Biology:Serpin
| Serpin (serine protease inhibitor) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Serpin, SERPIN (root symbol of family) | ||||||||||
| Pfam | PF00079 | ||||||||||
| InterPro | IPR000215 | ||||||||||
| PROSITE | PDOC00256 | ||||||||||
| SCOP2 | 1hle / SCOPe / SUPFAM | ||||||||||
| CDD | cd00172 | ||||||||||
| |||||||||||
Serpins are a superfamily of proteins with similar structures that were first identified for their protease inhibition activity and are found in all kingdoms of life.[1][2] The acronym serpin was originally coined because the first serpins to be identified act on chymotrypsin-like serine proteases (serine protease inhibitors).[3][4][5] They are notable for their unusual mechanism of action, in which they irreversibly inhibit their target protease by undergoing a large conformational change to disrupt the target's active site.[6][7] This contrasts with the more common competitive mechanism for protease inhibitors that bind to and block access to the protease active site.[8][9]
Protease inhibition by serpins controls an array of biological processes, including coagulation and inflammation, and consequently these proteins are the target of medical research.[10] Their unique conformational change also makes them of interest to the structural biology and protein folding research communities.[7][8] The conformational-change mechanism confers certain advantages, but it also has drawbacks: serpins are vulnerable to mutations that can result in serpinopathies such as protein misfolding and the formation of inactive long-chain polymers.[11][12] Serpin polymerisation not only reduces the amount of active inhibitor, but also leads to accumulation of the polymers, causing cell death and organ failure.[10]
Although most serpins control proteolytic cascades, some proteins with a serpin structure are not enzyme inhibitors, but instead perform diverse functions such as storage (as in egg white—ovalbumin), transport as in hormone carriage proteins (thyroxine-binding globulin, cortisol-binding globulin) and molecular chaperoning (HSP47).[9] The term serpin is used to describe these members as well, despite their non-inhibitory function, since they are evolutionarily related.[1]
History
Protease inhibitory activity in blood plasma was first reported in the late 1800s,[13] but it was not until the 1950s that the serpins antithrombin and alpha 1-antitrypsin were isolated,[14] with the subsequent recognition of their close family homology in 1979.[15][16] That they belonged to a new protein family became apparent on their further alignment with the non-inhibitory egg-white protein ovalbumin, to give what was initially called the alpha1-antitrypsin-antithrombin III-ovalbumin superfamily of serine proteinase inhibitors,[17] but was subsequently succinctly renamed as the Serpins.[18] The initial characterisation of the new family centred on alpha1-antitrypsin, a serpin present in high concentration in blood plasma, the common genetic disorder of which was shown to cause a predisposition to the lung disease emphysema[19] and to liver cirrhosis.[20] The identification of the S and Z mutations[21][22] responsible for the genetic deficiency and the subsequent sequence alignments of alpha1-antitrypsin and antithrombin in 1982 led to the recognition of the close homologies of the active sites of the two proteins,[23][24] centred on a methionine[25] in alpha1-antitrypsin as an inhibitor of tissue elastase and on arginine in antithrombin[26] as an inhibitor of thrombin.[27]
The critical role of the active centre residue in determining the specificity of inhibition of serpins was unequivocally confirmed by the finding that a natural mutation of the active centre methionine in alpha1-antitrypsin to an arginine, as in antithrombin, resulted in a severe bleeding disorder.[28] This active-centre specificity of inhibition was also evident in the many other families of protease inhibitors[7] but the serpins differed from them in being much larger proteins and also in possessing what was soon apparent as an inherent ability to undergo a change in shape. The nature of this conformational change was revealed with the determination in 1984 of the first crystal structure of a serpin, that of post-cleavage alpha1-antitrypsin.[29] This together with the subsequent solving of the structure of native (uncleaved) ovalbumin[30] indicated that the inhibitory mechanism of the serpins involved a remarkable conformational shift, with the movement of the exposed peptide loop containing the reactive site and its incorporation as a middle strand in the main beta-pleated sheet that characterises the serpin molecule.[31][32] Early evidence of the essential role of this loop movement in the inhibitory mechanism came from the finding that even minor aberrations in the amino acid residues that form the hinge of the movement in antithrombin resulted in thrombotic disease.[31][33] Ultimate confirmation of the linked displacement of the target protease by this loop movement was provided in 2000 by the structure of the post-inhibitory complex of alpha1-antitrypsin with trypsin,[6] showing how the displacement results in the deformation and inactivation of the attached protease. Subsequent structural studies have revealed an additional advantage of the conformational mechanism[34] in allowing the subtle modulation of inhibitory activity, as notably seen at tissue level[35] with the functionally diverse serpins in human plasma.
Over 1000 serpins have now been identified, including 36 human proteins, as well as molecules in all kingdoms of life—animals, plants, fungi, bacteria, and archaea—and some viruses.[36][37][38] The central feature of all is a tightly conserved framework, which allows the precise alignment of their key structural and functional components based on the template structure of alpha1-antitrypsin.[39] In the 2000s, a systematic nomenclature was introduced in order to categorise members of the serpin superfamily based on their evolutionary relationships.[1] Serpins are therefore the largest and most diverse superfamily of protease inhibitors.[40]
Activity

Most serpins are protease inhibitors, targeting extracellular, chymotrypsin-like serine proteases. These proteases possess a nucleophilic serine residue in a catalytic triad in their active site. Examples include thrombin, trypsin, and human neutrophil elastase.[41] Serpins act as irreversible, suicide inhibitors by trapping an intermediate of the protease's catalytic mechanism.[6]
Some serpins inhibit other protease classes, typically cysteine proteases, and are termed "cross-class inhibitors". These enzymes differ from serine proteases in that they use a nucleophilic cysteine residue, rather than a serine, in their active site.[42] Nonetheless, the enzymatic chemistry is similar, and the mechanism of inhibition by serpins is the same for both classes of protease.[43] Examples of cross-class inhibitory serpins include serpin B4 a squamous cell carcinoma antigen 1 (SCCA-1) and the avian serpin myeloid and erythroid nuclear termination stage-specific protein (MENT), which both inhibit papain-like cysteine proteases.[44][45][46]
Biological function and localization
Protease inhibition
Approximately two-thirds of human serpins perform extracellular roles, inhibiting proteases in the bloodstream in order to modulate their activities. For example, extracellular serpins regulate the proteolytic cascades central to blood clotting (antithrombin), the inflammatory and immune responses (antitrypsin, antichymotrypsin, and C1-inhibitor) and tissue remodelling (PAI-1).[9] By inhibiting signalling cascade proteases, they can also affect development.[47][48] The table of human serpins (below) provides examples of the range of functions performed by human serpin, as well as some of the diseases that result from serpin deficiency.
The protease targets of intracellular inhibitory serpins have been difficult to identify, since many of these molecules appear to perform overlapping roles. Further, many human serpins lack precise functional equivalents in model organisms such as the mouse. Nevertheless, an important function of intracellular serpins may be to protect against the inappropriate activity of proteases inside the cell.[49] For example, one of the best-characterised human intracellular serpins is Serpin B9, which inhibits the cytotoxic granule protease granzyme B. In doing so, Serpin B9 may protect against inadvertent release of granzyme B and premature or unwanted activation of cell death pathways.[50]
Some viruses use serpins to disrupt protease functions in their host. The cowpox viral serpin CrmA (cytokine response modifier A) is used in order to avoid inflammatory and apoptotic responses of infected host cells. CrmA increases infectivity by suppressing its host's inflammatory response through inhibition of IL-1 and IL-18 processing by the cysteine protease caspase-1.[51] In eukaryotes, a plant serpin inhibits both metacaspases[52] and a papain-like cysteine protease.[53]
Non-inhibitory roles
Non-inhibitory extracellular serpins also perform a wide array of important roles. Thyroxine-binding globulin and transcortin transport the hormones thyroxine and cortisol, respectively.[54][55] The non-inhibitory serpin ovalbumin is the most abundant protein in egg white. Its exact function is unknown, but it is thought to be a storage protein for the developing foetus.[56] Heat shock serpin 47 is a chaperone, essential for proper folding of collagen. It acts by stabilising collagen's triple helix whilst it is being processed in the endoplasmic reticulum.[57]
Some serpins are both protease inhibitors and perform additional roles. For example, the nuclear cysteine protease inhibitor MENT, in birds also acts as a chromatin remodelling molecule in a bird's red blood cells.[45][58]
Structure

All serpins share a common structure (or fold), despite their varied functions. All typically have three β-sheets (named A, B and C) and eight or nine α-helices (named hA–hI).[29][30] The most significant regions to serpin function are the A-sheet and the reactive centre loop (RCL). The A-sheet includes two β-strands that are in a parallel orientation with a region between them called the 'shutter', and upper region called the 'breach'. The RCL forms the initial interaction with the target protease in inhibitory molecules. Structures have been solved showing the RCL either fully exposed or partially inserted into the A-sheet, and serpins are thought to be in dynamic equilibrium between these two states.[8] The RCL also only makes temporary interactions with the rest of the structure, and is therefore highly flexible and exposed to the solvent.[8]
The serpin structures that have been determined cover several different conformations, which has been necessary for the understanding of their multiple-step mechanism of action. Structural biology has therefore played a central role in the understanding of serpin function and biology.[8]
Conformational change and inhibitory mechanism
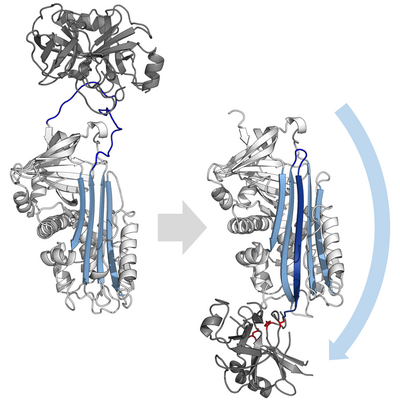
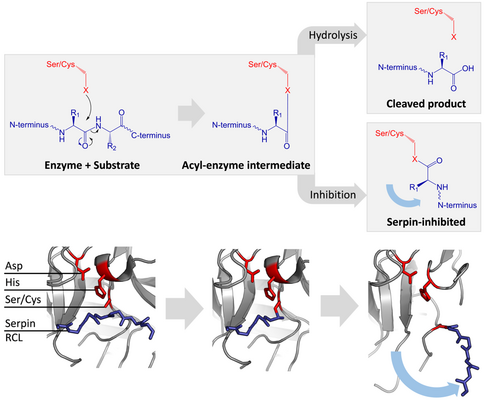
Inhibitory serpins do not inhibit their target proteases by the typical competitive (lock-and-key) mechanism used by most small protease inhibitors (e.g. Kunitz-type inhibitors). Instead, serpins use an unusual conformational change, which disrupts the structure of the protease and prevents it from completing catalysis. The conformational change involves the RCL moving to the opposite end of the protein and inserting into β-sheet A, forming an extra antiparallel β-strand. This converts the serpin from a stressed state, to a lower-energy relaxed state (S to R transition).[7][8][61]
Serine and cysteine proteases catalyse peptide bond cleavage by a two-step process. Initially, the catalytic residue of the active site triad performs a nucleophilic attack on the peptide bond of the substrate. This releases the new N-terminus and forms a covalent ester-bond between the enzyme and the substrate.[7] This covalent complex between enzyme and substrate is called an acyl-enzyme intermediate. For standard substrates, the ester bond is hydrolysed and the new C-terminus is released to complete catalysis. However, when a serpin is cleaved by a protease, it rapidly undergoes the S to R transition before the acyl-enzyme intermediate is hydrolysed.[7] The efficiency of inhibition depends on fact that the relative kinetic rate of the conformational change is several orders of magnitude faster than hydrolysis by the protease.
