Biology:Triosephosphate isomerase
| triosephosphate isomerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
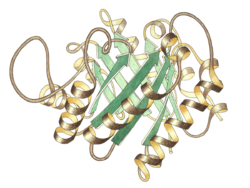 Side view of triose P isomerase monomer, active site at top center | |||||||||
| Identifiers | |||||||||
| EC number | 5.3.1.1 | ||||||||
| CAS number | 9023-78-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Triose-phosphate isomerase (TPI or TIM) is an enzyme (EC 5.3.1.1) that catalyzes the reversible interconversion of the triose phosphate isomers dihydroxyacetone phosphate and D-glyceraldehyde 3-phosphate.
| Dihydroxyacetone phosphate | triose phosphate isomerase | D-glyceraldehyde 3-phosphate | |
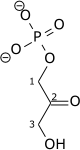
|
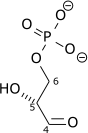
| ||
| |||
| triose phosphate isomerase | |||
Compound C00111 at KEGG Pathway Database.Enzyme 5.3.1.1 at KEGG Pathway Database.Compound C00118 at KEGG Pathway Database.
TPI plays an important role in glycolysis and is essential for efficient energy production. TPI has been found in nearly every organism searched for the enzyme, including animals such as mammals and insects as well as in fungi, plants, and bacteria. However, some bacteria that do not perform glycolysis, like ureaplasmas, lack TPI.
In humans, deficiencies in TPI are associated with a progressive, severe neurological disorder called triose phosphate isomerase deficiency. Triose phosphate isomerase deficiency is characterized by chronic hemolytic anemia. While there are various mutations that cause this disease, most include the replacement of glutamic acid at position 104 with an aspartic acid.[1]
Triose phosphate isomerase is a highly efficient enzyme, performing the reaction billions of times faster than it would occur naturally in solution. The reaction is so efficient that it is said to be catalytically perfect: It is limited only by the rate the substrate can diffuse into and out of the enzyme's active site.[2][3]
Mechanism
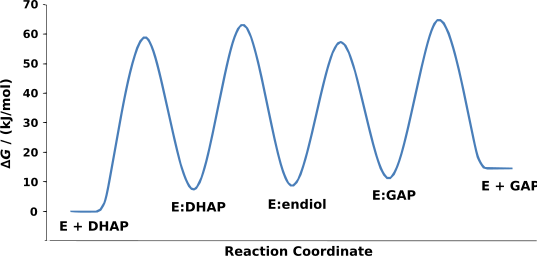
The mechanism involves the intermediate formation of an enediol. The relative free energy of each ground state and transition state has been determined experimentally, and is displayed in the figure.[2]
The structure of TPI facilitates the conversion between dihydroxyacetone phosphate (DHAP) and glyceraldehyde 3-phosphate (GAP). The nucleophilic glutamate 165 residue of TPI deprotonates the substrate,[4] and the electrophilic histidine 95 residue donates a proton to form the enediol intermediate.[5][6] When deprotonated, the enediolate then collapses and, abstracting a proton from protonated glutamate 165, forms the GAP product. Catalysis of the reverse reaction proceeds analogously, forming the same enediol but with enediolate collapse from the oxygen at C2.[7]
TPI is diffusion-limited. In terms of thermodynamics, DHAP formation is favored 20:1 over GAP production.[8] However, in glycolysis, the use of GAP in the subsequent steps of metabolism drives the reaction toward its production. TPI is inhibited by sulfate, phosphate, and arsenate ions, which bind to the active site.[9] Other inhibitors include 2-phosphoglycolate, a transition state analog, and D-glycerol-1-phosphate, a substrate analog.[10]
Structure
| Triosephosphate isomerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | TIM | ||||||||
| Pfam | PF00121 | ||||||||
| Pfam clan | CL0036 | ||||||||
| InterPro | IPR000652 | ||||||||
| PROSITE | PDOC00155 | ||||||||
| SCOP2 | 1tph / SCOPe / SUPFAM | ||||||||
| |||||||||
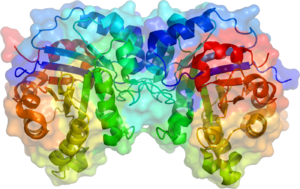
Triose phosphate isomerase is a dimer of identical subunits, each of which is made up of about 250 amino acid residues. The three-dimensional structure of a subunit contains eight α-helices on the outside and eight parallel β-strands on the inside. In the illustration, the ribbon backbone of each subunit is colored in blue to red from N-terminus to C-terminus. This structural motif is called an αβ-barrel, or a TIM-barrel, and is by far the most commonly observed protein fold. The active site of this enzyme is in the center of the barrel. A glutamic acid residue and a histidine are involved in the catalytic mechanism. The sequence around the active site residues is conserved in all known triose phosphate isomerases.
