Biology:Light-independent reactions
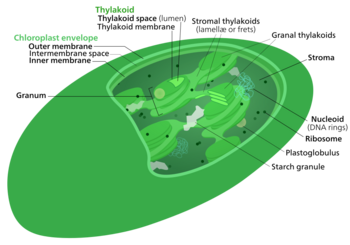
The light-independent reactions, or dark reactions,[1] of photosynthesis are chemical reactions that convert carbon dioxide and other compounds into glucose. These reactions occur in the stroma, the fluid-filled area of a chloroplast outside the thylakoid membranes. These reactions take the products (ATP and NADPH) of light-dependent reactions and perform further chemical processes on them.
Coupling to other Metabolic Pathways
These reactions are closely coupled to the thylakoid electron transport chain as reducing power provided by NADPH produced in the photosystem I is actively needed. The process of photorespiration, also known as C2 cycle, is also coupled to the dark reactions, as it results from an alternative reaction of the RuBisCO enzyme, and its final byproduct is also another glyceraldehyde-3-P.
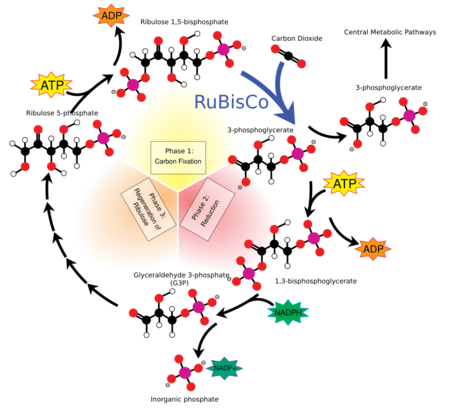
The Calvin cycle, Calvin–Benson–Bassham (CBB) cycle, reductive pentose phosphate cycle or C3 cycle is a series of biochemical redox reactions that take place in the stroma of chloroplast in photosynthetic organisms.
The cycle was discovered by Melvin Calvin, James Bassham, and Andrew Benson at the University of California, Berkeley[2] by using the radioactive isotope carbon-14.
Photosynthesis occurs in two stages in a cell. In the first stage, light-dependent reactions capture the energy of light and use it to make the energy-storage and transport molecules ATP and NADPH. The Calvin cycle uses the energy from short-lived electronically excited carriers to convert carbon dioxide and water into organic compounds[3] that can be used by the organism (and by animals that feed on it). This set of reactions is also called carbon fixation. The key enzyme of the cycle is called RuBisCO. In the following biochemical equations, the chemical species (phosphates and carboxylic acids) exist in equilibria among their various ionized states as governed by the pH.
The enzymes in the Calvin cycle are functionally equivalent to most enzymes used in other metabolic pathways such as gluconeogenesis and the pentose phosphate pathway, but they are found in the chloroplast stroma instead of the cell cytosol, separating the reactions. They are activated in the light (which is why the name "dark reaction" is misleading), and also by products of the light-dependent reaction. These regulatory functions prevent the Calvin cycle from being respired to carbon dioxide. Energy (in the form of ATP) would be wasted in carrying out these reactions that have no net productivity.
The sum of reactions in the Calvin cycle is the following:
- 3 CO2 + 6 NADPH + 6 H+ + 9 ATP → glyceraldehyde-3-phosphate (G3P) + 6 NADP+ + 9 ADP + 3 H2O + 8 Pi (Pi = inorganic phosphate)
Hexose (six-carbon) sugars are not a product of the Calvin cycle. Although many texts list a product of photosynthesis as C6H12O6, this is mainly a convenience to counter the equation of respiration, where six-carbon sugars are oxidized in mitochondria. The carbohydrate products of the Calvin cycle are three-carbon sugar phosphate molecules, or "triose phosphates", namely, glyceraldehyde-3-phosphate (G3P).
Steps
In the first stage of the Calvin cycle, a CO
2 molecule is incorporated into one of two three-carbon molecules (glyceraldehyde 3-phosphate or G3P), where it uses up two molecules of ATP and two molecules of NADPH, which had been produced in the light-dependent stage. The three steps involved are:
- The enzyme RuBisCO catalyses the carboxylation of ribulose-1,5-bisphosphate, RuBP, a 5-carbon compound, by carbon dioxide (a total of 6 carbons) in a two-step reaction.[4] The product of the first step is enediol-enzyme complex that can capture CO2 or O2. Thus, enediol-enzyme complex is the real carboxylase/oxygenase. The CO2 that is captured by enediol in second step produces an unstable six-carbon compound called 2-carboxy 3-keto 1,5-biphosphoribotol (or 3-keto-2-carboxyarabinitol 1,5-bisphosphate) that immediately splits into 2 molecules of 3-phosphoglycerate, or 3-PGA, a 3-carbon compound[5] (also: 3-phosphoglyceric acid, PGA, 3PGA).