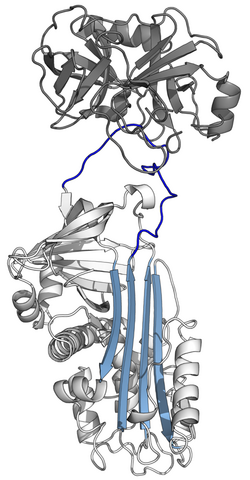
Since the RCL is still covalently attached to the protease via the ester bond, the S to R transition pulls protease from the top to the bottom of the serpin and distorts the catalytic triad. The distorted protease can only hydrolyse the acyl enzyme intermediate extremely slowly and so the protease remains covalently attached for days to weeks.[6] Serpins are classed as irreversible inhibitors and as suicide inhibitors since each serpin protein permanently inactivates a single protease, and can only function once.[7]
Allosteric activation
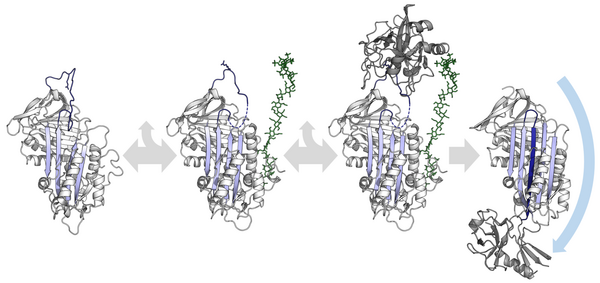
The conformational mobility of serpins provides a key advantage over static lock-and-key protease inhibitors.[34] In particular, the function of inhibitory serpins can be regulated by allosteric interactions with specific cofactors. The X-ray crystal structures of antithrombin, heparin cofactor II, MENT and murine antichymotrypsin reveal that these serpins adopt a conformation wherein the first two amino acids of the RCL are inserted into the top of the A β-sheet. The partially inserted conformation is important because co-factors are able to conformationally switch certain partially inserted serpins into a fully expelled form.[62][63] This conformational rearrangement makes the serpin a more effective inhibitor.
The archetypal example of this situation is antithrombin, which circulates in plasma in a partially inserted relatively inactive state. The primary specificity determining residue (the P1 arginine) points toward the body of the serpin and is unavailable to the protease. Upon binding a high-affinity pentasaccharide sequence within long-chain heparin, antithrombin undergoes a conformational change, RCL expulsion, and exposure of the P1 arginine. The heparin pentasaccharide-bound form of antithrombin is, thus, a more effective inhibitor of thrombin and factor Xa.[64][65] Furthermore, both of these coagulation proteases also contain binding sites (called exosites) for heparin. Heparin, therefore, also acts as a template for binding of both protease and serpin, further dramatically accelerating the interaction between the two parties. After the initial interaction, the final serpin complex is formed and the heparin moiety is released. This interaction is physiologically important. For example, after injury to the blood vessel wall, heparin is exposed, and antithrombin is activated to control the clotting response. Understanding of the molecular basis of this interaction enabled the development of Fondaparinux, a synthetic form of Heparin pentasaccharide used as an anti-clotting drug.[66][67]
Latent conformation

Certain serpins spontaneously undergo the S to R transition without having been cleaved by a protease, to form a conformation termed the latent state. Latent serpins are unable to interact with proteases and so are no longer protease inhibitors. The conformational change to latency is not exactly the same as the S to R transition of a cleaved serpin. Since the RCL is still intact, the first strand of the C-sheet has to peel off to allow full RCL insertion.[68]
Regulation of the latency transition can act as a control mechanism in some serpins, such as PAI-1. Although PAI-1 is produced in the inhibitory S conformation, it "auto-inactivates" by changing to the latent state unless it is bound to the cofactor vitronectin.[68] Similarly, antithrombin can also spontaneously convert to the latent state, as an additional modulation mechanism to its allosteric activation by heparin.[69] Finally, the N-terminus of tengpin, a serpin from Thermoanaerobacter tengcongensis, is required to lock the molecule in the native inhibitory state. Disruption of interactions made by the N-terminal region results in spontaneous conformational change of this serpin to the latent conformation.[70][71]
Conformational change in non-inhibitory functions
Certain non-inhibitory serpins also use the serpin conformational change as part of their function. For example, the native (S) form of thyroxine-binding globulin has high affinity for thyroxine, whereas the cleaved (R) form has low affinity. Similarly, transcortin has higher affinity for cortisol when in its native (S) state, than its cleaved (R) state. Thus, in these serpins, RCL cleavage and the S to R transition has been commandeered to allow for ligand release, rather than protease inhibition.[54][55][72]
In some serpins, the S to R transition can activate cell signalling events. In these cases, a serpin that has formed a complex with its target protease, is then recognised by a receptor. The binding event then leads to downstream signalling by the receptor.[73] The S to R transition is therefore used to alert cells to the presence of protease activity.[73] This differs from the usual mechanism whereby serpins affect signalling simply by inhibiting proteases involved in a signalling cascade.[47][48]
Degradation
When a serpin inhibits a target protease, it forms a permanent complex, which needs to be disposed of. For extracellular serpins, the final serpin-enzyme complexes are rapidly cleared from circulation. One mechanism by which this occurs in mammals is via the low-density lipoprotein receptor-related protein (LRP), which binds to inhibitory complexes made by antithrombin, PA1-1, and neuroserpin, causing cellular uptake.[73][74] Similarly, the Drosophila necrotic serpin is degraded in the lysosome after being trafficked into the cell by the Lipophorin Receptor-1 (homologous to the mammalian LDL receptor family).[75]
Disease and serpinopathies
Serpins are involved in a wide array of physiological functions, and so mutations in genes encoding them can cause a range of diseases. Mutations that change the activity, specificity or aggregation properties of serpins all affect how they function. The majority of serpin-related diseases are the result of serpin polymerisation into aggregates, though several other types of disease-linked mutations also occur.[8][76] The disorder alpha-1 antitrypsin deficiency is one of the most common hereditary diseases.[11][77]
Inactivity or absence

Since the stressed serpin fold is high-energy, mutations can cause them to incorrectly change into their lower-energy conformations (e.g. relaxed or latent) before they have correctly performed their inhibitory role.[10]
Mutations that affect the rate or the extent of RCL insertion into the A-sheet can cause the serpin to undergo its S to R conformational change before having engaged a protease. Since a serpin can only make this conformational change once, the resulting misfired serpin is inactive and unable to properly control its target protease.[10][78] Similarly, mutations that promote inappropriate transition to the monomeric latent state cause disease by reducing the amount of active inhibitory serpin. For example, the disease-linked antithrombin variants wibble and wobble,[79] both promote formation of the latent state.
The structure of the disease-linked mutant of antichymotrypsin (L55P) revealed another, inactive "δ-conformation". In the δ-conformation, four residues of the RCL are inserted into the top of β-sheet A. The bottom half of the sheet is filled as a result of one of the α-helices (the F-helix) partially switching to a β-strand conformation, completing the β-sheet hydrogen bonding.[80] It is unclear whether other serpins can adopt this conformer, and whether this conformation has a functional role, but it is speculated that the δ-conformation may be adopted by Thyroxine-binding globulin during thyroxine release.[55] The non-inhibitory proteins related to serpins can also cause diseases when mutated. For example, mutations in SERPINF1 cause osteogenesis imperfecta type VI in humans.[81]
In the absence of a required serpin, the protease that it normally would regulate is over-active, leading to pathologies.[10] Consequently, simple deficiency of a serpin (e.g. a null mutation) can result in disease.[82] Gene knockouts, particularly in mice, are used experimentally to determine the normal functions of serpins by the effect of their absence.[83]
Specificity change
In some rare cases, a single amino acid change in a serpin's RCL alters its specificity to target the wrong protease. For example, the Antitrypsin-Pittsburgh mutation (M358R) causes the α1-antitrypsin serpin to inhibit thrombin, causing a bleeding disorder.[28]
Polymerisation and aggregation
The majority of serpin diseases are due to protein aggregation and are termed "serpinopathies".[12][80] Serpins are vulnerable to disease-causing mutations that promote formation of misfolded polymers due to their inherently unstable structures.[80] Well-characterised serpinopathies include α1-antitrypsin deficiency (alpha-1), which may cause familial emphysema, and sometimes liver cirrhosis, certain familial forms of thrombosis related to antithrombin deficiency, types 1 and 2 hereditary angioedema (HAE) related to deficiency of C1-inhibitor, and familial encephalopathy with neuroserpin inclusion bodies (FENIB; a rare type of dementia caused by neuroserpin polymerisation).[11][12][84]
Each monomer of the serpin aggregate exists in the inactive, relaxed conformation (with the RCL inserted into the A-sheet). The polymers are therefore hyperstable to temperature and unable to inhibit proteases. Serpinopathies therefore cause pathologies similarly to other proteopathies (e.g. prion diseases) via two main mechanisms.[11][12] First, the lack of active serpin results in uncontrolled protease activity and tissue destruction. Second, the hyperstable polymers themselves clog up the endoplasmic reticulum of cells that synthesize serpins, eventually resulting in cell death and tissue damage. In the case of antitrypsin deficiency, antitrypsin polymers cause the death of liver cells, sometimes resulting in liver damage and cirrhosis. Within the cell, serpin polymers are slowly removed via degradation in the endoplasmic reticulum.[85] However, the details of how serpin polymers cause cell death remains to be fully understood.[11]
Physiological serpin polymers are thought to form via domain swapping events, where a segment of one serpin protein inserts into another.[86] Domain-swaps occur when mutations or environmental factors interfere with the final stages of serpin folding to the native state, causing high-energy intermediates to misfold.[87] Both dimer and trimer domain-swap structures have been solved. In the dimer (of antithrombin), the RCL and part of the A-sheet incorporates into the A-sheet of another serpin molecule.[86] The domain-swapped trimer (of antitrypsin) forms via the exchange of an entirely different region of the structure, the B-sheet (with each molecule's RCL inserted into its own A-sheet).[88] It has also been proposed that serpins may form domain-swaps by inserting the RCL of one protein into the A-sheet of another (A-sheet polymerisation).[84][89] These domain-swapped dimer and trimer structures are thought to be the building blocks of the disease-causing polymer aggregates, but the exact mechanism is still unclear.[86][87][88][90]
Therapeutic strategies
Several therapeutic approaches are in use or under investigation to treat the most common serpinopathy: antitrypsin deficiency.[11] Antitrypsin augmentation therapy is approved for severe antitrypsin deficiency-related emphysema.[91] In this therapy, antitrypsin is purified from the plasma of blood donors and administered intravenously (first marketed as Prolastin).[11][92] To treat severe antitrypsin deficiency-related disease, lung and liver transplantation has proven effective.[11][93] In animal models, gene targeting in induced pluripotent stem cells has been successfully used to correct an antitrypsin polymerisation defect and to restore the ability of the mammalian liver to secrete active antitrypsin.[94] Small molecules have also been developed that block antitrypsin polymerisation in vitro.[95][96]
Evolution
