing next-generation sequencing to ascertain the ancestral lineage of an Ancient Egyptian individual. The research was led by Carsten Pusch of the University of Tübingen in Germany and Rabab Khairat, who released their findings in the Journal of Applied Genetics. DNA was extracted from the heads of five Egyptian mummies that were housed at the institution. All the specimens were dated to between 806 BC and 124 AD, a timeframe corresponding with the Late Dynastic and Ptolemaic periods. The researchers observed that one of the mummified individuals likely belonged to the mtDNA haplogroup I2, a maternal clade that is believed to have originated in Western Asia.[76]
In 2013, Iberomaurusian skeletons from the prehistoric sites of Taforalt and Afalou in the Maghreb were analyzed for ancient DNA. All of the specimens belonged to maternal clades associated with either North Africa or the northern and southern Mediterranean littoral, indicating gene flow between these areas since the Epipaleolithic.[77] The ancient Taforalt individuals carried the mtDNA haplogroups U6, H, JT and V, which points to population continuity in the region dating from the Iberomaurusian period.[78]
The E1b1b-M81 (~44%), R-M269 (~44%), and E-M132/E1a (~6%) paternal haplogroups have been found in ancient Guanche (Bimbapes) fossils excavated in Punta Azul, El Hierro, Canary Islands, which are dated to the 10th century. Maternally, the specimens all belong to the H1 clade. These locally born individuals carried the H1-16260 haplotype, which is exclusive to the Canary Islands and Algeria.[79] In 2018, DNA analysis of Later Stone Age individuals from the site of Taforalt (Iberomaurusian, 15 000 BP) revealed that the Iberomaurusians carried haplogroup E-M78* and Cardial culture bearers from the site of Ifri N' Ammar (7 000 BP) carried the Levantine marker E-L19 indicating a break in continuity in the region. These studies confirmed a break in continuity in the region showing that Mesolithic Moroccans did not contribute paternally to Later Stone Age individuals and to present-day Maghrebi populations.[3][80]
A 2019 study seeking to determine if North Africans descend from strictly Palaeolithic groups (Taforalt), or subsequent migrations, discovered that most of the genetic variation in the region was shaped during the Neolithic. While the ancient samples had more of the Taforalt component, it is most frequent today in Western North Africans (Saharawi, Moroccans, Algerians) and Berbers, and suggested a continuity of this autochronous North African component. The consideration of Berber-speaking groups as the autochthonous peoples of North Africa was reinforced by these results.[81]
See also
- Genetic history of Africa
- Y-DNA haplogroups in populations of North Africa
- Y-DNA haplogroups in populations of the Near East
- Population history of Egypt
- Ethnic groups of North Africa
- African admixture in Europe
- Genetic studies on Jews
- Genetic studies on Arabs
- Genetic history of the Iberian Peninsula
- Genetic studies on Moroccans
- Genetic history of the Middle East
References
- ↑ 1.0 1.1 Arauna, Lara R.; Mendoza-Revilla, Javier; Mas-Sandoval, Alex; Izaabel, Hassan; Bekada, Asmahan; Benhamamouch, Soraya; Fadhlaoui-Zid, Karima; Zalloua, Pierre et al. (February 2017). "Recent Historical Migrations Have Shaped the Gene Pool of Arabs and Berbers in North Africa". Molecular Biology and Evolution 34 (2): 318–329. doi:10.1093/molbev/msw218. PMID 27744413.
- ↑ Arauna, L. R.; Hellenthal, G.; Comas, D. (May 2019). "Dissecting human North African gene-flow into its western coastal surroundings". Proceedings of the Royal Society B: Biological Sciences 286 (1902). doi:10.1098/rspb.2019.0471. PMID 31039721.
- ↑ 3.0 3.1 3.2 van de Loosdrecht, Marieke; Bouzouggar, Abdeljalil; Humphrey, Louise; Posth, Cosimo; Barton, Nick; Aximu-Petri, Ayinuer; Nickel, Birgit; Nagel, Sarah et al. (4 May 2018). "Pleistocene North African genomes link Near Eastern and sub-Saharan African human populations". Science 360 (6388): 548–552. doi:10.1126/science.aar8380. PMID 29545507. Bibcode: 2018Sci...360..548V.
- ↑ Lieberman, Leonard; Jackson, Fatimah Linda C. (1995). "Race and Three Models of Human Origin". American Anthropologist 97 (2): 231–242. doi:10.1525/aa.1995.97.2.02a00030. ISSN 0002-7294. https://www.jstor.org/stable/681958.
- ↑ "It is not clear to what degree certain genetic systems usually interpreted as non-African may in fact be native to Africa. Much depends on how "African" is defined and the model of interpretation. The various genetic studies usually suffer from what is called categorical thinking, specifically, racial thinking. Many investigators still think of "African" in a stereotyped, nonscientific (evolutionary) fashion, not acknowledging a range of genetic variants or traits as equally African".Celenko, Theodore (1996). "The Geographical Origins and Population Relationships of Early Ancient Egyptians" In Egypt in Africa. Indianapolis, Ind.: Indianapolis Museum of Art. pp. 20–33. ISBN 0936260645.
- ↑ Ryan A.Brown and George J. Armelagos (2001). "Apportionment of racial diversity: A review". Evolutionary Anthropology 10, Issue 1 (34–40): 34–40. doi:10.1002/1520-6505(2001)10:1<34::AID-EVAN1011>3.0.CO;2-P.
