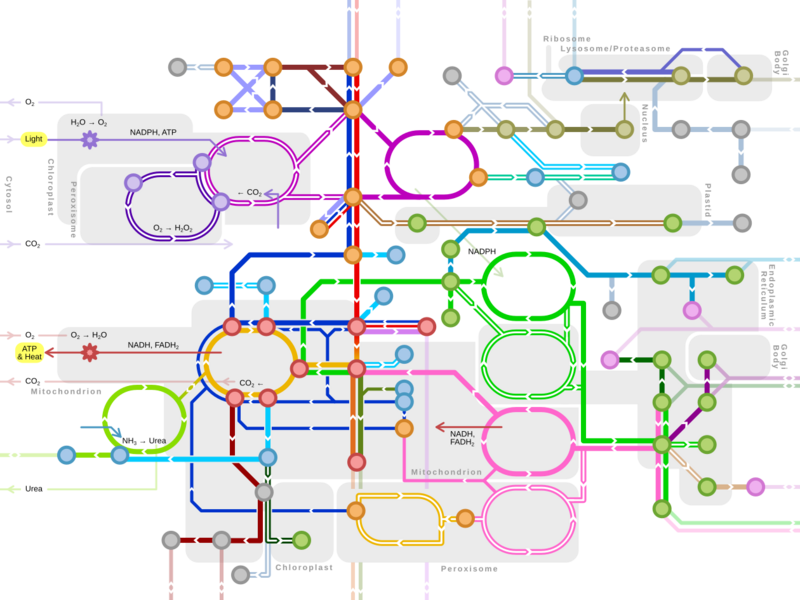
Earth:Biological carbon fixation

Biological carbon fixation or сarbon assimilation is the process by which inorganic carbon (particularly in the form of carbon dioxide) is converted to organic compounds by living organisms.[1] The compounds are then used to store energy and as structure for other biomolecules. Carbon is primarily fixed through photosynthesis, but some organisms use a process called chemosynthesis in the absence of sunlight.
Organisms that grow by fixing carbon are called autotrophs, which include photoautotrophs (which use sunlight), and lithoautotrophs (which use inorganic oxidation). Heterotrophs are not themselves capable of carbon fixation but are able to grow by consuming the carbon fixed by autotrophs or other heterotrophs. "Fixed carbon", "reduced carbon", and "organic carbon" may all be used interchangeably to refer to various organic compounds.[2] Chemosynthesis is carbon fixation driven by chemical energy, rather than from sunlight. Sulfur- and hydrogen-oxidizing bacteria often use the Calvin cycle or the reductive citric acid cycle.[3]
Net vs. gross CO2 fixation
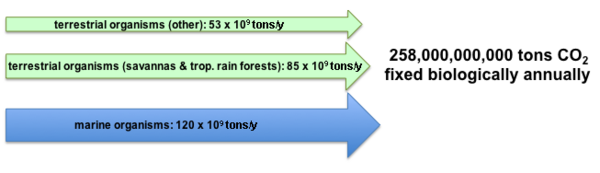
The primary form of inorganic carbon that is fixed is carbon dioxide (CO2). It is estimated that approximately 250 billion tons of carbon dioxide are converted by photosynthesis annually. The majority of the fixation occurs in terrestrial environments, especially the tropics. The gross amount of carbon dioxide fixed is much larger since approximately 40% is consumed by respiration following photosynthesis.[2][4] Historically it is estimated that approximately 2×1011 billion tons of carbon has been fixed since the origin of life.[5]
Overview of pathways
Seven autotrophic carbon fixation pathways are known. The Calvin cycle fixes carbon in the chloroplasts of plants and algae, and in the cyanobacteria. It also fixes carbon in the anoxygenic photosynthesis in one type of Pseudomonadota called purple bacteria, and in some non-phototrophic Pseudomonadota.[6]
Of the five other autotrophic pathways, two are known only in bacteria (the reductive citric acid cycle and the 3-hydroxypropionate cycle), two only in archaea (two variants of the 3-hydroxypropionate cycle), and one in both bacteria and archaea (the reductive acetyl CoA pathway).