Serpins are the most widely distributed and largest superfamily of protease inhibitors.[1][40] They were initially believed to be restricted to eukaryote organisms, but have since been found in bacteria, archaea and some viruses.[36][37][97] It remains unclear whether prokaryote genes are the descendants of an ancestral prokaryotic serpin or the product of horizontal gene transfer from eukaryotes. Most intracellular serpins belong to a single phylogenetic clade, whether they come from plants or animals, indicating that the intracellular and extracellular serpins may have diverged before the plants and animals.[98] Exceptions include the intracellular heat shock serpin HSP47, which is a chaperone essential for proper folding of collagen, and cycles between the cis-Golgi and the endoplasmic reticulum.[57]
Protease-inhibition is thought to be the ancestral function, with non-inhibitory members the results of evolutionary neofunctionalisation of the structure. The S to R conformational change has also been adapted by some binding serpins to regulate affinity for their targets.[55]
Distribution
Animal
Human
The human genome encodes 16 serpin clades, termed serpinA through serpinP, including 29 inhibitory and 7 non-inhibitory serpin proteins.[9][83] The human serpin naming system is based upon a phylogenetic analysis of approximately 500 serpins from 2001, with proteins named serpinXY, where X is the clade of the protein and Y the number of the protein within that clade.[1][36][83] The functions of human serpins have been determined by a combination of biochemical studies, human genetic disorders, and knockout mouse models.[83]
| Gene name | Common name | Localisation | Function / Activity[9][83] | Effect of deficiency[9][83] | Human disease | Chromosomal location | Protein structure |
|---|---|---|---|---|---|---|---|
| SERPINA1 | α1-antitrypsin | Extracellular | Inhibitor of human neutrophil elastase.[99] The C-terminal fragment of cleaved SERPINA1 may inhibit HIV-1 infection.[100] | Deficiency results in emphysema, polymerisation results in cirrhosis (serpinopathy).[11][101] | 14q32.1 | 1QLP, 7API, 1D5S | |
| SERPINA2 | Antitrypsin-related protein | Extracellular | Possible pseudogene.[102] | 14q32.1 | |||
| SERPINA3 | α1-antichymotrypsin | Extracellular | Inhibitor of cathepsin G.[103] Additional roles in chromatin condensation in hepatic cells.[104] | Mis-regulation results in Alzheimer's disease (serpinopathy).[105] | 14q32.1 | 1YXA, 2ACH | |
| SERPINA4 | Kallistatin | Extracellular | Inhibitor of kallikrein, regulator of vascular function.[106][107] | Depletion in hypertensive rats exacerbates renal and cardiovascular injury.[108] | 14q32.1 | ||
| SERPINA5 | Protein C inhibitor | Extracellular | Inhibitor of active protein C.[109] Intracellular role in preventing phagocytosis of bacteria.[110] | Knockout in male mice causes infertility.[111] Accumulation occurs in chronic active plaques in multiple sclerosis.[112] | 14q32.1 | 2OL2, 3B9F | |
| SERPINA6 | Transcortin | Extracellular | Non-inhibitory. Cortisol binding.[54] | 14q32.1 | 2V6D, 2VDX, 2VDY | ||
| SERPINA7 | Thyroxine-binding globulin | Extracellular | Non-inhibitory. Thyroxine binding.[55] | Deficiency causes hypothyroidism.[113][114] | Xq22.2 | 2CEO, 2RIV, 2RIW | |
| SERPINA8 | Angiotensinogen | Extracellular | Non-inhibitory, cleavage by renin results in release of angiotensin I.[115] | Knockout in mice causes hypotension.[116] | 1q42-q43 | 2X0B, 2WXW, 2WXX, 2WXY, 2WXZ, 2WY0, 2WY1 | |
| SERPINA9 | Centerin / GCET1 | Extracellular | Strongly expressed in most B-cell lymphomas.[117][118] | 14q32.1 | |||
| SERPINA10 | Protein Z-related protease inhibitor | Extracellular | Binds protein Z and inactivates factor Xa and factor XIa.[119] | 14q32.1 | 3F1S, 3H5C | ||
| SERPINA11 | – | Probably extracellular | Unknown | 14q32.13 | |||
| SERPINA12 | Vaspin | Extracellular | Inhibitor of Kallikrein-7. Insulin-sensitizing adipocytokine.[120] | High plasma levels associated with type II diabetes.[121] | 14q32.1 | 4IF8 | |
| SERPINA13 | – | Probably extracellular | Unknown | 14q32 | |||
| SERPINB1 | Monocyte neutrophil elastase inhibitor | Intracellular | Inhibitor of neutrophil elastase.[122] | Knockout in mice causes neutrophil survival defect and immune deficiency.[123] | 6p25 | 1HLE | |
| SERPINB2 | Plasminogen activator inhibitor-2 | Intracellular/extracellular | Inhibitor of extracellular uPA. Intracellular function unclear, but may protect against viral infection.[124] | Deficiency in mice reduces immune response to nematode infection.[125] Knockout in mice causes no obvious phenotype.[126] | 18q21.3 | 1BY7 | |
| SERPINB3 | Squamous cell carcinoma antigen-1 (SCCA-1) | Intracellular | Inhibitor of papain-like cysteine proteases[44] and cathepsins K, L and S.[127][128] | Knockout in mice of Serpinb3a (the murine homolog of both human SERPINB3 and SERPINB4) have reduced mucus production in a murine model of asthma.[129] | 18q21.3 | 2ZV6 | |
| SERPINB4 | Squamous cell carcinoma antigen-2 (SCCA-2) | Intracellular | Inhibitor of chymotrypsin-like serine proteases, cathepsin G and chymase.[128][130] | Knockout in mice of Serpinb3a (the murine homolog of both human SERPINB3 and SERPINB4) have reduced mucus production in a murine model of asthma.[129] | 18q21.3 | ||
| SERPINB5 | Maspin | Intracellular | Non-inhibitory, function unclear[131][132][133] (see also maspin) | Knockout in mice originally reported as lethal,[134] but subsequently shown to have no obvious phenotype.[133] Expression may be a prognostic indicator that reflects expression of a neighbouring tumour suppressor gene (the phosphatase PHLPP1).[133] | 18q21.3 | 1WZ9 | |
| SERPINB6 | PI-6 | Intracellular | Inhibitor of cathepsin G.[135] | Knockout in mice causes hearing loss[136] and mild neutropenia.[137] | Deficiency associated with hearing loss.[138] | 6p25 | |
| SERPINB7 | Megsin | Intracellular | Involved in megakaryocyte maturation.[139] | Over-expression in mice causes kidney disease.[140] Knockout in mice does not cause histological abnormalities.[140] | Mutations associated with Nagashima-type Palmoplantar Keratosis.[141] | 18q21.3 | |
| SERPINB8 | PI-8 | Intracellular | Possible inhibitor of furin.[142] | 18q21.3 | |||
| SERPINB9 | PI-9 | Intracellular | Inhibitor of the cytotoxic granule protease granzyme B.[143] | Knockout in mice causes immune dysfunction.[144][145] | 6p25 | ||
| SERPINB10 | Bomapin | Intracellular | Unknown[146] | Knockout in mice causes no obvious phenotype (C57/BL6; lab strain BC069938). | 18q21.3 | ||
| SERPINB11 | Intracellular | Unknown[147] | Murine Serpinb11 is an active inhibitor whereas the human orthalogue is inactive.[147] Deficiency in ponies is associated with hoof wall separation disease.[148] | 18q21.3 | |||
| SERPINB12 | Yukopin | Intracellular | Unknown[149] | 18q21.3 | |||
| SERPINB13 | Hurpin/Headpin | Intracellular | Inhibitor of papain-like cysteine proteases.[150] | 18q21.3 | |||
| SERPINC1 | Antithrombin | Extracellular | Inhibitor of coagulation, specifically factor X, factor IX and thrombin.[34] | Knockouts in mice are lethal.[151] | Deficiency results in thrombosis and other clotting disorders (serpinopathy).[152][153] | 1q23-q21 | 2ANT, 2ZNH, 1AZX, 1TB6, 2GD4, 1T1F |
| SERPIND1 | Heparin cofactor II | Extracellular | Inhibitor of thrombin.[154] | Knockouts in mice are lethal.[155] | 22q11 | 1JMJ, 1JMO | |
| SERPINE1 | Plasminogen activator inhibitor 1 | Extracellular | Inhibitor of thrombin, uPA and TPa.[156] | 7q21.3-q22 | 1DVN, 1OC0 | ||
| SERPINE2 | Glia derived nexin / Protease nexin I | Extracellular | Inhibitor of uPA and tPA.[157] | Abnormal expression leads to male infertility.[158] Knockout in mice causes epilepsy.[159] | 2q33-q35 | 4DY0 | |
| SERPINF1 | Pigment epithelium derived factor | Extracellular | Non-inhibitory, potent anti-angiogenic molecule.[160] PEDF has been reported to bind the glycosaminoglycan hyaluronan.[161] | Knockout in mice affects the vasculature and mass of the pancreas and the prostate.[160] Promotes Notch–dependent renewal of adult periventricular neural stem cells.[162] Mutations in humans cause osteogenesis imperfecta type VI.[81] | 17p13.3 | 1IMV | |
| SERPINF2 | α2-antiplasmin | Extracellular | Inhibitor of plasmin, inhibitor of fibrinolysis.[163] | Knockouts in mice show increased mice show increased fibrinolysis but no bleeding disorder.[164] | Deficiency causes a rare bleeding disorder.[165][166] | 17pter-p12 | 2R9Y |
| SERPING1 | Complement 1-inhibitor | Extracellular | Inhibitor of C1 esterase.[167] | Several polymorphisms associated with macular degeneration[168] and hereditary angeoedema.[169] | 11q11-q13.1 | 2OAY | |
| SERPINH1 | 47 kDa Heat shock protein (HSP47) | Intracellular | Non-inhibitory, molecular chaperone in collagen folding.[57] | Knockouts in mice are lethal.[170] | Mutation in humans causes severe osteogenesis imperfecta.[171][172] | 11p15 | 4AXY |
| SERPINI1 | Neuroserpin | Extracellular | Inhibitor of tPA, uPA and plasmin.[173] | Mutation causes FENIB dementia (serpinopathy).[174][175] | 3q26 | 1JJO, 3FGQ, 3F5N, 3F02 | |
| SERPINI2 | Pancpin | Extracellular | Unknown[176] | Deficiency in mice causes pancreatic insufficiency via acinar cell loss.[177] | 3q26 |
Specialised mammalian serpins
Many mammalian serpins have been identified that share no obvious orthology with a human serpin counterpart. Examples include numerous rodent serpins (particularly some of the murine intracellular serpins) as well as the uterine serpins. The term uterine serpin refers to members of the serpin A clade that are encoded by the SERPINA14 gene. Uterine serpins are produced by the endometrium of a restricted group of mammals in the Laurasiatheria clade under the influence of progesterone or estrogen.[178] They are probably not functional proteinase inhibitors and may function during pregnancy to inhibit maternal immune responses against the conceptus or to participate in transplacental transport.[179]
Insect
The Drosophila melanogaster genome contains 29 serpin encoding genes. Amino acid sequence analysis has placed 14 of these serpins in serpin clade Q and three in serpin clade K with the remaining twelve classified as orphan serpins not belonging to any clade.[180] The clade classification system is difficult to use for Drosophila serpins and instead a nomenclature system has been adopted that is based on the position of serpin genes on the Drosophila chromosomes. Thirteen of the Drosophila serpins occur as isolated genes in the genome (including Serpin-27A, see below), with the remaining 16 organised into five gene clusters that occur at chromosome positions 28D (2 serpins), 42D (5 serpins), 43A (4 serpins), 77B (3 serpins) and 88E (2 serpins).[180][181][182]
Studies on Drosophila serpins reveal that Serpin-27A inhibits the Easter protease (the final protease in the Nudel, Gastrulation Defective, Snake and Easter proteolytic cascade) and thus controls dorsoventral patterning. Easter functions to cleave Spätzle (a chemokine-type ligand), which results in toll-mediated signaling. As well as its central role in embryonic patterning, toll signaling is also important for the innate immune response in insects. Accordingly, serpin-27A also functions to control the insect immune response.[48][183][184] In Tenebrio molitor (a large beetle), a protein (SPN93) comprising two discrete tandem serpin domains functions to regulate the toll proteolytic cascade.[185]
Serpins have been found in tick saliva, suppressing T lymphocyte production and inhibiting expression of TNF-α, IFN-γ, and IL-6.[186]
Nematode
The genome of the nematode worm C. elegans contains 9 serpins, all of which lack signal sequences and so are likely intracellular.[187] However, only 5 of these serpins appear to function as protease inhibitors.[187] One, SRP-6, performs a protective function and guards against stress-induced calpain-associated lysosomal disruption. Further, SRP-6 inhibits lysosomal cysteine proteases released after lysosomal rupture. Accordingly, worms lacking SRP-6 are sensitive to stress. Most notably, SRP-6 knockout worms die when placed in water (the hypo-osmotic stress lethal phenotype or Osl). It has therefore been suggested that lysosomes play a general and controllable role in determining cell fate.[188]
Plant
Plant serpins were amongst the first members of the superfamily that were identified.[189] The serpin barley protein Z is highly abundant in barley grain, and one of the major protein components in beer. The genome of the model plant, Arabidopsis thaliana contain 18 serpin-like genes, although only 8 of these are full-length serpin sequences.