- ↑ Eltis, David; Bradley, Keith R.; Perry, Craig; Engerman, Stanley L.; Cartledge, Paul; Richardson, David (12 August 2021) (in en). The Cambridge World History of Slavery: Volume 2, AD 500-AD 1420. Cambridge University Press. p. 150. ISBN 978-0-521-84067-5. https://books.google.com/books?id=DskwEAAAQBAJ&dq=gourdine+critique+their+methods&pg=PA150.
- ↑ Candelora 2022, pp. 101–122.
- ↑ Keita, S. O. Y.; Kittles, Rick A. (1997). "The Persistence of Racial Thinking and the Myth of Racial Divergence". American Anthropologist 99 (3): 534–544. doi:10.1525/aa.1997.99.3.534. ISSN 0002-7294. https://www.jstor.org/stable/681741.
- ↑ Ehret, Christopher (20 June 2023) (in en). Ancient Africa: A Global History, to 300 CE. Princeton: Princeton University Press. pp. 83–86, 167–169. ISBN 978-0-691-24409-9. https://books.google.com/books?id=Q5KjEAAAQBAJ&q=ancient+africa:+a+global+history,+to+300+ce+christopher+ehret.
- ↑ Callaway, Ewen (7 June 2017). "Oldest Homo sapiens fossil claim rewrites our species' history". Nature. doi:10.1038/nature.2017.22114.
- ↑ Hublin, Jean-Jacques; McPherron, Shannon (2012-03-31) (in en). Modern Origins: A North African Perspective. Springer Science & Business Media. pp. 180. ISBN 9789400729285. https://books.google.com/books?id=bI-egtJvrn8C&pg=PA180.
- ↑ Woolmer, Mark (2017-04-30) (in en). A Short History of the Phoenicians. Bloomsbury Publishing. pp. 201. ISBN 978-1-78672-217-1. https://books.google.com/books?id=lbmKDwAAQBAJ&dq=300+colonies+phoenician+strabo&pg=PA201.
- ↑ 14.0 14.1 Penninx, Wim. The male lines of the Maghreb: Phoenicians, Carthage, Muslim conquest and Berbers. https://www.academia.edu/39673436.
- ↑ Rando, J. C.; Pinto, F.; Gonzalez, A. M.; Hernandez, M.; Larruga, J. M.; Cabrera, V. M.; Bandelt, H.-J. (November 1998). "Mitochondrial DNA analysis of Northwest African populations reveals genetic exchanges with European, Near-Eastern, and sub-Saharan populations". Annals of Human Genetics 62 (6): 531–550. doi:10.1046/j.1469-1809.1998.6260531.x. PMID 10363131.
- ↑ Arauna, Lara R.; Mendoza-Revilla, Javier; Mas-Sandoval, Alex; Izaabel, Hassan; Bekada, Asmahan; Benhamamouch, Soraya; Fadhlaoui-Zid, Karima; Zalloua, Pierre et al. (February 2017). "Recent Historical Migrations Have Shaped the Gene Pool of Arabs and Berbers in North Africa". Molecular Biology and Evolution 34 (2): 318–329. doi:10.1093/molbev/msw218. PMID 27744413.
- ↑ 17.0 17.1 17.2 17.3 17.4 Nebel, Almut; Landau-Tasseron, Ella; Filon, Dvora; Oppenheim, Ariella; Faerman, Marina (June 2002). "Genetic Evidence for the Expansion of Arabian Tribes into the Southern Levant and North Africa". The American Journal of Human Genetics 70 (6): 1594–1596. doi:10.1086/340669. PMID 11992266.
- ↑ 18.0 18.1 18.2 18.3 18.4 Cite error: Invalid
<ref>tag; no text was provided for refs namedvan de Loosdrecht 20182 - ↑ 19.0 19.1 19.2 19.3 Simões, Luciana G.; Günther, Torsten; Martínez-Sánchez, Rafael M.; Vera-Rodríguez, Juan Carlos; Iriarte, Eneko; Rodríguez-Varela, Ricardo; Bokbot, Youssef; Valdiosera, Cristina et al. (June 2023). "Northwest African Neolithic initiated by migrants from Iberia and Levant" (in en). Nature 618 (7965): 550–556. doi:10.1038/s41586-023-06166-6. ISSN 1476-4687. PMID 37286608. Bibcode: 2023Natur.618..550S.
- ↑ 20.0 20.1 20.2 20.3 Fregel, Rosa; Méndez, Fernando L.; Bokbot, Youssef; Martín-Socas, Dimas; Camalich-Massieu, María D.; Santana, Jonathan; Morales, Jacob; Ávila-Arcos, María C. et al. (2018-06-26). "Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe". Proceedings of the National Academy of Sciences of the United States of America 115 (26): 6774–6779. doi:10.1073/pnas.1800851115. ISSN 0027-8424. PMID 29895688. Bibcode: 2018PNAS..115.6774F.
- ↑ Drosou, Konstantina; Price, Campbell; Brown, Terence A. (2018-02-01). "The kinship of two 12th Dynasty mummies revealed by ancient DNA sequencing". Journal of Archaeological Science: Reports 17: 793–797. doi:10.1016/j.jasrep.2017.12.025. ISSN 2352-409X. Bibcode: 2018JArSR..17..793D. https://www.sciencedirect.com/science/article/pii/S2352409X17305631.