List of pathways
Calvin cycle
The Calvin cycle accounts for 90% of biological carbon fixation. Consuming ATP and NADPH, the Calvin cycle in plants accounts for the preponderance of carbon fixation on land. In algae and cyanobacteria, it accounts for the preponderance of carbon fixation in the oceans. The Calvin cycle converts carbon dioxide into sugar, as triose phosphate (TP), which is glyceraldehyde 3-phosphate (GAP) together with dihydroxyacetone phosphate (DHAP):
- 3 CO2 + 12 e− + 12 H+ + Pi → TP + 4 H2O
An alternative perspective accounts for NADPH (source of e−) and ATP:
- 3 CO2 + 6 NADPH + 6 H+ + 9 ATP + 5 H2O → TP + 6 NADP+ + 9 ADP + 8 Pi
The formula for inorganic phosphate (Pi) is HOPO32− + 2H+. Formulas for triose and TP are C2H3O2-CH2OH and C2H3O2-CH2OPO32− + 2H+
Reverse Krebs cycle
The reverse Krebs cycle, also known as reverse TCA cycle (rTCA) or reductive citric acid cycle, is an alternative to the standard Calvin-Benson cycle for carbon fixation. It has been found in strict anaerobic or microaerobic bacteria (as Aquificales) and anaerobic archea. It was discovered by Evans, Buchanan and Arnon in 1966 working with the photosynthetic green sulfur bacterium Chlorobium limicola.[7] In particular, it is one of the most used pathways in hydrothermal vents by the Campylobacterota.[8] This feature is very important in oceans. Without it, there would be no primary production in aphotic environments, which would lead to habitats without life. So this kind of primary production is called "dark primary production".[9]
The cycle involves the biosynthesis of acetyl-CoA from two molecules of CO2.[10] The key steps of the reverse Krebs cycle are:
- Oxaloacetate to malate, using NADH + H+
- Fumarate to succinate, catalyzed by an oxidoreductase, Fumarate reductase
- Succinate to succinyl-CoA, an ATP dependent step
- Succinyl-CoA to alpha-ketoglutarate, using one molecule of CO2
- Alpha-ketoglutarate to isocitrate, using NADPH + H+ and another molecule of CO2
- Citrate converted into oxaloacetate and acetyl-CoA, this is an ATP dependent step and the key enzyme is the ATP citrate lyase
This pathway is cyclic due to the regeneration of the oxaloacetate.[11]
The bacteria Gammaproteobacteria and Riftia pachyptila switch from the Calvin-Benson cycle to the rTCA cycle in response to concentrations of H2S.[12]
Reductive acetyl CoA pathway
The reductive acetyl CoA pathway (CoA) pathway, also known as the Wood-Ljungdahl pathway uses CO2 as electron acceptor and carbon source, and H2 as an electron donor to form acetic acid.[13] This metabolism is wide spread within the phylum Bacillota, especially in the Clostridia.[14]
The pathway is also used by methanogens, which are mainly Euryarchaeota, and several anaerobic chemolithoautotrophs, such as sulfate-reducing bacteria and archaea. It is probably performed also by the Brocadiales, an order of Planctomycetota that oxidize ammonia in anaerobic condition.[10] [14][15] Hydrogenotrophic methanogenesis, which is only found in certain archaea and accounts for 80% of global methanogenesis, is also based on the reductive acetyl CoA pathway.
The Carbon Monoxide Dehydrogenase/Acetyl-CoA Synthase is the oxygen-sensitive enzyme that permits the reduction of CO2 to CO and the synthesis of acetyl-CoA in several reactions.[16]
One branch of this pathway, the methyl branch, is similar but non-homologous between bacteria and archaea. In this branch happens the reduction of CO2 to a methyl residue bound to a cofactor. The intermediates are formate for bacteria and formyl-methanofuran for archaea, and also the carriers, tetrahydrofolate and tetrahydropterins respectively in bacteria and archaea, are different, such as the enzymes forming the cofactor-bound methyl group.[10]
Otherwise, the carbonyl branch is homologous between the two domains and consists of the reduction of another molecule of CO2 to a carbonyl residue bound to an enzyme, catalyzed by the CO dehydrogenase/acetyl-CoA synthase. This key enzyme is also the catalyst for the formation of acetyl-CoA starting from the products of the previous reactions, the methyl and the carbonyl residues.[16]
This carbon fixation pathway requires only one molecule of ATP for the production of one molecule of pyruvate, which makes this process one of the main choice for chemolithoautotrophs limited in energy and living in anaerobic conditions.[10]
3-Hydroxypropionate bicycle
The 3-Hydroxypropionate bicycle, also known as 3-HP/malyl-CoA cycle, discovered only in 1989, is utilized by green non-sulfur phototrophs of Chloroflexaceae family, including the maximum exponent of this family Chloroflexus auranticus by which this way was discovered and demonstrated.[17] The 3-Hydroxipropionate bicycle is composed of two cycles and the name of this way comes from the 3-Hydroxyporopionate which corresponds to an intermediate characteristic of it.