Plant serpins are potent inhibitors of mammalian chymotrypsin-like serine proteases in vitro, the best-studied example being barley serpin Zx (BSZx), which is able to inhibit trypsin and chymotrypsin as well as several blood coagulation factors.[190] However, close relatives of chymotrypsin-like serine proteases are absent in plants. The RCL of several serpins from wheat grain and rye contain poly-Q repeat sequences similar to those present in the prolamin storage proteins of the endosperm.[191][192] It has therefore been suggested that plant serpins may function to inhibit proteases from insects or microbes that would otherwise digest grain storage proteins. In support of this hypothesis, specific plant serpins have been identified in the phloem sap of pumpkin (CmPS-1)[193] and cucumber plants.[194][195] Although an inverse correlation between up-regulation of CmPS-1 expression and aphid survival was observed, in vitro feeding experiments revealed that recombinant CmPS-1 did not appear to affect insect survival.[193]
Alternative roles and protease targets for plant serpins have been proposed. The Arabidopsis serpin, AtSerpin1 (At1g47710; 3LE2), mediates set-point control over programmed cell death by targeting the 'Responsive to Desiccation-21' (RD21) papain-like cysteine protease.[53][196] AtSerpin1 also inhibits metacaspase-like proteases in vitro.[52] Two other Arabidopsis serpins, AtSRP2 (At2g14540) and AtSRP3 (At1g64030) appear to be involved in responses to DNA damage.[197]
Fungal
A single fungal serpin has been characterized to date: celpin from Piromyces spp. strain E2. Piromyces is a genus of anaerobic fungi found in the gut of ruminants and is important for digesting plant material. Celpin is predicted to be inhibitory and contains two N-terminal dockerin domains in addition to its serpin domain. Dockerins are commonly found in proteins that localise to the fungal cellulosome, a large extracellular multiprotein complex that breaks down cellulose.[38] It is therefore suggested that celpin may protect the cellulosome against plant proteases. Certain bacterial serpins similarly localize to the cellulosome.[198]
Prokaryotic
Predicted serpin genes are sporadically distributed in prokaryotes. In vitro studies on some of these molecules have revealed that they are able to inhibit proteases, and it is suggested that they function as inhibitors in vivo. Several prokaryote serpins are found in extremophiles. Accordingly, and in contrast to mammalian serpins, these molecules possess elevated resistance to heat denaturation.[199][200] The precise role of most bacterial serpins remains obscure, although Clostridium thermocellum serpin localises to the cellulosome. It is suggested that the role of cellulosome-associated serpins may be to prevent unwanted protease activity against the cellulosome.[198]
Viral
Serpins are also expressed by viruses as a way to evade the host's immune defense.[201] In particular, serpins expressed by pox viruses, including cow pox (vaccinia) and rabbit pox (myxoma), are of interest because of their potential use as novel therapeutics for immune and inflammatory disorders as well as transplant therapy.[202][203] Serp1 suppresses the TLR-mediated innate immune response and allows indefinite cardiac allograft survival in rats.[202][204] Crma and Serp2 are both cross-class inhibitors and target both serine (granzyme B; albeit weakly) and cysteine proteases (caspase 1 and caspase 8).[205][206] In comparison to their mammalian counterparts, viral serpins contain significant deletions of elements of secondary structure. Specifically, crmA lacks the D-helix as well as significant portions of the A- and E-helices.[207]
References
- ↑ 1.0 1.1 1.2 1.3 1.4 "The serpins are an expanding superfamily of structurally similar but functionally diverse proteins. Evolution, mechanism of inhibition, novel functions, and a revised nomenclature". The Journal of Biological Chemistry 276 (36): 33293–33296. September 2001. doi:10.1074/jbc.R100016200. PMID 11435447.
- ↑ "A Comprehensive Phylogenetic Analysis of the Serpin Superfamily". Molecular Biology and Evolution 38 (7): 2915–2929. June 2021. doi:10.1093/molbev/msab081. PMID 33744972.
- ↑ "Serpins: the superfamily of plasma serine proteinase inhibitors" (in English). Proteinase Inhibitors. Research monographs in cell and tissue physiology. 12. Amsterdam: Elsevier Science Publishers BV. 1986. pp. 403–420. ISBN 0-444-80763-2.
- ↑ "Serpins flex their muscle: I. Putting the clamps on proteolysis in diverse biological systems". The Journal of Biological Chemistry 285 (32): 24299–24305. August 2010. doi:10.1074/jbc.R110.112771. PMID 20498369.
- ↑ "Serpins flex their muscle: II. Structural insights into target peptidase recognition, polymerization, and transport functions". The Journal of Biological Chemistry 285 (32): 24307–24312. August 2010. doi:10.1074/jbc.R110.141408. PMID 20498368.
- ↑ 6.0 6.1 6.2 6.3 "Structure of a serpin-protease complex shows inhibition by deformation". Nature 407 (6806): 923–926. October 2000. doi:10.1038/35038119. PMID 11057674. Bibcode: 2000Natur.407..923H.
- ↑ 7.0 7.1 7.2 7.3 7.4 7.5 7.6 "Serpin structure, mechanism, and function". Chemical Reviews 102 (12): 4751–4804. December 2002. doi:10.1021/cr010170. PMID 12475206.
- ↑ 8.0 8.1 8.2 8.3 8.4 8.5 8.6 "Molecular gymnastics: serpin structure, folding and misfolding". Current Opinion in Structural Biology 16 (6): 761–768. December 2006. doi:10.1016/j.sbi.2006.10.005. PMID 17079131.
- ↑ 9.0 9.1 9.2 9.3 9.4 9.5 "An overview of the serpin superfamily". Genome Biology 7 (5): 216. 2006. doi:10.1186/gb-2006-7-5-216. PMID 16737556.
- ↑ 10.0 10.1 10.2 10.3 10.4 "What do dysfunctional serpins tell us about molecular mobility and disease?". Nature Structural Biology 2 (2): 96–113. February 1995. doi:10.1038/nsb0295-96. PMID 7749926.
- ↑ 11.0 11.1 11.2 11.3 11.4 11.5 11.6 11.7 11.8 "The discovery of α1-antitrypsin and its role in health and disease". Respiratory Medicine 105 (8): 1129–1139. August 2011. doi:10.1016/j.rmed.2011.02.002. PMID 21367592.
- ↑ 12.0 12.1 12.2 12.3 "Conformational disease". Lancet 350 (9071): 134–138. July 1997. doi:10.1016/S0140-6736(97)02073-4. PMID 9228977.
- ↑ "Untersuchungen uber die enzyme, Vergleichende Studie" (in de). Zeitschrift für Hygiene und Infektionskrankheiten (18): 83–89. December 1894. doi:10.1007/BF02216836. https://ia800708.us.archive.org/view_archive.php?archive=/22/items/crossref-pre-1909-scholarly-works/10.1007%252Fbf02214664.zip&file=10.1007%252Fbf02216836.pdf.
- ↑ "Zur Kenntnis der alpha-globulin des menschlichen normal serums" (in de). Zeitschrift für Naturforschung B 10 (8): 463. August 1955. doi:10.1515/znb-1955-0810.
- ↑ "Primary structure of antithrombin III (heparin cofactor): partial homology between alpha-1-antitrypsin and antithrombin III". The Physiological Inhibitors of Coagulation and Fibrinolysis. Amsterdam: Elsevier. 1979. pp. 43–54.
- ↑ "Carboxy terminal fragment of human alpha-1-antitrypsin from hydroxylamine cleavage: homology with antithrombin III". Biochemical and Biophysical Research Communications 91 (3): 1032–1037. December 1979. doi:10.1016/0006-291X(79)91983-1. PMID 316698. Bibcode: 1979BBRC...91.1032C.
- ↑ "A surprising new protein superfamily containing ovalbumin, antithrombin-III, and alpha 1-proteinase inhibitor". Biochemical and Biophysical Research Communications 95 (2): 864–871. July 1980. doi:10.1016/0006-291X(80)90867-0. PMID 6968211.
- ↑ "α1-Antitrypsin and the serpins: variation and countervariation". Trends in Biochemical Sciences 10: 20–24. 1985. doi:10.1016/0968-0004(85)90011-8.
- ↑ "The electrophoretic α1-globulin pattern of serum in α1-antitrypsin deficiency. 1963". Copd 10 (Suppl 1): 3–8. March 2013. doi:10.3109/15412555.2013.771956. PMID 23527532.
- ↑ "Cirrhosis associated with alpha-1-antitrypsin deficiency: a previously unrecognized inherited disorder". The Journal of Laboratory and Clinical Medicine 73 (6): 934–939. June 1969. PMID 4182334.
- ↑ "Alpha-1-antitrypsin: molecular abnormality of S variant". British Medical Journal 1 (6002): 130–131. January 1976. doi:10.1136/bmj.1.6002.130-a. PMID 1082356.
- ↑ "Amino acid substitution Glu leads to Lys alpha1-antitrypsin PiZ". FEBS Letters 65 (2): 195–197. June 1976. doi:10.1016/0014-5793(76)80478-4. PMID 1084290.
- ↑ "Structure and variation of human alpha 1-antitrypsin". Nature 298 (5872): 329–334. July 1982. doi:10.1038/298329a0. PMID 7045697. Bibcode: 1982Natur.298..329C.
- ↑ "Active site of alpha 1-antitrypsin: homologous site in antithrombin-III". Biochemical and Biophysical Research Communications 93 (2): 399–402. March 1980. doi:10.1016/0006-291X(80)91090-6. PMID 6966929.
- ↑ "Structural evidence for methionine at the reactive site of human alpha-1-proteinase inhibitor". The Journal of Biological Chemistry 253 (20): 7142–7144. October 1978. doi:10.1016/S0021-9258(17)34475-7. PMID 701239.
- ↑ "The thrombin cleavage site in bovine antithrombin". FEBS Letters 106 (2): 358–362. October 1979. doi:10.1016/0014-5793(79)80532-3. PMID 499520. Bibcode: 1979FEBSL.106..358J.
