Biology:Dicer
Dicer, also known as endoribonuclease Dicer or helicase with RNase motif, is an enzyme that in humans is encoded by the DICER1 gene. Being part of the RNase III family, Dicer cleaves double-stranded RNA (dsRNA) and pre-microRNA (pre-miRNA) into short double-stranded RNA fragments called small interfering RNA and microRNA, respectively. These fragments are approximately 20–25 base pairs long with a two-base overhang on the 3′-end. Dicer facilitates the activation of the RNA-induced silencing complex (RISC), which is essential for RNA interference. RISC has a catalytic component Argonaute, which is an endonuclease capable of degrading messenger RNA (mRNA).[1]
Discovery
Dicer was given its name in 2001 by Stony Brook PhD student Emily Bernstein while conducting research in Gregory Hannon's lab at Cold Spring Harbor Laboratory. Bernstein sought to discover the enzyme responsible for generating small RNA fragments from double-stranded RNA. Dicer's ability to generate around 22 nucleotide RNA fragments was discovered by separating it from the RISC enzyme complex after initiating the RNAi pathway with dsRNA transfection. This experiment showed that RISC was not responsible for generating the observable small nucleotide fragments. Subsequent experiments testing RNase III family enzymes abilities to create RNA fragments narrowed the search to Drosophila CG4792, now named Dicer.[2]
Dicer orthologs are present in many other organisms.[3] In the moss Physcomitrella patens DCL1b, one of four DICER proteins, is not involved in miRNA biogenesis but in dicing miRNA target transcripts. Thus, a novel mechanism for regulation of gene expression, the epigenetic silencing of genes by miRNAs, was discovered.[4]
In terms of crystal structure, the first Dicer to be explored was that from the protozoan Giardia intestinalis. The work was done by Ian MacRae while conducting research as a postdoctoral fellow in Jennifer Doudna's lab at the University of California, Berkeley. A PAZ domain and two RNase III domains were discovered by X-ray crystallography. The protein size is 82 kDa, representing the conserved functional core that has subsequently been found in larger Dicer proteins in other organisms; for example, it is 219 kDa in humans. The difference in size from humans to G. intestinalis Dicer is due to at least five different domains being present within human Dicer. These domains are important in Dicer activity regulation, dsRNA processing, and RNA interference protein factor functioning.[5]
Functional domains
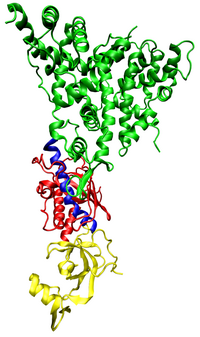
Human Dicer (also known as hsDicer or DICER1) is classified a Ribonuclease III because it cleaves double-stranded RNA. In addition to two RNaseIII domains, it contains a helicase domain, a PAZ (Piwi/Argonaute/Zwille) domain,[7][8] and two double stranded RNA binding domains (DUF283 and dsRBD).[5][9]
Current research suggests the PAZ domain is capable of binding the 2 nucleotide 3' overhang of dsRNA while the RNaseIII catalytic domains form a pseudo-dimer around the dsRNA to initiate cleavage of the strands. This results in a functional shortening of the dsRNA strand. The distance between the PAZ and RNaseIII domains is determined by the angle of the connector helix and influences the length of the micro RNA product.[6] The dsRBD domain binds the dsRNA, although the specific binding site of the domain has not been defined. It is possible that this domain works as part of a complex with other regulator proteins (TRBP in humans, R2D2, Loqs in Drosophila) in order to effectively position the RNaseIII domains and thus control the specificity of the sRNA products.[10] The helicase domain has been implicated in processing long substrates.[10]
Roles in RNA interference
MicroRNAs
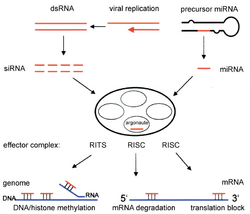
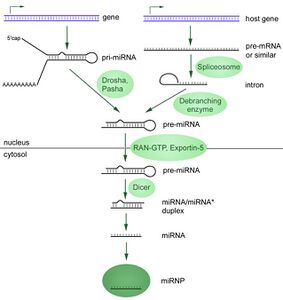
RNA interference is a process where the breakdown of RNA molecules into a microRNA (miRNA) inhibits gene expression of specific host mRNA sequences. miRNA is produced within the cell starting from primary miRNA (pri-miRNA) in the nucleus. These long sequences are cleaved into smaller precursor miRNA (pre-miRNA), which are usually 70 nucleotides with a hairpin structure. Pri-miRNA are identified by DGCR8 and cleaved by Drosha to form the pre-miRNA, a process that occurs in the nucleus. These pre-miRNA are then exported to the cytoplasm, where they are cleaved by Dicer to form mature miRNA.[12]
Small Interfering RNAs