- The enzyme phosphoglycerate kinase catalyses the phosphorylation of 3-PGA by ATP (which was produced in the light-dependent stage). 1,3-Bisphosphoglycerate (1,3BPGA, glycerate-1,3-bisphosphate) and ADP are the products. (However, note that two 3-PGAs are produced for every CO2 that enters the cycle, so this step utilizes two ATP per CO2 fixed.)
- The enzyme glyceraldehyde 3-phosphate dehydrogenase catalyses the reduction of 1,3BPGA by NADPH (which is another product of the light-dependent stage). Glyceraldehyde 3-phosphate (also called G3P, GP, TP, PGAL, GAP) is produced, and the NADPH itself is oxidized and becomes NADP+. Again, two NADPH are utilized per CO2 fixed.
The next stage in the Calvin cycle is to regenerate RuBP. Five G3P molecules produce three RuBP molecules, using up three molecules of ATP. Since each CO
2 molecule produces two G3P molecules, three CO
2 molecules produce six G3P molecules, of which five are used to regenerate RuBP, leaving a net gain of one G3P molecule per three CO
2 molecules (as would be expected from the number of carbon atoms involved).
The regeneration stage can be broken down into steps.
- Triose phosphate isomerase converts all of the G3P reversibly into dihydroxyacetone phosphate (DHAP), also a 3-carbon molecule.
- Aldolase and fructose-1,6-bisphosphatase convert a G3P and a DHAP into fructose 6-phosphate (6C). A phosphate ion is lost into solution.
- Then fixation of another CO2 generates two more G3P.
- F6P has two carbons removed by transketolase, giving erythrose-4-phosphate. The two carbons on transketolase are added to a G3P, giving the ketose xylulose-5-phosphate (Xu5P).
- E4P and a DHAP (formed from one of the G3P from the second CO2 fixation) are converted into sedoheptulose-1,7-bisphosphate (7C) by aldolase enzyme.
- Sedoheptulose-1,7-bisphosphatase (one of only three enzymes of the Calvin cycle that are unique to plants) cleaves sedoheptulose-1,7-bisphosphate into sedoheptulose-7-phosphate, releasing an inorganic phosphate ion into solution.
- Fixation of a third CO2 generates two more G3P. The ketose S7P has two carbons removed by transketolase, giving ribose-5-phosphate (R5P), and the two carbons remaining on transketolase are transferred to one of the G3P, giving another Xu5P. This leaves one G3P as the product of fixation of 3 CO2, with generation of three pentoses that can be converted to Ru5P.
- R5P is converted into ribulose-5-phosphate (Ru5P, RuP) by phosphopentose isomerase. Xu5P is converted into RuP by phosphopentose epimerase.
- Finally, phosphoribulokinase (another plant-unique enzyme of the pathway) phosphorylates RuP into RuBP, ribulose-1,5-bisphosphate, completing the Calvin cycle. This requires the input of one ATP.
Thus, of six G3P produced, five are used to make three RuBP (5C) molecules (totaling 15 carbons), with only one G3P available for subsequent conversion to hexose. This requires nine ATP molecules and six NADPH molecules per three CO2 molecules. The equation of the overall Calvin cycle is shown diagrammatically below.
RuBisCO also reacts competitively with O2 instead of CO2 in photorespiration. The rate of photorespiration is higher at high temperatures. Photorespiration turns RuBP into 3-PGA and 2-phosphoglycolate, a 2-carbon molecule that can be converted via glycolate and glyoxalate to glycine. Via the glycine cleavage system and tetrahydrofolate, two glycines are converted into serine +CO2. Serine can be converted back to 3-phosphoglycerate. Thus, only 3 of 4 carbons from two phosphoglycolates can be converted back to 3-PGA. It can be seen that photorespiration has very negative consequences for the plant, because, rather than fixing CO2, this process leads to loss of CO2. C4 carbon fixation evolved to circumvent photorespiration, but can occur only in certain plants native to very warm or tropical climates—corn, for example.
Products
The immediate products of one turn of the Calvin cycle are 2 glyceraldehyde-3-phosphate (G3P) molecules, 3 ADP, and 2 NADP+. (ADP and NADP+ are not really "products." They are regenerated and later used again in the Light-dependent reactions). Each G3P molecule is composed of 3 carbons. For the Calvin cycle to continue, RuBP (ribulose 1,5-bisphosphate) must be regenerated. So, 5 out of 6 carbons from the 2 G3P molecules are used for this purpose. Therefore, there is only 1 net carbon produced to play with for each turn. To create 1 surplus G3P requires 3 carbons, and therefore 3 turns of the Calvin cycle. To make one glucose molecule (which can be created from 2 G3P molecules) would require 6 turns of the Calvin cycle. Surplus G3P can also be used to form other carbohydrates such as starch, sucrose, and cellulose, depending on what the plant needs.[6]
Light-dependent regulation
These reactions do not occur in the dark or at night. There is a light-dependent regulation of the cycle enzymes, as the third step requires reduced NADP.
