Chemistry:Propionyl-CoA
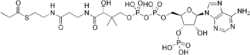
| |
| Names | |
|---|---|
| IUPAC name
3′-O-Phosphonoadenosine 5′-{(3R)-3-hydroxy-2,2-dimethyl-4-oxo-4-[(3-oxo-3-{[2-(propanoylsulfanyl)ethyl]amino}propyl)amino]butyl dihydrogen diphosphate}
| |
| Systematic IUPAC name
O1-{[(2R,3S,4R,5R)-5-(6-Amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methyl} O3-{(3R)-3-hydroxy-2,2-dimethyl-4-oxo-4-[(3-oxo-3-{[2-(propanoylsulfanyl)ethyl]amino}propyl)amino]butyl} dihydrogen diphosphate | |
| Other names
Propionyl Coenzyme A; Propanoyl Coenzyme A
| |
| Identifiers | |
3D model (JSmol)
|
|
| ChEBI | |
| ChemSpider | |
| MeSH | propionyl-coenzyme+A |
PubChem CID
|
|
| UNII | |
| |
| |
| Properties | |
| C24H40N7O17P3S | |
| Molar mass | 823.60 g/mol |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| Infobox references | |
Propionyl-CoA is a coenzyme A derivative of propionic acid. It is composed of a 24 total carbon chain (without the coenzyme, it is a 3 carbon structure) and its production and metabolic fate depend on which organism it is present in.[1] Several different pathways can lead to its production, such as through the catabolism of specific amino acids or the oxidation of odd-chain fatty acids.[2] It later can be broken down by propionyl-CoA carboxylase or through the methylcitrate cycle.[3] In different organisms, however, propionyl-CoA can be sequestered into controlled regions, to alleviate its potential toxicity through accumulation.[4] Genetic deficiencies regarding the production and breakdown of propionyl-CoA also have great clinical and human significance.[5]
Production
There are several different pathways through which propionyl-CoA can be produced:
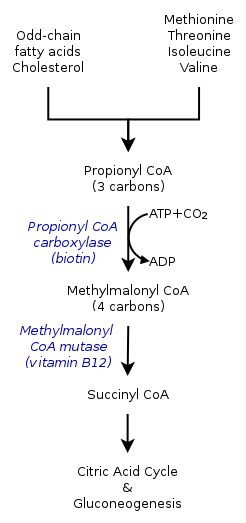
- Propionyl-CoA, a three-carbon structure, is considered to be a minor species of propionic acid. Therefore, odd-number chains of fatty acids are oxidized to yield both propionyl-CoA as well as acetyl-CoA. Propionyl-CoA is later converted into succinyl-CoA through biotin-dependant propionyl-CoA carboxylase (PCC) and b12-dependant methylmalonyl-CoA mutase (MCM), sequentially.[2]
- Propionyl-CoA is not only produced from the oxidation of odd-chain fatty acids, but also by the oxidation of amino acids including methionine, valine, isoleucine, and threonine. Furthermore, catabolism of amino acids can also be a result of the conversion of propionyl-CoA to methylmalonyl-CoA by propionyl-CoA carboxylase.[1]
- Cholesterol oxidation, which forms bile acids, also forms propionyl-CoA as a side product. In an experiment performed by Suld et al., when combining liver mitochondria and propionic acid with the addition of coenzyme A, labeled isotopes of psionic acid were degraded. However, following 5β-cholestane-3α,7α,12α,26-tetrol-26,27-C14 incubation, propionyl CoA was able to be rescued along with the formation of bile.[6]
Metabolic fate
The metabolic (catabolic fate) of propionyl-CoA depends on what environment it is being synthesized in. Therefore, propionyl-CoA in an anaerobic environment could have a different fate than that in an aerobic organism. The multiple pathways, either catabolism by propionyl-CoA carboxylase or methylcitrate synthase, also depend on the presence of various genes.[7]
Reaction with propionyl-CoA carboxylase
Within the citric acid cycle in humans, propionyl-CoA, which interacts with oxaloacetate to form methylcitrate, can also catalyzed into methylmalonyl-CoA through carboxylation by propionyl-CoA carboxylase (PCC). Methylmalonyl-CoA is later transformed to succinyl-CoA to be further used in the tricarboxylic acid cycle. PCC not only catalyzes the carboxylation of propionyl-CoA to methylmalonyl-CoA, but also acts on several different acyl-CoAs. Nevertheless, its highest binding affinity is to propionyl-CoA. It was further shown that propionyl-CoA transformation is inhibited by the absence of several TCA markers, such as glutamate. The mechanism is shown by the figure to the left.[2]
Mechanism
In mammals, propionyl-CoA is converted to (S)-methylmalonyl-CoA by propionyl-CoA carboxylase, a biotin-dependent enzyme also requiring bicarbonate and ATP.
This product is converted to (R)-methylmalonyl-CoA by methylmalonyl-CoA racemase.
(R)-Methylmalonyl-CoA is converted to succinyl-CoA, an intermediate in the tricarboxylic acid cycle, by methylmalonyl-CoA mutase, an enzyme requiring
cobalamin to catalyze the carbon-carbon bond migration.