Small interfering RNA (siRNA) are produced and function in a similar manner to miRNA by cleaving double-stranded RNA with Dicer into smaller fragments, 21 to 23 nucleotides in length.[10] Both miRNAs and siRNAs activate the RNA-induced silencing complex (RISC), which finds the complementary target mRNA sequence and cleaves the RNA using RNase.[13] This in turn silences the particular gene by RNA interference.[14] siRNAs and miRNAs differ in the fact that siRNAs are typically specific to the mRNA sequence while miRNAs aren't completely complementary to the mRNA sequence. miRNAs can interact with targets that have similar sequences, which inhibits translation of different genes.[15] In general, RNA interference is an essential part of normal processes within organisms such as humans, and it is an area being researched as a diagnostic and therapeutic tool for cancer targets.[12]
Roles in mammalian disease
Macular degeneration
Age related macular degeneration is a prominent cause of blindness in developed countries. Dicer's role in this disease became apparent after it was discovered that affected patients showed decreased levels of Dicer in their retinal pigment epithelium (RPE). Mice with Dicer knocked out, lacking Dicer only in their RPE, exhibited similar symptoms. However, other mice lacking important RNAi pathway proteins like Drosha and Pasha, did not have symptoms of macular degeneration as Dicer-knockout mice. This observation suggested a Dicer specific role in retinal health that was independent of the RNAi pathway and thus not a function of si/miRNA generation. A form of RNA called Alu RNA (the RNA transcripts of alu elements)) was found to be elevated in patients with insufficient Dicer levels. These non coding strands of RNA can loop forming dsRNA structures that would be degraded by Dicer in a healthy retina. However, with insufficient Dicer levels, the accumulation of alu RNA leads to the degeneration of RPE as a result of inflammation.[16][17]
Cancer
Altered miRNA expression profiles in malignant cancers suggest a pivotal role of miRNA and thus dicer in cancer development and prognosis. miRNAs can function as tumor suppressors and therefore their altered expression may result in tumorigenesis.[18] In analysis of lung and ovarian cancer, poor prognosis and decreased patient survival times correlate with decreased dicer and drosha expression. Decreased Dicer mRNA levels correlate with advanced tumor stage. However, high dicer expression in other cancers, like prostate[19] and esophageal, has been shown to correlate with poor patient prognosis. This discrepancy between cancer types suggests unique RNAi regulatory processes involving dicer differ amongst different tumor types.[12]
Dicer is also involved in DNA repair. DNA damage increases in mammalian cells with decreased Dicer expression as a result of decreased efficiency of DNA damage repair and other mechanisms. For example, siRNA from double strand breaks (produced by Dicer) may act as guides for protein complexes involved in the double strand break repair mechanisms and can also direct chromatin modifications. Additionally, miRNAs expression patterns change as a result of DNA damage caused by ionizing or ultraviolet radiation. RNAi mechanisms are responsible for transposon silencing and in their absence, as when Dicer is knocked out/down, can lead to activated transposons that cause DNA damage. Accumulation of DNA damage may result in cells with oncogenic mutations and thus the development of a tumor.[12]
Other conditions
Multinodular goiter with schwannomatosis has been shown to be an autosomal dominant condition associated with mutations in this gene.[20]
Viral pathogenesis
Infection by RNA viruses can trigger the RNAi cascade. It is likely Dicer is involved in viral immunity as viruses that infect both plant and animal cells contain proteins designed to inhibit the RNAi response. In humans, the viruses HIV-1, influenza, and vaccinia encode such RNAi suppressing proteins. Inhibition of dicer is beneficial to the virus as dicer is able to cleave viral dsRNA and load the product onto RISC resulting in targeted degradation of viral mRNA; thus fighting the infection. Another potential mechanism for viral pathogenesis is the blockade of dicer as a way to inhibit cellular miRNA pathways.[21]
Diagnostic and therapeutic applications
Dicer can be used to identify whether tumors are present within the body based on the expression level of the enzyme. A study showed that many patients that had cancer had decreased expression levels of Dicer. The same study showed that lower Dicer expression correlated with lower patient survival length.[12] Along with being a diagnostic tool, Dicer can be used for treating patients by injecting foreign siRNA intravenously to cause gene silencing.[22]