- ↑ 22.0 22.1 22.2 22.3 22.4 22.5 22.6 "Error: no
|title=specified when using {{Cite web}}". https://bia.unibz.it/esploro/outputs/bookChapter/Maternal-and-Paternal-Lineages-in-King/991005930750801241. - ↑ 23.0 23.1 23.2 Schuenemann, Verena J.; Peltzer, Alexander; Welte, Beatrix; van Pelt, W. Paul; Molak, Martyna; Wang, Chuan-Chao; Furtwängler, Anja; Urban, Christian et al. (2017-05-30). "Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods". Nature Communications 8 (1): 15694. doi:10.1038/ncomms15694. ISSN 2041-1723. PMID 28556824. Bibcode: 2017NatCo...815694S.
- ↑ 24.0 24.1 24.2 24.3 Moots, Hannah M.; Antonio, Margaret; Sawyer, Susanna; Spence, Jeffrey P.; Oberreiter, Victoria; Weiß, Clemens L.; Lucci, Michaela; Cherifi, Yahia Mehdi Seddik et al. (September 2023). "A genetic history of continuity and mobility in the Iron Age central Mediterranean" (in en). Nature Ecology & Evolution 7 (9): 1515–1524. doi:10.1038/s41559-023-02143-4. ISSN 2397-334X. PMID 37592021. Bibcode: 2023NatEE...7.1515M. https://www.nature.com/articles/s41559-023-02143-4.
- ↑ Antonio, Margaret; Weiß, Clemens; Gao, Ziyue; Sawyer, Susanna; Oberreiter, Victoria; Moots, Hannah; Spence, Jeffrey; Cheronet, Olivia et al. (2023). Stable population structure in Europe since the Iron Age, despite high mobilit (Report). doi:10.1101/2022.05.15.491973. https://doi.org/10.1101/2022.05.15.491973.
- ↑ Arredi, Barbara; Poloni, Estella S.; Paracchini, Silvia; Zerjal, Tatiana; Fathallah, Dahmani M.; Makrelouf, Mohamed; Pascali, Vincenzo L.; Novelletto, Andrea et al. (August 2004). "A predominantly neolithic origin for Y-chromosomal DNA variation in North Africa". American Journal of Human Genetics 75 (2): 338–345. doi:10.1086/423147. ISSN 0002-9297. PMID 15202071.
- ↑ Sahakyan, Hovhannes; Margaryan, Ashot; Saag, Lauri; Karmin, Monika; Flores, Rodrigo; Haber, Marc; Kushniarevich, Alena; Khachatryan, Zaruhi et al. (2021-03-23). "Origin and diffusion of human Y chromosome haplogroup J1-M267" (in en). Scientific Reports 11 (1): 6659. doi:10.1038/s41598-021-85883-2. ISSN 2045-2322. PMID 33758277. Bibcode: 2021NatSR..11.6659S.
- ↑ 28.0 28.1 28.2 28.3 Cruciani, Fulvio; La Fratta, Roberta; Santolamazza, Piero; Sellitto, Daniele; Pascone, Roberto; Moral, Pedro; Watson, Elizabeth; Guida, Valentina et al. (May 2004). "Phylogeographic Analysis of Haplogroup E3b (E-M215) Y Chromosomes Reveals Multiple Migratory Events Within and Out Of Africa". The American Journal of Human Genetics 74 (5): 1014–1022. doi:10.1086/386294. PMID 15042509.
- ↑ 29.0 29.1 29.2 29.3 29.4 Semino, Ornella; Magri, Chiara; Benuzzi, Giorgia; Lin, Alice A.; Al-Zahery, Nadia; Battaglia, Vincenza; Maccioni, Liliana; Triantaphyllidis, Costas et al. (May 2004). "Origin, Diffusion, and Differentiation of Y-Chromosome Haplogroups E and J: Inferences on the Neolithization of Europe and Later Migratory Events in the Mediterranean Area". The American Journal of Human Genetics 74 (5): 1023–1034. doi:10.1086/386295. PMID 15069642.
- ↑ Kujanová, Martina; Pereira, Luísa; Fernandes, Verónica; Pereira, Joana B.; Černý, Viktor (October 2009). "Near Eastern Neolithic genetic input in a small oasis of the Egyptian Western Desert". American Journal of Physical Anthropology 140 (2): 336–346. doi:10.1002/ajpa.21078. PMID 19425100.
- ↑ Fadhlaoui-Zid, Karima; Martinez-Cruz, Begoña; Khodjet-el-khil, Houssein; Mendizabal, Isabel; Benammar-Elgaaied, Amel; Comas, David (October 2011). "Genetic structure of Tunisian ethnic groups revealed by paternal lineages". American Journal of Physical Anthropology 146 (2): 271–280. doi:10.1002/ajpa.21581. PMID 21915847.
- ↑ Myles, Sean; Bouzekri, Nourdine; Haverfield, Eden; Cherkaoui, Mohamed; Dugoujon, Jean-Michel; Ward, Ryk (1 June 2005). "Genetic evidence in support of a shared Eurasian-North African dairying origin". Human Genetics 117 (1): 34–42. doi:10.1007/s00439-005-1266-3. PMID 15806398.
