The beta clamp is a specific DNA clamp and a subunit of the DNA polymerase III holoenzyme found in bacteria. Two beta subunits are assembled around the DNA by the gamma subunit and ATP hydrolysis; this assembly is called the pre-initiation complex. After assembly around the DNA, the beta subunits' affinity for the gamma subunit is replaced by an affinity for the alpha and epsilon subunits, which together create the complete holoenzyme.[2][3][4] DNA polymerase III is the primary enzyme complex involved in prokaryotic DNA replication.
The gamma complex of DNA polymerase III, composed of γδδ'χψ subunits, catalyzes ATP to chaperone two beta subunits to bind to DNA. Once bound to DNA, the beta subunits can freely slide along double stranded DNA. The beta subunits in turn bind the αε polymerase complex. The α subunit possesses DNA polymerase activity and the ε subunit is a 3'-5' exonuclease.[4]
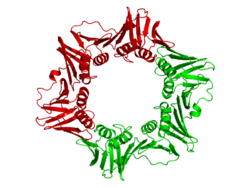
The beta chain of bacterial DNA polymerase III is composed of three topologically equivalent domains (N-terminal, central, and C-terminal). Two beta chain molecules are tightly associated to form a closed ring encircling duplex DNA.
| DNA polymerase III, beta chain (whole protein) |
|---|
| Identifiers |
|---|
| Symbol | DNA_polIII_beta |
|---|
| InterPro | IPR001001 |
|---|
| SMART | SM00480 |
|---|
| SCOP2 | 2pol / SCOPe / SUPFAM |
|---|
| Available protein structures: |
|---|
| Pfam
| |
|---|
| PDB | 1jqj, 1jql, 1mmi, 1ok7, 1unn, 1vpk, 2pol, 3bep, 3d1e, 3d1f, 3d1g |
|---|
|
|
As a drug target
Certain NSAIDs (carprofen, bromfenac, and vedaprofen) exhibit some suppression of bacterial DNA replication by inhibiting bacterial DNA clamp.[5]
Eukaryotic and archaeal
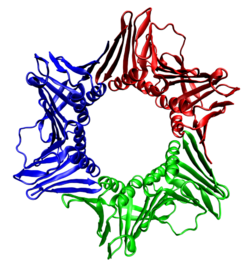
The sliding clamp in eukaryotes is assembled from a specific subunit of DNA polymerase delta called the proliferating cell nuclear antigen (PCNA). The N-terminal and C-terminal domains of PCNA are topologically identical. Three PCNA molecules are tightly associated to form a closed ring encircling duplex DNA.
The sequence of PCNA is well conserved between plants, animals and fungi, indicating a strong selective pressure for structure conservation, and suggesting that this type of DNA replication mechanism is conserved throughout eukaryotes.[7][8] In eukaryotes, a homologous, heterotrimeric "9-1-1 clamp" made up of RAD9-RAD1-HUS1 (911) is responsible for DNA damage checkpoint control.[9] This 9-1-1 clamp mounts onto DNA in the opposite direction.[10]
Archaea, probable evolutionary precursor of eukaryotes, also universally have at least one PCNA gene. This PCNA ring works with PolD, the single eukaryotic-like DNA polymerase in archaea responsible for multiple functions from replication to repair. Some unusual species have two or even three PCNA genes, forming heterotrimers or distinct specialized homotrimers.[11] Archaeons also share with eukaryotes the PIP (PCNA-interacting protein) motif, but a wider variety of such proteins performing different functions are found.[12]
PCNA is also appropriated by some viruses. The giant virus genus Chlorovirus, with PBCV-1 as a representative, carries in its genome two PCNA genes (Q84513, O41056) and a eukaryotic-type DNA polymerase.[13] Members of Baculoviridae also encode a PCNA homolog (P11038).[14]
Caudoviral
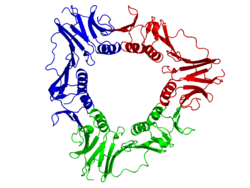
The viral gp45 sliding clamp subunit protein contains two domains. Each domain consists of two alpha helices and two beta sheets – the fold is duplicated and has internal pseudo two-fold symmetry.[16] Three gp45 molecules are tightly associated to form a closed ring encircling duplex DNA.
Herpesviral
Some members of Herpesviridae encode a protein that has a DNA clamp fold but does not associate into a ring clamp. The two-domain protein does, however, associate with the viral DNA polymerase and also acts to increase processivity.[17] As it does not form a ring, it does not need a clamp loader to be attached to DNA.[18]
Assembly
Sliding clamps are loaded onto their associated DNA template strands by specialized proteins known as "sliding clamp loaders", which also disassemble the clamps after replication has completed. The binding sites for these initiator proteins overlap with the binding sites for the DNA polymerase, so the clamp cannot simultaneously associate with a clamp loader and with a polymerase. Thus the clamp will not be actively disassembled while the polymerase remains bound. DNA clamps also associate with other factors involved in DNA and genome homeostasis, such as nucleosome assembly factors, Okazaki fragment ligases, and DNA repair proteins. All of these proteins also share a binding site on the DNA clamp that overlaps with the clamp loader site, ensuring that the clamp will not be removed while any enzyme is still working on the DNA. The activity of the clamp loader requires ATP hydrolysis to "close" the clamp around the DNA.
References
Further reading
External links
Lua error: Internal error: The interpreter has terminated with signal "24".
Lua error: Internal error: The interpreter has terminated with signal "24".
Lua error: Internal error: The interpreter has terminated with signal "24".