The siRNA was shown to be delivered in two ways in mammalian species such as mice. One way would be to directly inject into the system, which would not require Dicer function. Another way would be to introduce it by plasmids that encode for short hairpin RNA, which are cleaved by Dicer into siRNA.[23]
One of the advantages of using Dicer to produce siRNA therapeutically would be the specificity and diversity of targets it can affect compared to what is currently being used such as antibodies or small molecular inhibitors. In general, small molecular inhibitors are difficult in terms of specificity along with unendurable side effects. Antibodies are as specific as siRNA, but it is limited by only being able to be used against ligands or surface receptors. On the other hand, low efficiency of intracellular uptake is the main obstacle of injection of siRNA.[12] Injected SiRNA has poor stability in blood and causes stimulations of non-specific immunity.[24] Also, producing miRNA therapeutically lacks in specificity because only 6-8 nucleotide base pairing is required for miRNA to attach to mRNA.[25]
Dicer proteins in insects
In Drosophila, Dicer-1 generates microRNAs (miRNAs) by processing pre-miRNA, Dicer-2 is responsible for producing small interfering RNAs (siRNAs) from long double-stranded RNA (dsRNA).[26] Insects can use Dicer as a potent antiviral. This finding is especially significant given that mosquitoes are responsible for the transmission of many viral diseases including the potentially deadly arboviruses: West Nile virus, dengue fever and yellow fever.[27] While mosquitoes, more specifically the Aedes aegypti species, serve as the vectors for these viruses, they are not the intended host of the virus. Transmission occurs as a result of the female mosquito's need for vertebrate blood to develop her eggs. The RNAi pathway in insects is very similar to that of other animals; Dicer-2 cleaves viral RNA and loads it onto the RISC complex where one strand serves as a template for the production of RNAi products and the other is degraded. Insects with mutations leading to non-functional components of their RNAi pathway show increased viral loads for viruses they carry or increased susceptibility to viruses for which they are hosts. Similarly to humans, insect viruses have evolved mechanisms to avoid the RNAi pathway. As an example, Drosophila C virus encodes for protein 1A which binds to dsRNA thus protecting it from dicer cleavage as well as RISC loading. Heliothis virescens ascovirus 3a encodes an RNase III enzyme similar to the RNase III domains of dicer which may compete for dsRNA substrate as well as degrade siRNA duplexes to prevent RISC loading.[28]
Plant Dicer-like (DCL) proteins
Plant genomes encode Dicer-like proteins with similar functions and protein domains as animal and insect Dicers. For example, in the model organism Arabidopsis thaliana, four Dicer-like-encoding genes have been described and are designated DCL1 to DCL4; there are four Dicer family proteins encoded in a typical eudicot genome.[29] DCL1 is involved with miRNA generation and sRNA production from inverted repeats. DCL2 creates siRNA from cis-acting antisense transcripts which aid in viral immunity and defense.[30] DCL3 generates heterochromatic siRNAs which direct and reinforce DNA methylation and chromatin modifications; DCL4 is involved in trans-acting siRNA metabolism and transcript silencing at the post-transcriptional level.[30]
The genomes of most monocots including rice and maize encode a fifth DCL protein, Dicer-like 5 or DCL5.[31] Originally called DCL3b due to its emergence from a monocot-specific duplication of DCL3, DCL5 plays an important role in male reproductive development.[32][33] The name of DCL5 was originally proposed based on its divergence within rice and conservation in other grasses, which led to the speculation that it had a distinct function.[29]
See also
References
- ↑ "DICER1 rs3742330 and AGO1 rs636832 polymorphisms and acute lymphoblastic leukemia in Greek children and adolescents: A case-control study". Gene Reports 37 (December 2024). Sep 2024. doi:10.1016/j.genrep.2024.102043.
- ↑ "Role for a bidentate ribonuclease in the initiation step of RNA interference". Nature 409 (6818): 363–6. 2001. doi:10.1038/35053110. PMID 11201747. Bibcode: 2001Natur.409..363B.

- ↑ "Role of Dicer in Posttranscriptional RNA Silencing". RNA Interference. Current Topics in Microbiology and Immunology. 320. 2008. pp. 77–97. doi:10.1007/978-3-540-75157-1_4. ISBN 978-3-540-75156-4.