- ↑ Keita, S.O.Y. (ed Bengston, John) (3 December 2008) (in en). "Geography, selected Afro-Asiatic families, and Y Chromosome lineage variation: An exploration in linguistics and phylogeography" in In Hot Pursuit of Language in Prehistory: Essays in the four fields of anthropology. In honor of Harold Crane Fleming. John Benjamins Publishing. pp. 3–15. ISBN 978-90-272-8985-8. https://books.google.com/books?id=MV46AAAAQBAJ&dq=While+it+is+known+and+must+be+emphasised+that+language%2C+biology%2C+and+culture+do+not+travel+as+an+obligatory+package%2C+this+does+not+mean+that+congruence+never+exists+and+may+be+historically+meaningful.+In+this+paper+the+published+Y+chromosome+data+and+branches+of+the+Afro-Asiatic+language+family+for+which+good+genetic+data+exist+were+examined+in+an+exploratory+fashion+to+determine+if+there+are+patterns+suggestive+of+overlap.+Most+of+the+Afro-Asiatic+speakers+shared+the+lineage+defined+by+Yap+descendant+called+PN2%2F215%2FM35.+Keita&pg=PA3.
- ↑ Arredi, Barbara; Poloni, Estella S.; Paracchini, Silvia; Zerjal, Tatiana; Fathallah, Dahmani M.; Makrelouf, Mohamed; Pascali, Vincenzo L.; Novelletto, Andrea et al. (August 2004). "A Predominantly Neolithic Origin for Y-Chromosomal DNA Variation in North Africa". American Journal of Human Genetics 75 (2): 338–345. doi:10.1086/423147. ISSN 0002-9297. PMID 15202071.
- ↑ Arredi, Barbara; Poloni, Estella S.; Paracchini, Silvia; Zerjal, Tatiana; Fathallah, Dahmani M.; Makrelouf, Mohamed; Pascali, Vincenzo L.; Novelletto, Andrea et al. (August 2004). "A Predominantly Neolithic Origin for Y-Chromosomal DNA Variation in North Africa". The American Journal of Human Genetics 75 (2): 338–345. doi:10.1086/423147. PMID 15202071.
- ↑ 36.0 36.1 Flores, Carlos; Maca-Meyer, Nicole; González, Ana M.; Oefner, Peter J.; Shen, Peidong; Pérez, Jose A.; Rojas, Antonio; Larruga, Jose M. et al. (October 2004). "Reduced genetic structure of the Iberian peninsula revealed by Y-chromosome analysis: implications for population demography". European Journal of Human Genetics 12 (10): 855–863. doi:10.1038/sj.ejhg.5201225. PMID 15280900.
- ↑ See the remarks of genetic genealogist Robert Tarín for example. We can add 6.1 per cent (8 out of 132) in Cuba, Mendizabal et al. (2008); 5.4 per cent (6 out of 112) in Brazil (Rio de Janeiro), "The presence of chromosomes of North African origin (E3b1b-M81; Cruciani et al., 2004) can also be explained by a Portuguese-mediated influx, since this haplogroup reaches a frequency of 5.6 per cent in Portugal (Beleza et al., 2006), quite similar to the frequency found in Rio de Janeiro (5.4 per cent) among European contributors.", Silva et al. (2006)[verification needed]
- ↑ 2.4 per cent (7 out of 295) among Hispanic men from California and Hawaii, Paracchini et al. (2003)[verification needed]
- ↑ Y-DNA Haplogroup E and its Subclades - 2008
- ↑ Gonçalves, Rita; Freitas, Ana; Branco, Marta; Rosa, Alexandra; Fernandes, Ana T.; Zhivotovsky, Lev A.; Underhill, Peter A.; Kivisild, Toomas et al. (July 2005). "Y-chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry: Y-chromosome Lineages in Portugal and the Atlantic Islands". Annals of Human Genetics 69 (4): 443–454. doi:10.1111/j.1529-8817.2005.00161.x. PMID 15996172.
- ↑ Cruciani, F.; La Fratta, R.; Trombetta, B.; Santolamazza, P.; Sellitto, D.; Colomb, E. B.; Dugoujon, J. -M.; Crivellaro, F. et al. (2007). "Tracing Past Human Male Movements in Northern/Eastern Africa and Western Eurasia: New Clues from Y-Chromosomal Haplogroups E-M78 and J-M12". Molecular Biology and Evolution 24 (6): 1300–1311. doi:10.1093/molbev/msm049. PMID 17351267.
- ↑ Lacan, Marie; Keyser, Christine; Ricaut, François-Xavier; Brucato, Nicolas; Tarrús, Josep; Bosch, Angel; Guilaine, Jean; Crubézy, Eric et al. (8 November 2011). "Ancient DNA suggests the leading role played by men in the Neolithic dissemination". Proceedings of the National Academy of Sciences 108 (45): 18255–18259. doi:10.1073/pnas.1113061108. PMID 22042855. Bibcode: 2011PNAS..10818255L.
- ↑ Szecsenyi-Nagy, Anna (2015). Molecular genetic investigation of the Neolithic population history in the western Carpathian Basin (Thesis). Johannes Gutenberg-Universität Mainz. doi:10.25358/openscience-1856.
- ↑ 44.0 44.1 Bosch, Elena; Calafell, Francesc; Comas, David; Oefner, Peter J.; Underhill, Peter A.; Bertranpetit, Jaume (April 2001). "High-Resolution Analysis of Human Y-Chromosome Variation Shows a Sharp Discontinuity and Limited Gene Flow between Northwestern Africa and the Iberian Peninsula". American Journal of Human Genetics 68 (4): 1019–1029. doi:10.1086/319521. PMID 11254456.