The first cycle is a way of synthesis of glyoxylate. During this cycle, two equivalents of bicarbonate are fixed by the action of two enzymes: the Acetyl-CoA carboxylase catalyzes the carboxylation of the Acetyl-CoA to Malonyl-CoA and Propionyl-CoA carboxylase catalyses the carboxylation of propionyl-CoA to methylamalonyl-CoA. From this point a series of reactions lead to the formation of glyoxylate which will thus become part of the second cycle.[18][1]
In the second cycle, glyoxylate is approximately one equivalent of propionyl-CoA forming methylamalonyl-CoA. This, in turn, is then converted through a series of reactions into citramalyl-CoA. The citramalyl-CoA is split into pyruvate and Acetyl-CoA thanks to the enzyme MMC lyase. At this point the pyruvate is released, while the Acetyl-CoA is reused and carboxylated again at Malonyl-CoA thus reconstituting the cycle.[19]
A total of 19 reactions are involved in 3-hydroxypropionate bicycle and 13 multifunctional enzymes are used. The multifunctionality of these enzymes is an important feature of this pathway which thus allows the fixation of three bicarbonate molecules.[19]
It is a very expensive pathway: 7 ATP molecules are used for the synthesis of the new pyruvate and 3 ATP for the phosphate triose.[1]
An important characteristic of this cycle is that it allows the co-assimilation of numerous compounds making it suitable for the mixotrophic organisms.[1]
Cycles related to the 3-hydroxypropionate cycle
A variant of the 3-hydroxypropionate cycle was found to operate in the aerobic extreme thermoacidophile archaeon Metallosphaera sedula. This pathway is called the 3-hydroxypropionate/4-hydroxybutyrate cycle.[20]
Yet another variant of the 3-hydroxypropionate cycle is the dicarboxylate/4-hydroxybutyrate cycle. It was discovered in anaerobic archaea. It was proposed in 2008 for the hyperthermophile archeon Ignicoccus hospitalis.[21]
enoyl-CoA carboxylases/reductases
CO
2 fixation is catalyzed by enoyl-CoA carboxylases/reductases.[22]
Non-autotrophic pathways
Although no heterotrophs use carbon dioxide in biosynthesis, some carbon dioxide is incorporated in their metabolism.[23] Notably pyruvate carboxylase consumes carbon dioxide (as bicarbonate ions) as part of gluconeogenesis, and carbon dioxide is consumed in various anaplerotic reactions.
6-phosphogluconate dehydrogenase catalyzes the reductive carboxylation of ribulose 5-phosphate to 6-phosphogluconate in E. coli under elevated CO2 concentrations.[24]
Carbon isotope discrimination
Some carboxylases, particularly RuBisCO, preferentially bind the lighter carbon stable isotope carbon-12 over the heavier carbon-13. This is known as carbon isotope discrimination and results in carbon-12 to carbon-13 ratios in the plant that are higher than in the free air. Measurement of this ratio is important in the evaluation of water use efficiency in plants,[25][26][27] and also in assessing the possible or likely sources of carbon in global carbon cycle studies.
See also
References
- ↑ 1.0 1.1 1.2 1.3 "Ecological aspects of the distribution of different autotrophic CO2 fixation pathways". Applied and Environmental Microbiology 77 (6): 1925–36. March 2011. doi:10.1128/aem.02473-10. PMID 21216907. Bibcode: 2011ApEnM..77.1925B.
- ↑ 2.0 2.1 "Primary productivity of planet earth: biological determinants and physical constraints in terrestrial and aquatic habitats". Global Change Biology 7 (8): 849–882. 2001. doi:10.1046/j.1365-2486.2001.00448.x. Bibcode: 2001GCBio...7..849G.