- ↑ "Inherited antithrombin deficiency causing thrombophilia". Thrombosis et Diathesis Haemorrhagica 13 (2): 516–530. June 1965. doi:10.1055/s-0038-1656297. PMID 14347873.
- ↑ 28.0 28.1 "Mutation of antitrypsin to antithrombin. alpha 1-antitrypsin Pittsburgh (358 Met leads to Arg), a fatal bleeding disorder". The New England Journal of Medicine 309 (12): 694–698. September 1983. doi:10.1056/NEJM198309223091203. PMID 6604220.
- ↑ 29.0 29.1 "Human alpha 1-proteinase inhibitor. Crystal structure analysis of two crystal modifications, molecular model and preliminary analysis of the implications for function". Journal of Molecular Biology 177 (3): 531–557. August 1984. doi:10.1016/0022-2836(84)90298-5. PMID 6332197.
- ↑ 30.0 30.1 "Crystal structure of ovalbumin as a model for the reactive centre of serpins". Nature 347 (6288): 99–102. September 1990. doi:10.1038/347099a0. PMID 2395463. Bibcode: 1990Natur.347...99S.
- ↑ 31.0 31.1 "Mobile reactive centre of serpins and the control of thrombosis". Nature 353 (6344): 576–578. October 1991. doi:10.1038/353576a0. PMID 1922367. Bibcode: 1991Natur.353..576C.
- ↑ "Structural basis of latency in plasminogen activator inhibitor-1". Nature 355 (6357): 270–273. January 1992. doi:10.1038/355270a0. PMID 1731226. Bibcode: 1992Natur.355..270M.
- ↑ "Antithrombin-III-Hamilton, Ala 382 to Thr: an antithrombin-III variant that acts as a substrate but not an inhibitor of alpha-thrombin and factor Xa". Blood 77 (10): 2185–2189. May 1991. doi:10.1182/blood.V77.10.2185.2185. PMID 2029579.
- ↑ 34.0 34.1 34.2 "Shape-shifting serpins--advantages of a mobile mechanism". Trends in Biochemical Sciences 31 (8): 427–435. August 2006. doi:10.1016/j.tibs.2006.06.005. PMID 16820297.
- ↑ "How serpins transport hormones and regulate their release". Seminars in Cell & Developmental Biology 62: 133–141. February 2017. doi:10.1016/j.semcdb.2016.12.007. PMID 28027946. https://www.repository.cam.ac.uk/handle/1810/263922.
- ↑ 36.0 36.1 36.2 "Phylogeny of the serpin superfamily: implications of patterns of amino acid conservation for structure and function". Genome Research 10 (12): 1845–1864. December 2000. doi:10.1101/gr.GR-1478R. PMID 11116082.
- ↑ 37.0 37.1 "Serpins in prokaryotes". Molecular Biology and Evolution 19 (11): 1881–1890. November 2002. doi:10.1093/oxfordjournals.molbev.a004012. PMID 12411597.
- ↑ 38.0 38.1 "A serpin in the cellulosome of the anaerobic fungus Piromyces sp. strain E2". Mycological Research 112 (Pt 8): 999–1006. August 2008. doi:10.1016/j.mycres.2008.01.021. PMID 18539447. https://zenodo.org/record/852424.
- ↑ "Implications of the three-dimensional structure of alpha 1-antitrypsin for structure and function of serpins". Biochemistry 28 (23): 8951–8966. November 1989. doi:10.1021/bi00449a001. PMID 2690952.
- ↑ 40.0 40.1 "Evolutionary families of peptidase inhibitors". The Biochemical Journal 378 (Pt 3): 705–716. March 2004. doi:10.1042/BJ20031825. PMID 14705960.
- ↑ "Families and clans of serine peptidases". Archives of Biochemistry and Biophysics 318 (2): 247–250. April 1995. doi:10.1006/abbi.1995.1227. PMID 7733651.
- ↑ "Evolutionary lines of cysteine peptidases". Biological Chemistry 382 (5): 727–733. May 2001. doi:10.1515/BC.2001.088. PMID 11517925.
- ↑ "Evidence that serpin architecture intrinsically supports papain-like cysteine protease inhibition: engineering alpha(1)-antitrypsin to inhibit cathepsin proteases". Biochemistry 41 (15): 4998–5004. April 2002. doi:10.1021/bi0159985. PMID 11939796.
- ↑ 44.0 44.1 "The reactive site loop of the serpin SCCA1 is essential for cysteine proteinase inhibition". Proceedings of the National Academy of Sciences of the United States of America 95 (23): 13465–13470. November 1998. doi:10.1073/pnas.95.23.13465. PMID 9811823. Bibcode: 1998PNAS...9513465S.
- ↑ 45.0 45.1 "X-ray crystal structure of MENT: evidence for functional loop-sheet polymers in chromatin condensation". The EMBO Journal 25 (13): 3144–3155. July 2006. doi:10.1038/sj.emboj.7601201. PMID 16810322.
- ↑ "DNA accelerates the inhibition of human cathepsin V by serpins". The Journal of Biological Chemistry 282 (51): 36980–36986. December 2007. doi:10.1074/jbc.M706991200. PMID 17923478.
- ↑ 47.0 47.1 "The serpin PN1 is a feedback regulator of FGF signaling in germ layer and primary axis formation". Development 142 (6): 1146–1158. March 2015. doi:10.1242/dev.113886. PMID 25758225.
- ↑ 48.0 48.1 48.2 "Spatial regulation of developmental signaling by a serpin". Developmental Cell 5 (6): 945–950. December 2003. doi:10.1016/S1534-5807(03)00338-1. PMID 14667416.
- ↑ "Regulation of pro-apoptotic leucocyte granule serine proteinases by intracellular serpins". Immunology and Cell Biology 77 (1): 47–57. February 1999. doi:10.1046/j.1440-1711.1999.00787.x. PMID 10101686.
- ↑ "Selective regulation of apoptosis: the cytotoxic lymphocyte serpin proteinase inhibitor 9 protects against granzyme B-mediated apoptosis without perturbing the Fas cell death pathway". Molecular and Cellular Biology 18 (11): 6387–6398. November 1998. doi:10.1128/mcb.18.11.6387. PMID 9774654.
- ↑ "Viral inhibition of inflammation: cowpox virus encodes an inhibitor of the interleukin-1 beta converting enzyme". Cell 69 (4): 597–604. May 1992. doi:10.1016/0092-8674(92)90223-Y. PMID 1339309.
- ↑ 52.0 52.1 "Serpin1 of Arabidopsis thaliana is a suicide inhibitor for metacaspase 9". Journal of Molecular Biology 364 (4): 625–636. December 2006. doi:10.1016/j.jmb.2006.09.010. PMID 17028019.
- ↑ 53.0 53.1 "Arabidopsis AtSerpin1, crystal structure and in vivo interaction with its target protease RESPONSIVE TO DESICCATION-21 (RD21)". The Journal of Biological Chemistry 285 (18): 13550–13560. April 2010. doi:10.1074/jbc.M109.095075. PMID 20181955.
- ↑ 54.0 54.1 54.2 "Corticosteroid-binding globulin, a structural basis for steroid transport and proteinase-triggered release". The Journal of Biological Chemistry 282 (40): 29594–29603. October 2007. doi:10.1074/jbc.M705014200. PMID 17644521.
- ↑ 55.0 55.1 55.2 55.3 55.4 "Structural mechanism for the carriage and release of thyroxine in the blood". Proceedings of the National Academy of Sciences of the United States of America 103 (36): 13321–13326. September 2006. doi:10.1073/pnas.0604080103. PMID 16938877. Bibcode: 2006PNAS..10313321Z.
- ↑ "Structure and properties of ovalbumin". Journal of Chromatography. B, Biomedical Sciences and Applications 756 (1–2): 189–198. May 2001. doi:10.1016/S0378-4347(01)00108-6. PMID 11419711.
- ↑ 57.0 57.1 57.2 "Interactions of heat shock protein 47 with collagen and the stress response: an unconventional chaperone model?". Life Sciences 87 (19–22): 579–586. November 2010. doi:10.1016/j.lfs.2010.09.024. PMID 20888348.
- ↑ "MENT, a heterochromatin protein that mediates higher order chromatin folding, is a new serpin family member". The Journal of Biological Chemistry 274 (9): 5626–5636. February 1999. doi:10.1074/jbc.274.9.5626. PMID 10026180.
- ↑ "Inhibitory conformation of the reactive loop of alpha 1-antitrypsin". Nature Structural Biology 3 (8): 676–681. August 1996. doi:10.1038/nsb0896-676. PMID 8756325.
- ↑ "The murine orthologue of human antichymotrypsin: a structural paradigm for clade A3 serpins". The Journal of Biological Chemistry 280 (52): 43168–43178. December 2005. doi:10.1074/jbc.M505598200. PMID 16141197.
- ↑ "Conformational changes in serpins: I. The native and cleaved conformations of alpha(1)-antitrypsin". Journal of Molecular Biology 296 (2): 685–699. February 2000. doi:10.1006/jmbi.1999.3520. PMID 10669617.
- ↑ "The anticoagulant activation of antithrombin by heparin". Proceedings of the National Academy of Sciences of the United States of America 94 (26): 14683–14688. December 1997. doi:10.1073/pnas.94.26.14683. PMID 9405673. Bibcode: 1997PNAS...9414683J.
- ↑ "Conformational changes in serpins: II. The mechanism of activation of antithrombin by heparin". Journal of Molecular Biology 301 (5): 1287–1305. September 2000. doi:10.1006/jmbi.2000.3982. PMID 10966821.
- ↑ "Structure of the antithrombin-thrombin-heparin ternary complex reveals the antithrombotic mechanism of heparin". Nature Structural & Molecular Biology 11 (9): 857–862. September 2004. doi:10.1038/nsmb811. PMID 15311269.
- ↑ "Antithrombin-S195A factor Xa-heparin structure reveals the allosteric mechanism of antithrombin activation". The EMBO Journal 25 (9): 2029–2037. May 2006. doi:10.1038/sj.emboj.7601089. PMID 16619025.
- ↑ "Fondaparinux: a synthetic heparin pentasaccharide as a new antithrombotic agent". Expert Opinion on Investigational Drugs 11 (3): 397–407. March 2002. doi:10.1517/13543784.11.3.397. PMID 11866668.
- ↑ "A synthetic antithrombin III binding pentasaccharide is now a drug! What comes next?". Angewandte Chemie 43 (24): 3118–3133. June 2004. doi:10.1002/anie.200300640. PMID 15199558. Bibcode: 2004ACIE...43.3118P.
- ↑ 68.0 68.1 "Stability of plasminogen activator inhibitor 1 (PAI-1)". Thrombosis and Haemostasis 62 (2): 748–751. September 1989. doi:10.1055/s-0038-1646895. PMID 2479113.
- ↑ "Latent antithrombin and its detection, formation and turnover in the circulation". Journal of Thrombosis and Haemostasis 2 (12): 2170–2177. December 2004. doi:10.1111/j.1538-7836.2004.01047.x. PMID 15613023.
- ↑ "The N terminus of the serpin, tengpin, functions to trap the metastable native state". EMBO Reports 8 (7): 658–663. July 2007. doi:10.1038/sj.embor.7400986. PMID 17557112.
- ↑ "A structural basis for loop C-sheet polymerization in serpins". Journal of Molecular Biology 376 (5): 1348–1359. March 2008. doi:10.1016/j.jmb.2007.12.050. PMID 18234218.
- ↑ "Hormone binding globulins undergo serpin conformational change in inflammation". Nature 336 (6196): 257–258. November 1988. doi:10.1038/336257a0. PMID 3143075. Bibcode: 1988Natur.336..257P.