- ↑ Di Gaetano, Cornelia; Cerutti, Nicoletta; Crobu, Francesca; Robino, Carlo; Inturri, Serena; Gino, Sarah; Guarrera, Simonetta; Underhill, Peter A et al. (January 2009). "Differential Greek, Phoenician and northern African migrations to Sicily are supported by genetic evidence from the Y chromosome". European Journal of Human Genetics 17 (1): 91–99. doi:10.1038/ejhg.2008.120. PMID 18685561. "The co-occurrence of the Berber E3b1b-M81 (2.12 percent) and of the Mid-Eastern J1-M267 (3.81 percent) Hgs together with the presence of E3b1a1-V12, E3b1a3-V22, E3b1a4-V65 (5.5 percent) support the hypothesis of intrusion of North African genes. (...) These Hgs are common in Northern Africa and are observed only in Mediterranean Europe and together the presence of the E3b1b-M81 highlights the genetic relationships between northern Africa and Sicily. (...) Hg E3b1b-M81 network cluster confirms the genetic affinity between Sicily and North Africa."
- ↑ Adams, Susan M.; Bosch, Elena; Balaresque, Patricia L.; Ballereau, Stéphane J.; Lee, Andrew C.; Arroyo, Eduardo; López-Parra, Ana M.; Aler, Mercedes et al. (December 2008). "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula". The American Journal of Human Genetics 83 (6): 725–736. doi:10.1016/j.ajhg.2008.11.007. PMID 19061982. PMC 2668061. https://www.upf.edu/en/web/e-noticies/categories/-/asset_publisher/wEpPxsVRD6Vt/content/id/2509519.
- ↑ Fadhlaoui-Zid, K.; Haber, M.; Martínez-Cruz, B.; Zalloua, P.; Benammar Elgaaied, A.; Comas, D. (November 2013). "Genome-Wide and Paternal Diversity Reveal a Recent Origin of Human Populations in North Africa". PLOS ONE 8 (11): e80293. doi:10.1371/journal.pone.0080293. PMID 24312208. Bibcode: 2013PLoSO...880293F.
- ↑ Elkamel, Sarra; Marques, Sofia L.; Alvarez, Luis; Gomes, Veronica; Boussetta, Sami; Mourali-Chebil, Soufia; Khodjet-El-Khil, Houssein; Cherni, Lotfi et al. (August 2021). "Insights into the Middle Eastern paternal genetic pool in Tunisia: high prevalence of T-M70 haplogroup in an Arab population". Scientific Reports 11 (1): 15728. doi:10.1038/s41598-021-95144-x. PMID 34344940. Bibcode: 2021NatSR..1115728E.
- ↑ Semino, Ornella; Magri, Chiara; Benuzzi, Giorgia; Lin, Alice A.; Al-Zahery, Nadia; Battaglia, Vincenza; MacCioni, Liliana; Triantaphyllidis, Costas et al. (May 2004). "Origin, Diffusion, and Differentiation of Y-Chromosome Haplogroups E and J: Inferences on the Neolithization of Europe and Later Migratory Events in the Mediterranean Area". The American Journal of Human Genetics 74 (5): 1023–1034. doi:10.1086/386295. PMID 15069642.
- ↑ 50.0 50.1 Macaulay, Vincent; Richards, Martin; Hickey, Eileen; Vega, Emilce; Cruciani, Fulvio; Guida, Valentina; Scozzari, Rosaria; Bonné-Tamir, Batsheva et al. (January 1999). "The Emerging Tree of West Eurasian mtDNAs: A Synthesis of Control-Region Sequences and RFLPs". The American Journal of Human Genetics 64 (1): 232–249. doi:10.1086/302204. PMID 9915963.
- ↑ Fadhlaoui-Zid, K.; Plaza, S.; Calafell, F.; Ben Amor, M.; Comas, D.; Bennamar, A.; Gaaied, E. (2004). "Mitochondrial DNA Heterogeneity in Tunisian Berbers". Annals of Human Genetics 68 (3): 222–33. doi:10.1046/j.1529-8817.2004.00096.x. PMID 15180702.
- ↑ Esteban, E.; González-Pérez, E.; Harich, N.; López-Alomar, A.; Via, M.; Luna, F.; Moral, P. (2004). "Genetic relationships among Berbers and South Spaniards based on CD4 microsatellite/Alu haplotypes". Annals of Human Biology 31 (2): 202–212. doi:10.1080/03014460310001652275. PMID 15204363.
- ↑ Loueslati, B. Y.; Cherni, L.; Khodjet-Elkhil, H.; Ennafaa, H.; Pereira, L. S.; Amorim, A. N.; Ben Ayed, F.; Ben Ammar Elgaaied, A. (2006). "Islands Inside an Island: Reproductive Isolates on Jerba Island". American Journal of Human Biology 18 (1): 149–153. doi:10.1002/ajhb.20473. PMID 16378336.
- ↑ Cherni, L.; Loueslati, B. Y.; Pereira, L.; Ennafaa, H.; Amorim, A.; Gaaied, A. B. A. E. (2005). "Female Gene Pools of Berber and Arab Neighboring Communities in Central Tunisia: Microstructure of mtDNA Variation in North Africa". Human Biology 77 (1): 61–70. doi:10.1353/hub.2005.0028. PMID 16114817.
- ↑ 55.0 55.1 Achilli, Alessandro; Rengo, Chiara; Battaglia, Vincenza; Pala, Maria; Olivieri, Anna; Fornarino, Simona; Magri, Chiara; Scozzari, Rosaria et al. (May 2005). "Saami and Berbers—An Unexpected Mitochondrial DNA Link". The American Journal of Human Genetics 76 (5): 883–886. doi:10.1086/430073. PMID 15791543.