- ↑ "Transcriptional control of gene expression by microRNAs". Cell 140 (1): 111–22. Jan 2010. doi:10.1016/j.cell.2009.12.023. PMID 20085706.
- ↑ 5.0 5.1 "Structure of the human Dicer-TRBP complex by electron microscopy". Structure 17 (10): 1326–32. Oct 2009. doi:10.1016/j.str.2009.08.013. PMID 19836333.
- ↑ 6.0 6.1 "Structural basis for double-stranded RNA processing by Dicer". Science 311 (5758): 195–8. Jan 2006. doi:10.1126/science.1121638. PMID 16410517. Bibcode: 2006Sci...311..195M.
- ↑ "Entrez Gene: DICER1 Dicer1, Dcr-1 homolog (Drosophila)". https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=ShowDetailView&TermToSearch=23405.
- ↑ "Molecular cloning and characterization of a novel human gene (HERNA) which encodes a putative RNA-helicase". Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression 1490 (1–2): 163–9. Jan 2000. doi:10.1016/S0167-4781(99)00221-3. PMID 10786632.
- ↑ "Dicing and slicing: the core machinery of the RNA interference pathway". FEBS Letters 579 (26): 5822–9. Oct 2005. doi:10.1016/j.febslet.2005.08.079. PMID 16214139. Bibcode: 2005FEBSL.579.5822H.
- ↑ 10.0 10.1 10.2 "Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease". Molecular Cell 42 (2): 172–84. Apr 2011. doi:10.1016/j.molcel.2011.03.002. PMID 21419681.
- ↑ "An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells". Nature 404 (6775): 293–6. Mar 2000. doi:10.1038/35005107. PMID 10749213. Bibcode: 2000Natur.404..293H.
- ↑ 12.0 12.1 12.2 12.3 12.4 12.5 "The dicey role of Dicer: implications for RNAi therapy". Cancer Research 70 (7): 2571–4. Apr 2010. doi:10.1158/0008-5472.CAN-09-2536. PMID 20179193.
- ↑ "The contributions of dsRNA structure to Dicer specificity and efficiency". RNA 11 (5): 674–82. May 2005. doi:10.1261/rna.7272305. PMID 15811921.
- ↑ Watson JD (2008). Molecular Biology of the Gene. San Francisco, CA: Cold Spring Harbor Laboratory Press. pp. 641–648. ISBN 978-0-8053-9592-1.
- ↑ "MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms". Proceedings of the National Academy of Sciences of the United States of America 100 (17): 9779–84. Aug 2003. doi:10.1073/pnas.1630797100. PMID 12902540. Bibcode: 2003PNAS..100.9779Z.
- ↑ "Vision: Dicer leaps into view". Nature 471 (7338): 308–9. Mar 2011. doi:10.1038/471308a. PMID 21412326. Bibcode: 2011Natur.471..308M.
- ↑ "DICER1 loss and Alu RNA induce age-related macular degeneration via the NLRP3 inflammasome and MyD88". Cell 149 (4): 847–59. May 2012. doi:10.1016/j.cell.2012.03.036. PMID 22541070.
- ↑ "The role of dicer in DNA damage repair". International Journal of Molecular Sciences 13 (12): 16769–78. 2012. doi:10.3390/ijms131216769. PMID 23222681.
- ↑ "Up-regulation of dicer, a component of the MicroRNA machinery, in prostate adenocarcinoma". The American Journal of Pathology 169 (5): 1812–20. Nov 2006. doi:10.2353/ajpath.2006.060480. PMID 17071602.
- ↑ Rivera B, Nadaf J, Fahiminiya S, Apellaniz-Ruiz M, Saskin A, Chong AS, Sharma S, Wagener R, Revil T, Condello V, Harra Z, Hamel N, Sabbaghian N, Muchantef K, Thomas C, de Kock L, Hébert-Blouin MN, Bassenden AV, Rabenstein H, Mete O, Paschke R, Pusztaszeri MP, Paulus W, Berghuis A, Ragoussis J, Nikiforov YE, Siebert R, Albrecht S, Turcotte R, Hasselblatt M, Fabian MR, Foulkes WD (2019) DGCR8 microprocessor defect characterizes familial multinodular goiter with schwannomatosis. J Clin Invest
- ↑ "The interplay between virus infection and the cellular RNA interference machinery". FEBS Letters 580 (12): 2896–902. May 2006. doi:10.1016/j.febslet.2006.02.070. PMID 16563388. Bibcode: 2006FEBSL.580.2896B.