- ↑ 73.0 73.1 73.2 "Endocytic receptor LRP together with tPA and PAI-1 coordinates Mac-1-dependent macrophage migration". The EMBO Journal 25 (9): 1860–1870. May 2006. doi:10.1038/sj.emboj.7601082. PMID 16601674.
- ↑ "Specificity of binding of the low density lipoprotein receptor-related protein to different conformational states of the clade E serpins plasminogen activator inhibitor-1 and proteinase nexin-1". The Journal of Biological Chemistry 284 (27): 17989–17997. July 2009. doi:10.1074/jbc.M109.009530. PMID 19439404.
- ↑ "Uptake of the necrotic serpin in Drosophila melanogaster via the lipophorin receptor-1". PLOS Genetics 5 (6). June 2009. doi:10.1371/journal.pgen.1000532. PMID 19557185.
- ↑ "Mechanisms of serpin dysfunction in disease". Expert Reviews in Molecular Medicine 8 (31): 1–19. December 2006. doi:10.1017/S1462399406000184. PMID 17156576.
- ↑ "Worldwide racial and ethnic distribution of alpha1-antitrypsin deficiency: summary of an analysis of published genetic epidemiologic surveys". Chest 122 (5): 1818–1829. November 2002. doi:10.1378/chest.122.5.1818. PMID 12426287.
- ↑ "Effects of mutations in the hinge region of serpins". Biochemistry 32 (30): 7650–7657. August 1993. doi:10.1021/bi00081a008. PMID 8347575.
- ↑ "Antithrombins Wibble and Wobble (T85M/K): archetypal conformational diseases with in vivo latent-transition, thrombosis, and heparin activation". Blood 92 (8): 2696–2706. October 1998. doi:10.1182/blood.V92.8.2696. PMID 9763552.
- ↑ 80.0 80.1 80.2 "Inactive conformation of the serpin alpha(1)-antichymotrypsin indicates two-stage insertion of the reactive loop: implications for inhibitory function and conformational disease". Proceedings of the National Academy of Sciences of the United States of America 97 (1): 67–72. January 2000. doi:10.1073/pnas.97.1.67. PMID 10618372. Bibcode: 2000PNAS...97...67G.
- ↑ 81.0 81.1 "Mutations in SERPINF1 cause osteogenesis imperfecta type VI". Journal of Bone and Mineral Research 26 (12): 2798–2803. December 2011. doi:10.1002/jbmr.487. PMID 21826736.
- ↑ "Human plasminogen activator inhibitor-1 (PAI-1) deficiency: characterization of a large kindred with a null mutation in the PAI-1 gene". Blood 90 (1): 204–208. July 1997. doi:10.1182/blood.V90.1.204. PMID 9207454.
- ↑ 83.0 83.1 83.2 83.3 83.4 83.5 "Update of the human and mouse SERPIN gene superfamily". Human Genomics 7 (1). October 2013. doi:10.1186/1479-7364-7-22. PMID 24172014.
- ↑ 84.0 84.1 "The mechanism of Z alpha 1-antitrypsin accumulation in the liver". Nature 357 (6379): 605–607. June 1992. doi:10.1038/357605a0. PMID 1608473. Bibcode: 1992Natur.357..605L.
- ↑ "Endoplasmic reticulum-associated degradation (ERAD) and autophagy cooperate to degrade polymerogenic mutant serpins". The Journal of Biological Chemistry 284 (34): 22793–22802. August 2009. doi:10.1074/jbc.M109.027102. PMID 19549782.
- ↑ 86.0 86.1 86.2 "Crystal structure of a stable dimer reveals the molecular basis of serpin polymerization". Nature 455 (7217): 1255–1258. October 2008. doi:10.1038/nature07394. PMID 18923394. Bibcode: 2008Natur.455.1255Y.
- ↑ 87.0 87.1 "The structural diversity in α1-antitrypsin misfolding". EMBO Reports 12 (10): 983–984. September 2011. doi:10.1038/embor.2011.187. PMID 21921939.
- ↑ 88.0 88.1 "Molecular basis of α1-antitrypsin deficiency revealed by the structure of a domain-swapped trimer". EMBO Reports 12 (10): 1011–1017. September 2011. doi:10.1038/embor.2011.171. PMID 21909074.
- ↑ "Importance of the release of strand 1C to the polymerization mechanism of inhibitory serpins". Protein Science 6 (1): 89–98. January 1997. doi:10.1002/pro.5560060110. PMID 9007980.
- ↑ "A novel monoclonal antibody to characterize pathogenic polymers in liver disease associated with alpha1-antitrypsin deficiency". Hepatology 52 (3): 1078–1088. September 2010. doi:10.1002/hep.23760. PMID 20583215.
- ↑ "alpha1-Antitrypsin deficiency . 6: new and emerging treatments for alpha1-antitrypsin deficiency". Thorax 59 (10): 904–909. October 2004. doi:10.1136/thx.2003.006551. PMID 15454659.
- ↑ "Expanding the clinical indications for α(1)-antitrypsin therapy". Molecular Medicine 18 (6): 957–970. September 2012. doi:10.2119/molmed.2011.00196. PMID 22634722.
- ↑ "Hereditary alpha-1-antitrypsin deficiency and its clinical consequences". Orphanet Journal of Rare Diseases 3. June 2008. doi:10.1186/1750-1172-3-16. PMID 18565211.
- ↑ "Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells". Nature 478 (7369): 391–394. October 2011. doi:10.1038/nature10424. PMID 21993621. Bibcode: 2011Natur.478..391Y.
- ↑ "Small molecules block the polymerization of Z alpha1-antitrypsin and increase the clearance of intracellular aggregates". Journal of Medicinal Chemistry 50 (22): 5357–5363. November 2007. doi:10.1021/jm070687z. PMID 17918823.
- ↑ "Automated high-content live animal drug screening using C. elegans expressing the aggregation prone serpin α1-antitrypsin Z". PLOS ONE 5 (11). November 2010. doi:10.1371/journal.pone.0015460. PMID 21103396. Bibcode: 2010PLoSO...515460G.
- ↑ "Aeropin from the extremophile Pyrobaculum aerophilum bypasses the serpin misfolding trap". The Journal of Biological Chemistry 282 (37): 26802–26809. September 2007. doi:10.1074/jbc.M705020200. PMID 17635906.
- ↑ "Serpin protease inhibitors in plant biology". Physiologia Plantarum 145 (1): 95–102. May 2012. doi:10.1111/j.1399-3054.2011.01540.x. PMID 22085334. Bibcode: 2012PPlan.145...95F.
- ↑ "Alpha1-antitrypsin deficiency". Lancet 365 (9478): 2225–2236. 2005. doi:10.1016/S0140-6736(05)66781-5. PMID 15978931.
- ↑ "Discovery and optimization of a natural HIV-1 entry inhibitor targeting the gp41 fusion peptide". Cell 129 (2): 263–275. April 2007. doi:10.1016/j.cell.2007.02.042. PMID 17448989.
- ↑ "The molecular and cellular pathology of α₁-antitrypsin deficiency". Trends in Molecular Medicine 20 (2): 116–127. February 2014. doi:10.1016/j.molmed.2013.10.007. PMID 24374162.
- ↑ "Sequence diversity at the proximal 14q32.1 SERPIN subcluster: evidence for natural selection favoring the pseudogenization of SERPINA2". Molecular Biology and Evolution 24 (2): 587–598. February 2007. doi:10.1093/molbev/msl187. PMID 17135331.
- ↑ "Alpha 1-antichymotrypsin". The International Journal of Biochemistry & Cell Biology 28 (9): 961–964. September 1996. doi:10.1016/1357-2725(96)00032-5. PMID 8930118.
- ↑ "Nuclear α1-antichymotrypsin promotes chromatin condensation and inhibits proliferation of human hepatocellular carcinoma cells". Gastroenterology 144 (4): 818–828.e4. April 2013. doi:10.1053/j.gastro.2012.12.029. PMID 23295442.
- ↑ "Multi-functional capability of proteins: alpha1-antichymotrypsin and the correlation with Alzheimer's disease". Journal of Alzheimer's Disease 4 (2): 115–122. April 2002. doi:10.3233/JAD-2002-4206. PMID 12214135.
- ↑ "Kallistatin is a potent new vasodilator". The Journal of Clinical Investigation 100 (1): 11–17. July 1997. doi:10.1172/JCI119502. PMID 9202051.
- ↑ "Kallistatin is a new inhibitor of angiogenesis and tumor growth". Blood 100 (9): 3245–3252. November 2002. doi:10.1182/blood-2002-01-0185. PMID 12384424.
- ↑ "Depletion of endogenous kallistatin exacerbates renal and cardiovascular oxidative stress, inflammation, and organ remodeling". American Journal of Physiology. Renal Physiology 303 (8): F1230–F1238. October 2012. doi:10.1152/ajprenal.00257.2012. PMID 22811485.
- ↑ "Protein C inhibitor, a serpin with functions in- and outside vascular biology". Thrombosis and Haemostasis 97 (3): 343–347. March 2007. doi:10.1160/th06-09-0488. PMID 17334499.
- ↑ "Phosphatidylethanolamine critically supports internalization of cell-penetrating protein C inhibitor". The Journal of Cell Biology 179 (4): 793–804. November 2007. doi:10.1083/jcb.200707165. PMID 18025309.
- ↑ "Disruption of the protein C inhibitor gene results in impaired spermatogenesis and male infertility". The Journal of Clinical Investigation 106 (12): 1531–1539. December 2000. doi:10.1172/JCI10768. PMID 11120760.
- ↑ "Proteomic analysis of active multiple sclerosis lesions reveals therapeutic targets". Nature 451 (7182): 1076–1081. February 2008. doi:10.1038/nature06559. PMID 18278032. Bibcode: 2008Natur.451.1076H.
- ↑ "Variations in thyroid hormone transport proteins and their clinical implications". Thyroid 2 (3): 237–245. 1992. doi:10.1089/thy.1992.2.237. PMID 1422238.
- ↑ "Clinical review: Central hypothyroidism: pathogenic, diagnostic, and therapeutic challenges". The Journal of Clinical Endocrinology and Metabolism 97 (9): 3068–3078. September 2012. doi:10.1210/jc.2012-1616. PMID 22851492.
- ↑ "The intracellular renin-angiotensin system: a new paradigm". Trends in Endocrinology and Metabolism 18 (5): 208–214. July 2007. doi:10.1016/j.tem.2007.05.001. PMID 17509892.
- ↑ "Angiotensinogen-deficient mice with hypotension". The Journal of Biological Chemistry 269 (50): 31334–31337. December 1994. doi:10.1016/S0021-9258(18)31697-1. PMID 7989296.
- ↑ "Expression of the serpin centerin defines a germinal center phenotype in B-cell lymphomas". American Journal of Clinical Pathology 130 (1): 117–126. July 2008. doi:10.1309/9QKE68QU7B825A3U. PMID 18550480.
- ↑ "An emerging role for Serine Protease Inhibitors in T lymphocyte immunity and beyond". Immunology Letters 152 (1): 65–76. April 2013. doi:10.1016/j.imlet.2013.04.004. PMID 23624075.
- ↑ "Characterization of the protein Z-dependent protease inhibitor". Blood 96 (9): 3049–3055. November 2000. doi:10.1182/blood.V96.9.3049. PMID 11049983.