The methylmalonyl-CoA mutase mechanism begins with the cleavage of the bond between the 5' CH2- of 5'-deoxyadenosyl and the cobalt, which is in its 3+ oxidation state (III), which produces a 5'-deoxyadenosyl radical and cobalamin in the reduced Co(II) oxidation state.
Next, this radical abstracts a hydrogen atom from the methyl group of methylmalonyl-CoA, which generates a methylmalonyl-CoA radical. It is believed that this radical forms a carbon-cobalt bond to the coenzyme, which is then followed by the rearrangement of the substrate's carbon skeleton, thus producing a succinyl-CoA radical. This radical then goes on to abstract a hydrogen from the previously produced 5'-deoxyadenosine, again creating a deoxyadenosyl radical, which attacks the coenzyme to reform the initial complex.
A defect in methylmalonyl-CoA mutase enzyme results in methylmalonic aciduria, a dangerous disorder that causes a lowering of blood pH.[8]
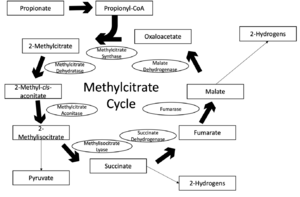
Methylcitrate cycle
Propionyl-CoA accumulation can prove toxic to different organisms. Since different cycles have been proposed regarding how propionyl-CoA is transformed into pyruvate, one studied mechanism is the methylcitrate cycle. The initial reaction is beta-oxidation to form the propionyl-CoA which is further broken down by the cycle. This pathway involves the enzymes both related to the methylcitrate cycle as well as the citric acid cycle. These all contribute to the overall reaction to detoxify the bacteria from harmful propionyl-CoA. It is also attributed as a resulting pathway due to the catabolism of fatty acids in mycobacteria.[3] In order to proceed, the prpC gene codes for methylcitrate synthase, and if not present, the methylcitrate cycle will not occur. Instead, catabolism proceeds through propionyl-CoA carboxylase.[7] This mechanism is shown below to the left along with the participating reactants, products, intermediates, and enzymes.
Bacterial metabolism
Mycobacterium tuberculosis metabolism
The oxidation of propionyl-CoA to form pyruvate is influenced by its necessity in Mycobacterium tuberculosis. Accumulation of propionyl-CoA can lead to toxic effects. In Mycobacterium tuberculosis, it has been suggested that the metabolism of propionyl-CoA is involved in cell wall biogenesis. A lack of such catabolism would therefore increase the susceptibility of the cell to various toxins, particularly to macrophage antimicrobial mechanisms. Another hypothesis regarding the fate of propionyl-CoA, in M. tuberculosisis, is that since propionyl-CoA is produced by beta odd chain fatty acid catabolism, the methylcitrate cycle is activated subsequently to negate any potential toxicity, acting as a buffering mechanism.[11]
Possible sequestration in R. sphaeroides
Propionyl-CoA has can have many adverse and toxic affects on different species, including bacterium. For example, inhibition of pyruvate dehydrogenase by an accumulation of propionyl-CoA in Rhodobacter sphaeroides can prove deadly. Furthermore, as with E. coli, an influx of propionyl-CoA in Myobacterial species can result in toxicity if not dealt with immediately. This toxicity is caused by a pathway involving the lipids that form the bacterial cell wall. Using esterification of long-chain fatty acids, excess propionyl-CoA can be sequestered and stored in the lipid, triacylglycerol (TAG), leading to regulation of elevated propionyl-CoA levels. Such a process of methyl branching of the fatty acids causes them to act as sinks for accumulating propion [4]
Escherichia coli metabolism
In an investigation performed by Luo et al., Escherichia coli strains were utilized to examine how the metabolism of propionyl-CoA could potentially lead to the production of 3-hydroxypropionic acid (3-HP). It was shown that a mutation in a key gene involved in the pathway, succinate CoA-transferase, led to a significant increase in 3-HP.[7] However, this is still a developing field and information on this topic is limited.[12]
Plant metabolism
Amino acid metabolism in plants has been deemed a controversial topic, due to the lack of concrete evidence for any particular pathway. However, it has been suggested that enzymes related to the production and use of propionyl-CoA are involved. Associated with this is the metabolism of isobutyryl-CoA. These two molecules are deemed to be intermediates in valine metabolism. As propionate consists in the form of propionyl-CoA, it was discovered that propionyl-CoA is converted to β-hydroxypropionate through a peroxisomal enzymatic β-oxidation pathway. Nevertheless, in the plant Arabidopsis, key enzymes in the conversion of valine to propionyl-CoA were not observed. Through different experiments performed by Lucas et al., it has been suggested that in plants, through peroxisomal enzymes, propionyl-CoA (and isobutyryl-CoA) are involved in the metabolism of many different substrates (currently being evaluated for identity), and not just valine.[13]
Fungi metabolism