- ↑ "Intravenous injection of siRNA directed against hypoxia-inducible factors prolongs survival in a Lewis lung carcinoma cancer model". Cancer Gene Therapy 16 (3): 195–205. Mar 2009. doi:10.1038/cgt.2008.71. PMID 18818708.
- ↑ "Gene silencing by RNA interference is being used routinely to study gene function in cultured mammalian cells". Life Technologies. https://www.lifetechnologies.com/us/en/home/references/ambion-tech-support/rnai-sirna/tech-notes/performing-rnai-experiments-in-animals.html.
- ↑ "Cancer siRNA therapy by tumor selective delivery with ligand-targeted sterically stabilized nanoparticle". Nucleic Acids Research 32 (19): e149. 2004. doi:10.1093/nar/gnh140. PMID 15520458.
- ↑ "Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps". Nature 460 (7254): 479–86. Jul 2009. doi:10.1038/nature08170. PMID 19536157. Bibcode: 2009Natur.460..479C.
- ↑ "Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease". Molecular Cell 42 (2): 172–184. April 2011. doi:10.1016/j.molcel.2011.03.002. PMID 21419681.
- ↑ "Mosquito-borne Diseases". National Center for Infections Disease, Center for Disease Control and Prevention. https://www.cdc.gov/ncidod/diseases/list_mosquitoborne.htm.
- ↑ "The long and short of antiviral defense: small RNA-based immunity in insects". Current Opinion in Virology 7: 19–28. Aug 2014. doi:10.1016/j.coviro.2014.03.010. PMID 24732439.
- ↑ 29.0 29.1 Margis, Rogerio; Fusaro, Adriana F.; Smith, Neil A.; Curtin, Shaun J.; Watson, John M.; Finnegan, E. Jean; Waterhouse, Peter M. (2006-05-01). "The evolution and diversification of Dicers in plants" (in en). FEBS Letters 580 (10): 2442–2450. doi:10.1016/j.febslet.2006.03.072. ISSN 0014-5793. https://febs.onlinelibrary.wiley.com/doi/10.1016/j.febslet.2006.03.072.
- ↑ 30.0 30.1 Axtell, Michael J. (2013-04-29). "Classification and Comparison of Small RNAs from Plants" (in en). Annual Review of Plant Biology 64 (1): 137–159. doi:10.1146/annurev-arplant-050312-120043. ISSN 1543-5008. https://www.annualreviews.org/doi/10.1146/annurev-arplant-050312-120043.
- ↑ You, Chenjiang; Cui, Jie; Wang, Hui; Qi, Xinping; Kuo, Li-Yaung; Ma, Hong; Gao, Lei; Mo, Beixin et al. (2017-12-01). "Conservation and divergence of small RNA pathways and microRNAs in land plants" (in en). Genome Biology 18 (1). doi:10.1186/s13059-017-1291-2. ISSN 1474-760X. PMID 28835265. PMC 5569507. https://genomebiology.biomedcentral.com/articles/10.1186/s13059-017-1291-2.
- ↑ Song, Xianwei; Li, Pingchuan; Zhai, Jixian; Zhou, Ming; Ma, Lijia; Liu, Bin; Jeong, Dong‐Hoon; Nakano, Mayumi et al. (2012-02-01). "Roles of DCL4 and DCL3b in rice phased small RNA biogenesis" (in en). The Plant Journal 69 (3): 462–474. doi:10.1111/j.1365-313X.2011.04805.x. ISSN 0960-7412. https://onlinelibrary.wiley.com/doi/10.1111/j.1365-313X.2011.04805.x.
- ↑ Teng, Chong; Zhang, Han; Hammond, Reza; Huang, Kun; Meyers, Blake C.; Walbot, Virginia (2020-06-09). "Dicer-like 5 deficiency confers temperature-sensitive male sterility in maize" (in en). Nature Communications 11 (1). doi:10.1038/s41467-020-16634-6. ISSN 2041-1723. PMID 32518237. PMC 7283321. https://www.nature.com/articles/s41467-020-16634-6.
External links
- Overview of all the structural information available in the PDB for UniProt: Q9UPY3 (Human Endoribonuclease Dicer) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: Q8R418 (Mouse Endoribonuclease Dicer) at the PDBe-KB.
 |