- ↑ Data from Achilli et al. 2005; Brakez et al. 2001; Cherni et al. 2005; Fadhlaoui-Zid et al. 2004; Krings et al.1999; Loueslati et al. 2006; Macaulay et al. 1999; Olivieri et al. 2006; Plaza et al. 2003; Rando et al. 1998; Stevanovitchet al. 2004; Coudray et al.2008; Cherni et al. 2008[improper synthesis?][verification needed]
- ↑ Harich, Nourdin; Costa, Marta D; Fernandes, Verónica; Kandil, Mostafa; Pereira, Joana B; Silva, Nuno M; Pereira, Luísa (December 2010). "The trans-Saharan slave trade - clues from interpolation analyses and high-resolution characterization of mitochondrial DNA lineages". BMC Evolutionary Biology 10 (1): 138. doi:10.1186/1471-2148-10-138. PMID 20459715. Bibcode: 2010BMCEE..10..138H.
- ↑ Frigi, Sabeh; Cherni, Lotfi; Fadhlaoui-zid, Karima; Benammar-Elgaaied, Amel (2010). "Ancient Local Evolution of African mtDNA Haplogroups in Tunisian Berber Populations". Human Biology 82 (4): 367–384. doi:10.3378/027.082.0402. Project MUSE 394730. PMID 21082907.
- ↑ Plaza, S.; Calafell, F.; Helal, A.; Bouzerna, N.; Lefranc, G.; Bertranpetit, J.; Comas, D. (2003). "Joining the Pillars of Hercules: MtDNA Sequences Show Multidirectional Gene Flow in the Western Mediterranean". Annals of Human Genetics 67 (4): 312–28. doi:10.1046/j.1469-1809.2003.00039.x. PMID 12914566. But very likely, most of the L mtDNA that has been found in minor amounts in Iberia, is actually pre-neolithic in origin, as it was demonstrated by María Cerezo et al., (Reconstructing ancient mitochondrial DNA links between Africa and Europe). "Haplogroup U6 is present at frequencies ranging from 0-7 percent in the various Iberian populations, with an average of 1.8 percent. Given that the frequency of U6 in NW Africa is 10 percent, the mtDNA contribution of NW Africa to Iberia can be estimated at 18 percent (though U6 has been found in many Iberian hunter-gatherer remains as well). This is larger than the contribution estimated with Y-chromosomal lineages (7 percent) (Bosch et al. 2001).
- ↑ Pereira, Luisa; Cunha, Carla; Alves, Cintia; Amorim, Antonio (2005). "African Female Heritage in Iberia: A Reassessment of mtDNA Lineage Distribution in Present Times". Human Biology 77 (2): 213–229. doi:10.1353/hub.2005.0041. PMID 16201138. "Although the absolute value of observed U6 frequency in Iberia is low, it reveals a discernible North African female contribution, if we keep in mind that haplogroup U6 is not very common in North Africa itself and virtually absent in the rest of Europe. Indeed, because the range of variation in western North Africa is 4-28 percent, the estimated minimum input is 8.54 percent"
- ↑ González, Ana M.; Brehm, Antonio; Pérez, José A.; Maca-Meyer, Nicole; Flores, Carlos; Cabrera, Vicente M. (April 2003). "Mitochondrial DNA affinities at the Atlantic fringe of Europe: Mitochondrial DNA in Atlantic Europe". American Journal of Physical Anthropology 120 (4): 391–404. doi:10.1002/ajpa.10168. PMID 12627534. "Our results clearly reinforce, extend, and clarify the preliminary clues of an 'important very ancient mtDNA contribution from northwest Africa into the Iberian Peninsula' (Côrte-Real et al., 1996; Rando et al., 1998; Flores et al., 2000a; Rocha et al., 1999)(...) Our own data allow us to make minimal estimates of the maternal African pre-Neolithic, Neolithic, and/or recent slave trade input into Iberia. For the former, we consider only the mean value of the U6 frequency in Northern African populations, excluding Saharans, Tuareg, and Mauritanians (16 percent), as the pre-Neolithic frequency in that area, and the present frequency in the whole Iberian Peninsula (2.3 percent) as the result of the northwest African gene flow at that time. The value obtained (14 percent) could be as high as 35 percent using the data of Corte-Real et al. (1996), or 27 percent with our north Portugal sample."
- ↑ Achilli, Alessandro; Olivieri, Anna; Pala, Maria; Metspalu, Ene; Fornarino, Simona; Battaglia, Vincenza; Accetturo, Matteo; Kutuev, Ildus et al. (April 2007). "Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans". The American Journal of Human Genetics 80 (4): 759–768. doi:10.1086/512822. PMID 17357081. "1.33% (3/226) in Calabria and 1.28 percent in Campania"
- ↑ Henn, B. M.; Botigué, L. R.; Gravel, S.; Wang, W.; Brisbin, A.; Byrnes, J. K.; Fadhlaoui-Zid, K.; Zalloua, P. A. et al. (2012). Schierup, Mikkel H. ed. "Genomic Ancestry of North Africans Supports Back-to-Africa Migrations". PLOS Genetics 8 (1): e1002397. doi:10.1371/journal.pgen.1002397. PMID 22253600.
- ↑ 64.0 64.1 64.2 Henn, B. M.; Botigué, L. R.; Gravel, S.; Wang, W.; Brisbin, A.; Byrnes, J. K.; Fadhlaoui-Zid, K.; Zalloua, P. A. et al. (2012). Schierup, Mikkel H. ed. "Genomic Ancestry of North Africans Supports Back-to-Africa Migrations". PLOS Genetics 8 (1): e1002397. doi:10.1371/journal.pgen.1002397. PMID 22253600.