- ↑ "Visceral adipose tissue-derived serine protease inhibitor: a unique insulin-sensitizing adipocytokine in obesity". Proceedings of the National Academy of Sciences of the United States of America 102 (30): 10610–10615. July 2005. doi:10.1073/pnas.0504703102. PMID 16030142. Bibcode: 2005PNAS..10210610H.
- ↑ "Higher vaspin levels in subjects with obesity and type 2 diabetes mellitus: a meta-analysis". Diabetes Research and Clinical Practice 106 (1): 88–94. October 2014. doi:10.1016/j.diabres.2014.07.026. PMID 25151227.
- ↑ "Sequence and molecular characterization of human monocyte/neutrophil elastase inhibitor". Proceedings of the National Academy of Sciences of the United States of America 89 (12): 5635–5639. June 1992. doi:10.1073/pnas.89.12.5635. PMID 1376927. Bibcode: 1992PNAS...89.5635R.
- ↑ "The neutrophil serine protease inhibitor serpinb1 preserves lung defense functions in Pseudomonas aeruginosa infection". The Journal of Experimental Medicine 204 (8): 1901–1909. August 2007. doi:10.1084/jem.20070494. PMID 17664292.
- ↑ "The serine proteinase inhibitor (serpin) plasminogen activation inhibitor type 2 protects against viral cytopathic effects by constitutive interferon alpha/beta priming". The Journal of Experimental Medicine 187 (11): 1799–1811. June 1998. doi:10.1084/jem.187.11.1799. PMID 9607921.
- ↑ "SerpinB2 is critical to Th2 immunity against enteric nematode infection". Journal of Immunology 190 (11): 5779–5787. June 2013. doi:10.4049/jimmunol.1200293. PMID 23630350.
- ↑ "The plasminogen activator inhibitor-2 gene is not required for normal murine development or survival". Proceedings of the National Academy of Sciences of the United States of America 96 (2): 686–691. January 1999. doi:10.1073/pnas.96.2.686. PMID 9892694. Bibcode: 1999PNAS...96..686D.
- ↑ "Squamous cell carcinoma antigen is a potent inhibitor of cysteine proteinase cathepsin L". FEBS Letters 359 (1): 78–80. February 1995. doi:10.1016/0014-5793(94)01456-b. PMID 7851535. Bibcode: 1995FEBSL.359...78T.
- ↑ 128.0 128.1 "SERPINB3 (serpin peptidase inhibitor, clade B (ovalbumin), member 3)". Atlas of Genetics and Cytogenetics in Oncology and Haematology 19 (3): 202–209. March 2015. doi:10.4267/2042/56413. PMID 25984243.
- ↑ 129.0 129.1 "A nonredundant role for mouse Serpinb3a in the induction of mucus production in asthma". The Journal of Allergy and Clinical Immunology 127 (1): 254–61, 261.e1–6. January 2011. doi:10.1016/j.jaci.2010.10.009. PMID 21126757.
- ↑ "Squamous cell carcinoma antigen 2 is a novel serpin that inhibits the chymotrypsin-like proteinases cathepsin G and mast cell chymase". The Journal of Biological Chemistry 272 (3): 1849–1855. January 1997. doi:10.1074/jbc.272.3.1849. PMID 8999871.
- ↑ "Maspin (SERPINB5) is an obligate intracellular serpin". The Journal of Biological Chemistry 285 (14): 10862–10869. April 2010. doi:10.1074/jbc.M109.073171. PMID 20123984.
- ↑ "Maspin, a serpin with tumor-suppressing activity in human mammary epithelial cells". Science 263 (5146): 526–529. January 1994. doi:10.1126/science.8290962. PMID 8290962. Bibcode: 1994Sci...263..526Z.
- ↑ 133.0 133.1 133.2 "Maspin is not required for embryonic development or tumour suppression". Nature Communications 5. 2014. doi:10.1038/ncomms4164. PMID 24445777. Bibcode: 2014NatCo...5.3164T.
- ↑ "Maspin plays an essential role in early embryonic development". Development 131 (7): 1479–1489. April 2004. doi:10.1242/dev.01048. PMID 14985257.
- ↑ "The intracellular serpin proteinase inhibitor 6 is expressed in monocytes and granulocytes and is a potent inhibitor of the azurophilic granule protease, cathepsin G". Blood 93 (6): 2089–2097. March 1999. doi:10.1182/blood.V93.6.2089.406k10_2089_2097. PMID 10068683.
- ↑ "Absence of SERPINB6A causes sensorineural hearing loss with multiple histopathologies in the mouse inner ear". The American Journal of Pathology 183 (1): 49–59. July 2013. doi:10.1016/j.ajpath.2013.03.009. PMID 23669344.
- ↑ "Targeted disruption of SPI3/Serpinb6 does not result in developmental or growth defects, leukocyte dysfunction, or susceptibility to stroke". Molecular and Cellular Biology 24 (9): 4075–4082. May 2004. doi:10.1128/MCB.24.9.4075-4082.2004. PMID 15082799.
- ↑ "A truncating mutation in SERPINB6 is associated with autosomal-recessive nonsyndromic sensorineural hearing loss". American Journal of Human Genetics 86 (5): 797–804. May 2010. doi:10.1016/j.ajhg.2010.04.004. PMID 20451170.
- ↑ "Overexpression of the serpin megsin induces progressive mesangial cell proliferation and expansion". The Journal of Clinical Investigation 109 (5): 585–593. March 2002. doi:10.1172/JCI14336. PMID 11877466.
- ↑ 140.0 140.1 "Megsin gene: its genomic analysis, pathobiological functions, and therapeutic perspectives". Current Genomics 8 (3): 203–208. May 2007. doi:10.2174/138920207780833856. PMID 18645605.
- ↑ "Nagashima-type palmoplantar keratosis: a common Asian type caused by SERPINB7 protease inhibitor deficiency". The Journal of Investigative Dermatology 134 (8): 2076–2079. August 2014. doi:10.1038/jid.2014.156. PMID 25029323.
- ↑ "Inhibition of soluble recombinant furin by human proteinase inhibitor 8". The Journal of Biological Chemistry 273 (4): 1851–1854. January 1998. doi:10.1074/jbc.273.4.1851. PMID 9442015.
- ↑ "A cytosolic granzyme B inhibitor related to the viral apoptotic regulator cytokine response modifier A is present in cytotoxic lymphocytes". The Journal of Biological Chemistry 271 (44): 27802–27809. November 1996. doi:10.1074/jbc.271.44.27802. PMID 8910377.
- ↑ "Serine protease inhibitor 6 protects cytotoxic T cells from self-inflicted injury by ensuring the integrity of cytotoxic granules". Immunity 24 (4): 451–461. April 2006. doi:10.1016/j.immuni.2006.02.002. PMID 16618603.
- ↑ "Serpinb9 (Spi6)-deficient mice are impaired in dendritic cell-mediated antigen cross-presentation". Immunology and Cell Biology 90 (9): 841–851. October 2012. doi:10.1038/icb.2012.29. PMID 22801574.
- ↑ "Expression of bomapin, a novel human serpin, in normal/malignant hematopoiesis and in the monocytic cell lines THP-1 and AML-193". Blood 91 (4): 1256–1262. February 1998. doi:10.1182/blood.V91.4.1256. PMID 9454755.
- ↑ 147.0 147.1 "SERPINB11 is a new noninhibitory intracellular serpin. Common single nucleotide polymorphisms in the scaffold impair conformational change". The Journal of Biological Chemistry 282 (34): 24948–24960. August 2007. doi:10.1074/jbc.M703182200. PMID 17562709.
- ↑ "SERPINB11 frameshift variant associated with novel hoof specific phenotype in Connemara ponies". PLOS Genetics 11 (4). April 2015. doi:10.1371/journal.pgen.1005122. PMID 25875171.
- ↑ "SERPINB12 is a novel member of the human ov-serpin family that is widely expressed and inhibits trypsin-like serine proteinases". The Journal of Biological Chemistry 276 (52): 49320–49330. December 2001. doi:10.1074/jbc.M108879200. PMID 11604408.
- ↑ "Hurpin is a selective inhibitor of lysosomal cathepsin L and protects keratinocytes from ultraviolet-induced apoptosis". Biochemistry 42 (24): 7381–7389. June 2003. doi:10.1021/bi027307q. PMID 12809493.
- ↑ "Complete antithrombin deficiency in mice results in embryonic lethality". The Journal of Clinical Investigation 106 (7): 873–878. October 2000. doi:10.1172/JCI10489. PMID 11018075.
- ↑ "Inherited antithrombin deficiency: a review". Haemophilia 14 (6): 1229–1239. November 2008. doi:10.1111/j.1365-2516.2008.01830.x. PMID 19141163.
- ↑ "Serpin structure, function and dysfunction". Journal of Thrombosis and Haemostasis 9 (Suppl 1): 26–34. July 2011. doi:10.1111/j.1538-7836.2011.04360.x. PMID 21781239.
- ↑ "Antithrombotic activity of dermatan sulfate in heparin cofactor II-deficient mice". Blood 104 (13): 3965–3970. December 2004. doi:10.1182/blood-2004-02-0598. PMID 15315969.
- ↑ "Strain-dependent embryonic lethality and exaggerated vascular remodeling in heparin cofactor II-deficient mice". The Journal of Clinical Investigation 117 (6): 1514–1526. June 2007. doi:10.1172/JCI27095. PMID 17549254.
- ↑ "Structure-function relationships of plasminogen activator inhibitor-1 and its potential as a therapeutic agent". Current Drug Targets 8 (9): 971–981. September 2007. doi:10.2174/138945007781662337. PMID 17896949.
- ↑ "Mice lacking protease nexin-1 show delayed structural and functional recovery after sciatic nerve crush". The Journal of Neuroscience 27 (14): 3677–3685. April 2007. doi:10.1523/JNEUROSCI.0277-07.2007. PMID 17409231.
- ↑ "Male fertility defects in mice lacking the serine protease inhibitor protease nexin-1". Proceedings of the National Academy of Sciences of the United States of America 98 (6): 3029–3033. March 2001. doi:10.1073/pnas.051630698. PMID 11248026. Bibcode: 2001PNAS...98.3029M.
- ↑ "Endogenous serine protease inhibitor modulates epileptic activity and hippocampal long-term potentiation". The Journal of Neuroscience 17 (12): 4688–4699. June 1997. doi:10.1523/JNEUROSCI.17-12-04688.1997. PMID 9169529.
- ↑ 160.0 160.1 "Pigment epithelium-derived factor regulates the vasculature and mass of the prostate and pancreas". Nature Medicine 9 (6): 774–780. June 2003. doi:10.1038/nm870. PMID 12740569.
- ↑ "Pigment epithelium-derived factor binds to hyaluronan. Mapping of a hyaluronan binding site". The Journal of Biological Chemistry 283 (48): 33310–33320. November 2008. doi:10.1074/jbc.M801287200. PMID 18805795.
- ↑ "Vascular niche factor PEDF modulates Notch-dependent stemness in the adult subependymal zone". Nature Neuroscience 12 (12): 1514–1523. December 2009. doi:10.1038/nn.2437. PMID 19898467.
- ↑ "On the mechanism of the reaction between human alpha 2-antiplasmin and plasmin". The Journal of Biological Chemistry 254 (18): 9291–9297. September 1979. doi:10.1016/S0021-9258(19)86843-6. PMID 158022.
- ↑ "Alpha2-antiplasmin gene deficiency in mice is associated with enhanced fibrinolytic potential without overt bleeding". Blood 93 (7): 2274–2281. April 1999. doi:10.1182/blood.V93.7.2274. PMID 10090937.