The structure of triose phosphate isomerase contributes to its function. Besides the precisely placed glutamate and histidine residues to form the enediol, a ten- or eleven-amino acid chain of TPI acts as a loop to stabilize the intermediate. The loop, formed by residues 166 to 176, closes and forms a hydrogen bond to the phosphate group of the substrate. This action stabilizes the enediol intermediate and the other transition states on the reaction pathway.[7]
In addition to making the reaction kinetically feasible, the TPI loop sequesters the reactive enediol intermediate to prevent decomposition to methylglyoxal and inorganic phosphate. The hydrogen bond between the enzyme and the phosphate group of the substrate makes such decomposition stereoelectronically unfavorable.[7] Methylglyoxal is a toxin and, if formed, is removed through the glyoxalase system.[11] The loss of a high-energy phosphate bond and the substrate for the rest of glycolysis makes formation of methylglyoxal inefficient.
Studies suggest that a lysine close to the active site (at position 12) is also crucial for enzyme function. The lysine, protonated at physiological pH, may help neutralize the negative charge of the phosphate group. When this lysine residue is replaced with a neutral amino acid, TPI loses all function, but variants with a different positively charged amino acid retain some function.[12]
See also
- TIM barrel
- Triose Phosphate Isomerase deficiency
- TPI1
- Triosephosphate isomerase in interactive 3D at Proteopedia
- Triosephosphate isomerase (TIM) family in PROSITE
References
- ↑ "Triosephosphate isomerase deficiency: facts and doubts". IUBMB Life 58 (12): 703–15. December 2006. doi:10.1080/15216540601115960. PMID 17424909.
- ↑ 2.0 2.1 "Free-energy profile of the reaction catalyzed by triosephosphate isomerase". Biochemistry 15 (25): 5627–31. December 1976. doi:10.1021/bi00670a031. PMID 999838.
- ↑ "Proton diffusion in the active site of triosephosphate isomerase". Biochemistry 29 (18): 4312–7. May 1990. doi:10.1021/bi00470a008. PMID 2161683.
- ↑ "On the three-dimensional structure and catalytic mechanism of triose phosphate isomerase". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 293 (1063): 159–71. June 1981. doi:10.1098/rstb.1981.0069. PMID 6115415.
- ↑ "Triosephosphate isomerase: removal of a putatively electrophilic histidine residue results in a subtle change in catalytic mechanism". Biochemistry 27 (16): 5948–60. August 1988. doi:10.1021/bi00416a019. PMID 2847777.
- ↑ "Electrophilic catalysis in triosephosphate isomerase: the role of histidine-95". Biochemistry 30 (12): 3011–9. March 1991. doi:10.1021/bi00226a005. PMID 2007138.
- ↑ 7.0 7.1 7.2 "Enzyme catalysis: not different, just better". Nature 350 (6314): 121–4. March 1991. doi:10.1038/350121a0. PMID 2005961.
- ↑ "Proton transfer in the mechanism of triosephosphate isomerase". Biochemistry 37 (47): 16828–38. November 1998. doi:10.1021/bi982089f. PMID 9843453.
- ↑ "Kinetic properties of triose-phosphate isomerase from Trypanosoma brucei brucei. A comparison with the rabbit muscle and yeast enzymes". European Journal of Biochemistry 168 (1): 69–74. October 1987. doi:10.1111/j.1432-1033.1987.tb13388.x. PMID 3311744.
- ↑ "Crystallographic analysis of the complex between triosephosphate isomerase and 2-phosphoglycolate at 2.5-A resolution: implications for catalysis". Biochemistry 29 (28): 6619–25. July 1990. doi:10.1021/bi00480a010. PMID 2204418.
- ↑ "Brief history of glyoxalase I and what we have learned about metal ion-dependent, enzyme-catalyzed isomerizations". Archives of Biochemistry and Biophysics 387 (1): 1–10. March 2001. doi:10.1006/abbi.2000.2253. PMID 11368170.
- ↑ "Triosephosphate isomerase requires a positively charged active site: the role of lysine-12". Biochemistry 33 (10): 2809–14. March 1994. doi:10.1021/bi00176a009. PMID 8130193.
External links
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Triosephosphate isomerase
{{Navbox
| name = Glycolysis enzymes | title = Metabolism: carbohydrate metabolism: [[Biology:Glycoglycolysis/gluconeogenesis enzymes | state = autocollapse | listclass = hlist
| group1 = Glycolysis
| list1 =
| |
| group2 = Gluconeogenesis only| list2 =
| to oxaloacetate: | |
|---|---|
| from lactate (Cori cycle): | |
| from alanine (Alanine cycle): | |
| from glycerol: |
| group4 = Regulatory | list4 =
}}
 |