- ↑ Encyclopedia of Microbiology. Academic Press. 2009. pp. 83–84. ISBN 9780123739445. https://books.google.com/books?id=rLhdW5YzuO4C&q=chemosynthesis+carbon+fixation&pg=RA2-PA83.
- ↑ Raghavendra, A. S. (2003-01-01), Thomas, Brian (ed.), "PHOTOSYNTHESIS AND PARTITIONING | C3 Plants", Encyclopedia of Applied Plant Sciences, Oxford: Elsevier, pp. 673–680, ISBN 978-0-12-227050-5, retrieved 2021-03-21
- ↑ Crockford, Peter W.; Bar On, Yinon M.; Ward, Luce M.; Milo, Ron; Halevy, Itay (November 2023). "The geologic history of primary productivity". Current Biology 33 (21): 4741–4750.e5. doi:10.1016/j.cub.2023.09.040. ISSN 0960-9822. https://doi.org/10.1016/j.cub.2023.09.040.
- ↑ "Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean". Science 333 (6047): 1296–300. September 2011. doi:10.1126/science.1203690. PMID 21885783. Bibcode: 2011Sci...333.1296S.
- ↑ "Alternative pathways of carbon dioxide fixation: insights into the early evolution of life?". Annual Review of Microbiology 65 (1): 631–58. 2011-10-13. doi:10.1146/annurev-micro-090110-102801. PMID 21740227.
- ↑ "Metagenome analysis of an extreme microbial symbiosis reveals eurythermal adaptation and metabolic flexibility". Proceedings of the National Academy of Sciences of the United States of America 105 (45): 17516–21. November 2008. doi:10.1073/pnas.0802782105. PMID 18987310. Bibcode: 2008PNAS..10517516G.
- ↑ "Is dark carbon fixation relevant for oceanic primary production estimates?". Biogeosciences. 2019-06-11. doi:10.5194/bg-2019-223. https://www.biogeosciences-discuss.net/bg-2019-223/bg-2019-223.pdf.
- ↑ 10.0 10.1 10.2 10.3 "Beyond the Calvin cycle: autotrophic carbon fixation in the ocean". Annual Review of Marine Science 3 (1): 261–89. 2011-01-15. doi:10.1146/annurev-marine-120709-142712. PMID 21329206. Bibcode: 2011ARMS....3..261H.
- ↑ "A reverse KREBS cycle in photosynthesis: consensus at last". Photosynthesis Research 24 (1): 47–53. April 1990. doi:10.1007/bf00032643. PMID 24419764.
- ↑ "Physiological proteomics of the uncultured endosymbiont of Riftia pachyptila". Science 315 (5809): 247–50. January 2007. doi:10.1126/science.1132913. OCLC 655249163. PMID 17218528. Bibcode: 2007Sci...315..247M.
- ↑ "A life with acetogens, thermophiles, and cellulolytic anaerobes". Annual Review of Microbiology 63 (1): 1–25. 2009. doi:10.1146/annurev.micro.091208.073617. PMID 19575555.
- ↑ 14.0 14.1 "Old acetogens, new light". Annals of the New York Academy of Sciences 1125 (1): 100–28. March 2008. doi:10.1196/annals.1419.016. PMID 18378590. Bibcode: 2008NYASA1125..100D.
- ↑ "Deciphering the evolution and metabolism of an anammox bacterium from a community genome". Nature 440 (7085): 790–4. April 2006. doi:10.1038/nature04647. PMID 16598256. Bibcode: 2006Natur.440..790S.
- ↑ 16.0 16.1 "Role of carbon monoxide dehydrogenase in the autotrophic pathway used by acetogenic bacteria". Proceedings of the National Academy of Sciences of the United States of America 81 (20): 6261–5. October 1984. doi:10.1073/pnas.81.20.6261. PMID 6436811. Bibcode: 1984PNAS...81.6261P.
- ↑ "Enzymes of a novel autotrophic CO2 fixation pathway in the phototrophic bacterium Chloroflexus aurantiacus, the 3-hydroxypropionate cycle". European Journal of Biochemistry 215 (3): 633–43. August 1993. doi:10.1111/j.1432-1033.1993.tb18074.x. PMID 8354269.
- ↑ "L-Malyl-coenzyme A lyase/beta-methylmalyl-coenzyme A lyase from Chloroflexus aurantiacus, a bifunctional enzyme involved in autotrophic CO2 fixation". Journal of Bacteriology 184 (21): 5999–6006. November 2002. doi:10.1128/jb.184.21.5999-6006.2002. PMID 12374834.