- ↑ "Alpha2-antiplasmin and its deficiency: fibrinolysis out of balance". Haemophilia 14 (6): 1250–1254. November 2008. doi:10.1111/j.1365-2516.2008.01766.x. PMID 19141165.
- ↑ "Congenital alpha(2)-plasmin inhibitor deficiencies: a review". British Journal of Haematology 114 (1): 4–10. July 2001. doi:10.1046/j.1365-2141.2001.02845.x. PMID 11472338.
- ↑ "C1 inhibitor serpin domain structure reveals the likely mechanism of heparin potentiation and conformational disease". The Journal of Biological Chemistry 282 (29): 21100–21109. July 2007. doi:10.1074/jbc.M700841200. PMID 17488724. Bibcode: 2007JBiCh.28221100B.
- ↑ "Complement analysis in the 21st century". Molecular Immunology 44 (16): 3838–3849. September 2007. doi:10.1016/j.molimm.2007.06.150. PMID 17768101.
- ↑ "The autoimmune side of hereditary angioedema: insights on the pathogenesis". Autoimmunity Reviews 14 (8): 665–669. August 2015. doi:10.1016/j.autrev.2015.03.006. PMID 25827463.
- ↑ "Embryonic lethality of molecular chaperone hsp47 knockout mice is associated with defects in collagen biosynthesis". The Journal of Cell Biology 150 (6): 1499–1506. September 2000. doi:10.1083/jcb.150.6.1499. PMID 10995453.
- ↑ "Osteogenesis imperfecta due to mutations in non-collagenous genes: lessons in the biology of bone formation". Current Opinion in Pediatrics 26 (4): 500–507. August 2014. doi:10.1097/MOP.0000000000000117. PMID 25007323.
- ↑ "Recessively inherited forms of osteogenesis imperfecta". Annual Review of Genetics 46: 475–497. 1 January 2012. doi:10.1146/annurev-genet-110711-155608. PMID 23145505.
- ↑ "The axonally secreted serine proteinase inhibitor, neuroserpin, inhibits plasminogen activators and plasmin but not thrombin". The Journal of Biological Chemistry 273 (4): 2312–2321. January 1998. doi:10.1074/jbc.273.4.2312. PMID 9442076.
- ↑ "Familial conformational diseases and dementias". Human Mutation 20 (1): 1–14. July 2002. doi:10.1002/humu.10100. PMID 12112652.
- ↑ "Protein misfolding and the serpinopathies". Prion 1 (1): 15–20. 1 March 2007. doi:10.4161/pri.1.1.3974. PMID 19164889.
- ↑ "Isolation and characterization of a novel human pancreas-specific gene, pancpin, that is down-regulated in pancreatic cancer cells". Genes, Chromosomes & Cancer 22 (3): 179–185. July 1998. doi:10.1002/(SICI)1098-2264(199807)22:3<179::AID-GCC3>3.0.CO;2-T. PMID 9624529.
- ↑ "Acinar cell apoptosis in Serpini2-deficient mice models pancreatic insufficiency". PLOS Genetics 1 (3). September 2005. doi:10.1371/journal.pgen.0010038. PMID 16184191.
- ↑ "The molecular phylogeny of uterine serpins and its relationship to evolution of placentation". FASEB Journal 24 (2): 526–537. February 2010. doi:10.1096/fj.09-138453. PMID 19825977.
- ↑ "Evolution and function of the uterine serpins (SERPINA14)". American Journal of Reproductive Immunology 64 (4): 265–274. October 2010. doi:10.1111/j.1600-0897.2010.00901.x. PMID 20678169.
- ↑ 180.0 180.1 "Tip of another iceberg: Drosophila serpins". Trends in Cell Biology 15 (12): 659–665. December 2005. doi:10.1016/j.tcb.2005.10.001. PMID 16260136.
- ↑ "A serpin that regulates immune melanization in the respiratory system of Drosophila". Developmental Cell 15 (4): 617–626. October 2008. doi:10.1016/j.devcel.2008.08.017. PMID 18854145.
- ↑ "Drosophila Serpin-28D regulates hemolymph phenoloxidase activity and adult pigmentation". Developmental Biology 323 (2): 189–196. November 2008. doi:10.1016/j.ydbio.2008.08.030. PMID 18801354.
- ↑ "Dorsoventral patterning: a serpin pinned down at last". Current Biology 14 (1): R16–R18. January 2004. doi:10.1016/j.cub.2003.12.015. PMID 14711428. Bibcode: 2004CBio...14..R16R.
- ↑ "A serpin regulates dorsal-ventral axis formation in the Drosophila embryo". Current Biology 13 (23): 2097–2102. December 2003. doi:10.1016/j.cub.2003.10.062. PMID 14654000. Bibcode: 2003CBio...13.2097L.
- ↑ "93-kDa twin-domain serine protease inhibitor (Serpin) has a regulatory function on the beetle Toll proteolytic signaling cascade". The Journal of Biological Chemistry 286 (40): 35087–35095. October 2011. doi:10.1074/jbc.M111.277343. PMID 21862574.
- ↑ Denisov, Stepan S.; Dijkgraaf, Ingrid (2021-10-13). "Immunomodulatory Proteins in Tick Saliva From a Structural Perspective" (in English). Frontiers in Cellular and Infection Microbiology 11. doi:10.3389/fcimb.2021.769574. ISSN 2235-2988. PMID 34722347.
- ↑ 187.0 187.1 "SRP-2 is a cross-class inhibitor that participates in postembryonic development of the nematode Caenorhabditis elegans: initial characterization of the clade L serpins". The Journal of Biological Chemistry 279 (15): 15448–15459. April 2004. doi:10.1074/jbc.M400261200. PMID 14739286.
- ↑ "An intracellular serpin regulates necrosis by inhibiting the induction and sequelae of lysosomal injury". Cell 130 (6): 1108–1119. September 2007. doi:10.1016/j.cell.2007.07.013. PMID 17889653.
- ↑ "Sequence homology between barley endosperm protein Z and protease inhibitors of the alpha-1-antitrypsin family". FEBS Lett. 180 (1): 89–94. 1985. doi:10.1016/0014-5793(85)80238-6. Bibcode: 1985FEBSL.180...89H.
- ↑ "Inhibition of coagulation factors by recombinant barley serpin BSZx". FEBS Letters 394 (2): 165–168. September 1996. doi:10.1016/0014-5793(96)00940-4. PMID 8843156. Bibcode: 1996FEBSL.394..165D.
- ↑ "Inhibitory serpins from rye grain with glutamine as P1 and P2 residues in the reactive center". FEBS Letters 488 (3): 149–153. January 2001. doi:10.1016/S0014-5793(00)02425-X. PMID 11163762. Bibcode: 2001FEBSL.488..149H.
- ↑ "Inhibitory serpins from wheat grain with reactive centers resembling glutamine-rich repeats of prolamin storage proteins. Cloning and characterization of five major molecular forms". The Journal of Biological Chemistry 275 (43): 33272–33279. October 2000. doi:10.1074/jbc.M004633200. PMID 10874043.
- ↑ 193.0 193.1 "Characterization of cucurbita maxima phloem serpin-1 (CmPS-1). A developmentally regulated elastase inhibitor". The Journal of Biological Chemistry 275 (45): 35122–35128. November 2000. doi:10.1074/jbc.M006060200. PMID 10960478.
- ↑ "Cucurbit phloem serpins are graft-transmissible and appear to be resistant to turnover in the sieve element-companion cell complex". Journal of Experimental Botany 56 (422): 3111–3120. December 2005. doi:10.1093/jxb/eri308. PMID 16246856.
- ↑ "Serpins in plants and green algae". Functional & Integrative Genomics 8 (1): 1–27. February 2008. doi:10.1007/s10142-007-0059-2. PMID 18060440.
- ↑ "Set-point control of RD21 protease activity by AtSerpin1 controls cell death in Arabidopsis". The Plant Journal 74 (3): 498–510. May 2013. doi:10.1111/tpj.12141. PMID 23398119. Bibcode: 2013PlJ....74..498L.
- ↑ "Serpin genes AtSRP2 and AtSRP3 are required for normal growth sensitivity to a DNA alkylating agent in Arabidopsis". BMC Plant Biology 9 (1). May 2009. doi:10.1186/1471-2229-9-52. PMID 19426562. Bibcode: 2009BMCPB...9...52A.
- ↑ 198.0 198.1 "The functional repertoire of prokaryote cellulosomes includes the serpin superfamily of serine proteinase inhibitors". Molecular Microbiology 60 (6): 1344–1354. June 2006. doi:10.1111/j.1365-2958.2006.05182.x. PMID 16796673.
- ↑ "The 1.5 A crystal structure of a prokaryote serpin: controlling conformational change in a heated environment". Structure 11 (4): 387–397. April 2003. doi:10.1016/S0969-2126(03)00057-1. PMID 12679017.
- ↑ "The high resolution crystal structure of a native thermostable serpin reveals the complex mechanism underpinning the stressed to relaxed transition". The Journal of Biological Chemistry 280 (9): 8435–8442. March 2005. doi:10.1074/jbc.M410206200. PMID 15590653.
- ↑ "Poxvirus immune modulators: functional insights from animal models". Virus Research 88 (1–2): 35–53. September 2002. doi:10.1016/S0168-1702(02)00119-3. PMID 12297326.
- ↑ 202.0 202.1 "Serpins, the vasculature, and viral therapeutics". Frontiers in Bioscience 11: 1042–1056. January 2006. doi:10.2741/1862. PMID 16146796.
- ↑ "Induction of indefinite cardiac allograft survival correlates with toll-like receptor 2 and 4 downregulation after serine protease inhibitor-1 (Serp-1) treatment". Transplantation 84 (9): 1158–1167. November 2007. doi:10.1097/01.tp.0000286099.50532.b0. PMID 17998872.
- ↑ "Serp-1, a viral anti-inflammatory serpin, regulates cellular serine proteinase and serpin responses to vascular injury". The Journal of Biological Chemistry 278 (20): 18563–18572. May 2003. doi:10.1074/jbc.M209683200. PMID 12637546.
- ↑ "Myxoma virus Serp2 is a weak inhibitor of granzyme B and interleukin-1beta-converting enzyme in vitro and unlike CrmA cannot block apoptosis in cowpox virus-infected cells". Journal of Virology 73 (8): 6394–6404. August 1999. doi:10.1128/JVI.73.8.6394-6404.1999. PMID 10400732.
- ↑ "Viral anti-inflammatory reagents: the potential for treatment of arthritic and vasculitic disorders". Endocrine, Metabolic & Immune Disorders Drug Targets 6 (4): 331–343. December 2006. doi:10.2174/187153006779025720. PMID 17214579.
- ↑ "Crystal structure of the apoptotic suppressor CrmA in its cleaved form". Structure 8 (7): 789–797. July 2000. doi:10.1016/S0969-2126(00)00165-9. PMID 10903953.
External links
- PDB Molecule of the Month Serpin
- Merops protease inhibitor claudication (Family I4)
- Serpins at the US National Library of Medicine Medical Subject Headings (MeSH)
- James Whisstock laboratory at Monash University
- Jim Huntington laboratory at University of Cambridge
- Frank Church laboratory at University of North Carolina at Chapel Hill
- Paul Declerck laboratory at Katholieke Universiteit Leuven
- Tom Roberts laboratory at University of Sydney
- Robert Fluhr laboratory at Weizmann Institute of Science
- Peter Gettins laboratory at University of Illinois at Chicago
- Overview of all the structural information available in the PDB for UniProt: P01009 (Human Alpha-1-antitrypsin) at the PDBe-KB.
 |