- ↑ Hodgson, Jason A. (2014). "Early Back-to-Africa Migration into the Horn of Africa". PLOS Genetics 10 (6): e1004393. doi:10.1371/journal.pgen.1004393. PMID 24921250. "The non-African ancestry in the HOA, which is primarily attributed to a novel Ethio-Somali inferred ancestry component, is significantly differentiated from all neighboring non-African ancestries in North Africa, the Levant, and Arabia. The Ethio-Somali ancestry is found in all admixed HOA ethnic groups, shows little inter-individual variance within these ethnic groups, is estimated to have diverged from all other non-African ancestries by at most 23 ka, and does not carry the unique Arabian lactase persistence allele that arose about 4 ka. Taking into account published mitochondrial, Y chromosome, paleoclimate, and archaeological data, we find that the time of the Ethio-Somali back-to-Africa migration is most likely pre-agricultural. ... While this Ethio-Somali IAC is found primarily in Africa, it has clear non-African affinities (Text S1). ... The most recent divergence date estimates for the Ethio-Somali ancestral population are with the Maghrebi and Arabian ancestral populations at 23 and 25 ka. ... In this model, later diversification and expansion within particular Afro-Asiatic language groups may be associated with agricultural expansions and transmissions, but the deep diversification of the group is pre-agricultural. We hypothesize that a population with substantial Ethio-Somali ancestry could be the proto-Afro-Asiatic speakers.".
- ↑ Dobon, Begoña; Hassan, Hisham Y.; Laayouni, Hafid; Luisi, Pierre; Ricaño-Ponce, Isis; Zhernakova, Alexandra; Wijmenga, Cisca; Tahir, Hanan et al. (September 2015). "The genetics of East African populations: a Nilo-Saharan component in the African genetic landscape". Scientific Reports 5 (1): 9996. doi:10.1038/srep09996. PMID 26017457. Bibcode: 2015NatSR...5E9996D.
- ↑ Arauna, Lara R.; Mendoza-Revilla, Javier; Mas-Sandoval, Alex; Izaabel, Hassan; Bekada, Asmahan; Benhamamouch, Soraya; Fadhlaoui-Zid, Karima; Zalloua, Pierre et al. (2017-02-01). "Recent Historical Migrations Have Shaped the Gene Pool of Arabs and Berbers in North Africa". Molecular Biology and Evolution 34 (2): 318–329. doi:10.1093/molbev/msw218. ISSN 1537-1719. PMID 27744413.
- ↑ Arauna, Lara R; Comas, David (2017-09-15). "Genetic Heterogeneity between Berbers and Arabs". eLS: 1–7. doi:10.1002/9780470015902.a0027485. ISBN 9780470016176. https://doi.org/10.1002/9780470015902.a0027485. "1. A back-to-Africa migration replaced the population of North Africa in pre-Holocene times. 2. North African populations are very heterogeneous and are composed of North African, Middle Eastern, sub-Saharan and European genetic components. 3. No genetic differences have been found between Arab and Berber groups. 4. The Arab expansion had an important cultural and genetic impact in North Africa. 5. The Berber people are genetically diverse and heterogeneous.".
- ↑ van de Loosdrecht (2018-03-15). "Pleistocene North African genomes link Near Eastern and sub-Saharan African human populations". Science 360 (6388): 548–552. doi:10.1126/science.aar8380. ISSN 0036-8075. PMID 29545507. Bibcode: 2018Sci...360..548V.
- ↑ Henn, Brenna M.; Botigué, Laura R.; Gravel, Simon; Wang, Wei; Brisbin, Abra; Byrnes, Jake K.; Fadhlaoui-Zid, Karima; Zalloua, Pierre A. et al. (2012-01-12). "Genomic Ancestry of North Africans Supports Back-to-Africa Migrations". PLOS Genetics 8 (1): e1002397. doi:10.1371/journal.pgen.1002397. ISSN 1553-7390. PMID 22253600.
- ↑ Fregel, Rosa; Méndez, Fernando L.; Bokbot, Youssef; Martín-Socas, Dimas; Camalich-Massieu, María D.; Santana, Jonathan; Morales, Jacob; Ávila-Arcos, María C. et al. (2018-06-12). "Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe". Proceedings of the National Academy of Sciences 115 (26): 6774–6779. doi:10.1073/pnas.1800851115. ISSN 0027-8424. PMID 29895688. Bibcode: 2018PNAS..115.6774F.
- ↑ Choudhury, Ananyo; Aron, Shaun; Sengupta, Dhriti; Hazelhurst, Scott; Ramsay, Michèle (2018-08-01). "African genetic diversity provides novel insights into evolutionary history and local adaptations". Human Molecular Genetics 27 (R2): R209–R218. doi:10.1093/hmg/ddy161. ISSN 0964-6906. PMID 29741686.
- ↑ Pakstis, Andrew J.; Gurkan, Cemal; Dogan, Mustafa; Balkaya, Hasan Emin; Dogan, Serkan; Neophytou, Pavlos I.; Cherni, Lotfi; Boussetta, Sami et al. (December 2019). "Genetic relationships of European, Mediterranean, and SW Asian populations using a panel of 55 AISNPs". European Journal of Human Genetics 27 (12): 1885–1893. doi:10.1038/s41431-019-0466-6. PMID 31285530.
- ↑ Lucas-Sánchez, Marcel; Serradell, Jose M.; Comas, David (2021-04-26). "Population history of North Africa based on modern and ancient genomes". Human Molecular Genetics 30 (R1): R17–R23. doi:10.1093/hmg/ddaa261. ISSN 1460-2083. PMID 33284971.