- ↑ 19.0 19.1 "Identifying the missing steps of the autotrophic 3-hydroxypropionate CO2 fixation cycle in Chloroflexus aurantiacus". Proceedings of the National Academy of Sciences of the United States of America 106 (50): 21317–22. December 2009. doi:10.1073/pnas.0908356106. PMID 19955419.
- ↑ "A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea". Science 318 (5857): 1782–6. December 2007. doi:10.1126/science.1149976. PMID 18079405. Bibcode: 2007Sci...318.1782B.
- ↑ "A dicarboxylate/4-hydroxybutyrate autotrophic carbon assimilation cycle in the hyperthermophilic Archaeum Ignicoccus hospitalis". Proceedings of the National Academy of Sciences of the United States of America 105 (22): 7851–6. June 2008. doi:10.1073/pnas.0801043105. PMID 18511565. Bibcode: 2008PNAS..105.7851H.
- ↑ Schwander, Thomas; Schada von Borzyskowski, Lennart; Burgener, Simon; Cortina, Niña Socorro; Erb, Tobias J. (2016). "A synthetic pathway for the fixation of carbon dioxide in vitro". Science 354 (6314): 900–904. doi:10.1126/science.aah5237. PMID 27856910. Bibcode: 2016Sci...354..900S.
- ↑ Nicole Kresge; Robert D. Simoni; Robert L. Hill (2005). "The Discovery of Heterotrophic Carbon Dioxide Fixation by Harland G. Wood". The Journal of Biological Chemistry 280 (18): e15. http://www.jbc.org/content/280/18/e15.full.
- ↑ "Awakening a latent carbon fixation cycle in Escherichia coli". Nature Communications 11 (1): 5812. November 2020. doi:10.1038/s41467-020-19564-5. PMID 33199707. Bibcode: 2020NatCo..11.5812S.
- ↑ "Genetic control of water use efficiency and leaf carbon isotope discrimination in sunflower (Helianthus annuus L.) subjected to two drought scenarios". PLOS ONE 9 (7): e101218. 3 July 2014. doi:10.1371/journal.pone.0101218. PMID 24992022. Bibcode: 2014PLoSO...9j1218A.
- ↑ "Carbon Isotope Discrimination and Photosynthesis". Annual Review of Plant Physiology and Plant Molecular Biology 40 (1): 503–537. June 1989. doi:10.1146/annurev.pp.40.060189.002443.
- ↑ "Carbon isotopes and water use efficiency: sense and sensitivity". Oecologia 155 (3): 441–54. March 2008. doi:10.1007/s00442-007-0932-7. PMID 18224341. Bibcode: 2008Oecol.155..441S.
Further reading
- "Diversity and evolutionary history of plastids and their hosts". American Journal of Botany 91 (10): 1481–93. October 2004. doi:10.3732/ajb.91.10.1481. PMID 21652304.
- "Chromalveolates and the evolution of plastids by secondary endosymbiosis". The Journal of Eukaryotic Microbiology 56 (1): 1–8. 2009. doi:10.1111/j.1550-7408.2008.00371.x. PMID 19335769. http://www.botany.ubc.ca/keeling/PDF/09KeelingTCS.pdf.
- "The endosymbiotic origin, diversification and fate of plastids". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 365 (1541): 729–48. March 2010. doi:10.1098/rstb.2009.0103. PMID 20124341.
- "Broad phylogenomic sampling and the sister lineage of land plants". PLOS ONE 7 (1): e29696. 2012. doi:10.1371/journal.pone.0029696. PMID 22253761. Bibcode: 2012PLoSO...7E9696T.
- "Evolution. Contemplating the first Plantae". Science 335 (6070): 809–10. February 2012. doi:10.1126/science.1218515. PMID 22344435. Bibcode: 2012Sci...335..809S.
- "Cyanophora paradoxa genome elucidates origin of photosynthesis in algae and plants". Science 335 (6070): 843–7. February 2012. doi:10.1126/science.1213561. PMID 22344442. Bibcode: 2012Sci...335..843P. http://cels.uri.edu/bio/lanelab/docs/Nic%20docs/PriceEtAl2012.pdf.