There are two regulation systems at work when the cycle must be turned on or off: the thioredoxin/ferredoxin activation system, which activates some of the cycle enzymes; and the RuBisCo enzyme activation, active in the Calvin cycle, which involves its own activase.
The thioredoxin/ferredoxin system activates the enzymes glyceraldehyde-3-P dehydrogenase, glyceraldehyde-3-P phosphatase, fructose-1,6-bisphosphatase, sedoheptulose-1,7-bisphosphatase, and ribulose-5-phosphatase kinase, which are key points of the process. This happens when light is available, as the ferredoxin protein is reduced in the photosystem I complex of the thylakoid electron chain when electrons are circulating through it.[7] Ferredoxin then binds to and reduces the thioredoxin protein, which activates the cycle enzymes by severing a cystine bond found in all these enzymes. This is a dynamic process as the same bond is formed again by other proteins that deactivate the enzymes. The implications of this process are that the enzymes remain mostly activated by day and are deactivated in the dark when there is no more reduced ferredoxin available.
The enzyme RuBisCo has its own, more complex activation process. It requires that a specific lysine amino acid be carbamylated to activate the enzyme. This lysine binds to RuBP and leads to a non-functional state if left uncarbamylated. A specific activase enzyme, called RuBisCo activase, helps this carbamylation process by removing one proton from the lysine and making the binding of the carbon dioxide molecule possible. Even then the RuBisCo enzyme is not yet functional, as it needs a magnesium ion bound to the lysine to function. This magnesium ion is released from the thylakoid lumen when the inner pH drops due to the active pumping of protons from the electron flow. RuBisCo activase itself is activated by increased concentrations of ATP in the stroma caused by its phosphorylation.
References
- Citations
- ↑ Silverstein, Alvin (2008). Photosynthesis. Twenty-First Century Books. p. 21. ISBN 9780822567981.
- ↑ "The path of carbon in photosynthesis". J Biol Chem 185 (2): 781–7. 1950. doi:10.2172/910351. PMID 14774424. http://www.jbc.org/cgi/reprint/185/2/781.pdf.
- ↑ Campbell, Neil A.; Brad Williamson; Robin J. Heyden (2006). Biology: Exploring Life. Boston, Massachusetts: Pearson Prentice Hall. ISBN 0-13-250882-6. http://www.phschool.com/el_marketing.html.
- ↑ Farazdaghi H (2009). "Modeling the Kinetics of Activation and Reaction of RuBisCO from Gas Exchange". Advances in Photosynthesis and Respiration 29 (IV): 275–294. doi:10.1007/978-1-4020-9237-4_12. http://www.springerlink.com/content/qu357422246r8870/.
- ↑ Campbell, and Reece Biology: 8th Edition, page 198. Benjamin Cummings, December 7, 2007.
- ↑ Russell, Wolfe et al.Biology: Exploring the Diversity of Life'.Toronto:Nelson College Indigenous,1st ed, Vol. 1, 2010, pg 151
- ↑ Besse, I; Buchanan, B (1997). "Thioredoxin-linked animal and plant processes: the new generation". Bot. Bull. Acad. Sin 38: 1–11.
- Bibliography
- Bassham JA (2003). "Mapping the carbon reduction cycle: a personal retrospective". Photosyn. Res. 76 (1-3): 35–52. doi:10.1023/A:1024929725022. PMID 16228564.
- Diwan, Joyce J. (2005). "Photosynthetic Dark Reaction". Biochemistry and Biophysics, Rensselaer Polytechnic Institute. http://www.rpi.edu/dept/bcbp/molbiochem/MBWeb/mb2/part1/dark.htm.
- Portis, Archie; Parry, Martin (2007). "Discoveries in Rubisco (Ribulose 1,5-bisphosphate carboxylase/oxygenase): a historical perspective". Photosynthesis Research 94 (1): 121–143. doi:10.1007/s11120-007-9225-6. PMID 17665149. Archived from the original on 2012-03-12. https://web.archive.org/web/20120312090903/http://ddr.nal.usda.gov/dspace/bitstream/10113/3976/1/IND43944177.pdf
Further reading
- Rubisco Activase, from the Plant Physiology Online website
- Thioredoxins, from the Plant Physiology Online website
External links
- The Biochemistry of the Calvin Cycle at Rensselaer Polytechnic Institute
- The Calvin Cycle and the Pentose Phosphate Pathway from Biochemistry, Fifth Edition by Jeremy M. Berg, John L. Tymoczko and Lubert Stryer. Published by W. H. Freeman and Company (2002).