- ↑ Sánchez-Quinto, Federico; Botigué, Laura R.; Civit, Sergi; Arenas, Conxita; Ávila-Arcos, María C.; Bustamante, Carlos D.; Comas, David; Lalueza-Fox, Carles (17 October 2012). "North African Populations Carry the Signature of Admixture with Neandertals". PLOS ONE 7 (10): e47765. doi:10.1371/journal.pone.0047765. PMID 23082212. Bibcode: 2012PLoSO...747765S.
- ↑ Khairat, Rabab; Ball, Markus; Chang, Chun-Chi Hsieh; Bianucci, Raffaella; Nerlich, Andreas G.; Trautmann, Martin; Ismail, Somaia; Shanab, Gamila M. L. et al. (August 2013). "First insights into the metagenome of Egyptian mummies using next-generation sequencing". Journal of Applied Genetics 54 (3): 309–325. doi:10.1007/s13353-013-0145-1. PMID 23553074.
- ↑ "MITOCHONDRIAL DNA AND PHYLOGENETIC ANALYSIS OF PREHISTORIC NORTH AFRICAN POPULATIONS". 8th ISABS Conference in Forensic, Anthropologic and Medical Genetics and Mayo Clinic Lectures in Translational Medicine. Split, Croatia: ISABS. June 2013. p. 232. ISBN 978-953-57695-0-7. http://www.isabs.hr/PDF/2013/ISABS-2013_book_of_abstracts.pdf. Retrieved 17 January 2016.
- ↑ Bernard Secher; Rosa Fregel; José M Larruga; Vicente M Cabrera; Phillip Endicott; José J Pestano; Ana M González (2014). "The history of the North African mitochondrial DNA haplogroup U6 gene flow into the African, Eurasian and American continents". BMC Evolutionary Biology 14 (1): 109. doi:10.1186/1471-2148-14-109. PMID 24885141. Bibcode: 2014BMCEE..14..109S.
- ↑ Ordóñez, Alejandra C.; Fregel, R.; Trujillo-Mederos, A.; Hervella, Montserrat; de-la-Rúa, Concepción; Arnay-de-la-Rosa, Matilde (February 2017). "Genetic studies on the prehispanic population buried in Punta Azul cave (El Hierro, Canary Islands)". Journal of Archaeological Science 78: 20–28. doi:10.1016/j.jas.2016.11.004. Bibcode: 2017JArSc..78...20O.
- ↑ Fregel, Rosa; Méndez, Fernando L.; Bokbot, Youssef; Martín-Socas, Dimas; Camalich-Massieu, María D.; Santana, Jonathan; Morales, Jacob; Ávila-Arcos, María C. et al. (26 June 2018). "Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe". Proceedings of the National Academy of Sciences 115 (26): 6774–6779. doi:10.1073/pnas.1800851115. PMID 29895688. Bibcode: 2018PNAS..115.6774F.
- ↑ Serra-Vidal, Gerard; Lucas-Sanchez, Marcel; Fadhlaoui-Zid, Karima; Bekada, Asmahan; Zalloua, Pierre; Comas, David (2019-11-18). "Heterogeneity in Palaeolithic Population Continuity and Neolithic Expansion in North Africa" (in English). Current Biology 29 (22): 3953–3959.e4. doi:10.1016/j.cub.2019.09.050. ISSN 0960-9822. PMID 31679935.
Bibliography
- Battaglia, Vincenza; Fornarino, Simona; Al-Zahery, Nadia; Olivieri, Anna; Pala, Maria; Myres, Natalie M; King, Roy J; Rootsi, Siiri et al. (2008), "Y-chromosomal evidence of the cultural diffusion of agriculture in southeast Europe", European Journal of Human Genetics 17 (6): 820–830, doi:10.1038/ejhg.2008.249, PMID 19107149
- Candelora, Danielle (2022). Candelora, Danielle; Ben-Marzouk, Nadia; Cooney, Kathyln. eds. Ancient Egyptian society: challenging assumptions, exploring approaches. Abingdon, Oxon: Routledge. ISBN 9780367434632.
- Mendizabal, Isabel; Sandoval, Karla; Berniell-Lee, Gemma; Calafell, Francesc; Salas, Antonio; Martinez-Fuentes, Antonio; Comas, David (2008), "Genetic origin, admixture, and asymmetry in maternal and paternal human lineages in Cuba", BMC Evol. Biol. 8 (1): 213, doi:10.1186/1471-2148-8-213, PMID 18644108, Bibcode: 2008BMCEE...8..213M
- Paracchini; Pearce, CL; Kolonel, LN; Altshuler, D; Henderson, BE; Tyler-Smith, C (2003), "A Y chromosomal influence on prostate cancer risk: the multi-ethnic cohort study", J Med Genet 40 (11): 815–819, doi:10.1136/jmg.40.11.815, PMID 14627670
- Silva; Carvalho, Elizeu; Costa, Guilherme; Tavares, Lígia; Amorim, António; Gusmão, Leonor (2006), "Y-chromosome genetic variation in Rio de Janeiro population", American Journal of Human Biology 18 (6): 829–837, doi:10.1002/ajhb.20567, PMID 17039481, http://www3.interscience.wiley.com/journal/113395738/abstract?CRETRY=1&SRETRY=0
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics 26 (3): 358–361. doi:10.1038/81685. PMID 11062480.
 |