Propionyl-CoA production through the catabolism of fatty acids is also associated with thioesterifcation. In a study concerning Aspergillus nidulans, it was found that with the inhibition of a methylcitrate synthase gene, mcsA, of the pathway described above, production of distinct polyketides was inhibited as well. Therefore, the utilization of propionyl-CoA through the methylcitrate cycle decreases its concentration, while subsequently increasing the concentration of polyketides. A polyketide is a structure commonly found in fungi that is made of acetyl- and malonyl-CoAs, providing a product with alternating carbonyl groups and methylene groups. Polyketides and polyketide derivatives are often highly structurally complex, and several are highly toxic. This has led to research on limiting polyketide toxicity to crops in agriculture through phytopathogenic fungi.[14]
Protein Propionylation
Propionyl-CoA is also a substrate for post-translational modification of proteins by reacting with lysine residues on proteins, a reaction called protein propionylation.[15][16] Due to structural similarities of Acetyl-CoA and Propionyl-CoA, propionylation reaction are thought to use many of the same enzymes used for protein acetylation.[16] Although functional consequences of protein propionylation are currently not completely understood, in vitro propionylation of the Propionyl-CoA Synthetase enzyme controls its activity.[17]
Human and clinical significance
Gen5
Similar to how plant peroxisomal enzymes bind propionyl-CoA and isobutyryl-CoA, Gen5, an acetyltransferase in humans, binds to propionyl-CoA and butyryl-CoA. These specifically bind to the catalytic domain of Gen5L2. This conserved acetyltransferase is responsible for the regulation of transcription by lysine acetylation of the histone N-terminal tails. This function of acetylation has a much higher reaction rate than propionylation or butyrylation. Because of the structure of propionyl-CoA, Gen5 distinguishes between different acyl-CoA molecules. In fact, it was found that the propyl group of butyrl-CoA cannot bind due to lack of stereospecificity to the active binding site of Gen5 due to the unsaturated acyl chains. On the other hand, the third carbon of propionyl-CoA can fit into the active site of Gen5 with the correct orientation.[18]
Propionic acidemia
In the neonatal developmental stages, propionic acidemia, which is a medical issue defined as the lack of propionyl-CoA carboxylase, can cause impairment, mental disability, and numerous other issues. This is caused by an accumulation of propionyl-CoA because it cannot be converted to methylmalonyl-CoA. Newborns are tested for elevated propionylcarnitine. Further ways of diagnosing this disease include urine samples. Medications used help to reverse and prevent recurring symptoms include using supplements to decrease propionate production.[5]
References
- ↑ 1.0 1.1 Dasgupta, Amitava (2019-01-01). "Chapter 2 - Biotin: Pharmacology, Pathophysiology, and Assessment of Biotin Status". in Dasgupta, Amitava. Biotin and Other Interferences in Immunoassays. Elsevier. pp. 17–35. doi:10.1016/B978-0-12-816429-7.00002-2. ISBN 9780128164297.
- ↑ 2.0 2.1 2.2 "Propionyl-CoA carboxylase - A review". Molecular Genetics and Metabolism 122 (4): 145–152. December 2017. doi:10.1016/j.ymgme.2017.10.002. PMID 29033250.
- ↑ 3.0 3.1 "Role of the methylcitrate cycle in propionate metabolism and detoxification in Mycobacterium smegmatis". Microbiology 153 (Pt 12): 3973–82. December 2007. doi:10.1099/mic.0.2007/011726-0. PMID 18048912.
- ↑ 4.0 4.1 "Loving the poison: the methylcitrate cycle and bacterial pathogenesis". Microbiology 164 (3): 251–259. March 2018. doi:10.1099/mic.0.000604. PMID 29458664.
- ↑ 5.0 5.1 "Propionic Acidemia". GeneReviews®. University of Washington, Seattle. 1993. http://www.ncbi.nlm.nih.gov/books/NBK92946/. Retrieved 2019-06-13.
- ↑ "Mechanism of formation of bile acids from cholesterol: oxidation of 5bita-choles-tane-3alpha,7alpha,12alpha-triol and formation of propionic acid from the side chain by rat liver mitochondria". The Journal of Biological Chemistry 237 (2): 338–44. February 1962. doi:10.1016/S0021-9258(18)93925-6. PMID 13918291. http://www.jbc.org/content/237/2/338.full.pdf.
- ↑ 7.0 7.1 7.2 "Production of 3-Hydroxypropionic Acid via the Propionyl-CoA Pathway Using Recombinant Escherichia coli Strains". PLOS ONE 11 (5): e0156286. 2016-05-26. doi:10.1371/journal.pone.0156286. PMID 27227837. Bibcode: 2016PLoSO..1156286L.
- ↑ "Comparative aspects of propionate metabolism". Comparative Biochemistry and Physiology. B, Comparative Biochemistry 92 (2): 227–31. 1989-01-01. doi:10.1016/0305-0491(89)90270-8. PMID 2647392.
- ↑ "The Nitrogen Regulator GlnR Directly Controls Transcription of the prpDBC Operon Involved in Methylcitrate Cycle in Mycobacterium smegmatis". Journal of Bacteriology 201 (8). April 2019. doi:10.1128/JB.00099-19. PMID 30745367.
- ↑ "Coupling Krebs cycle metabolites to signalling in immunity and cancer". Nature Metabolism 1 (1): 16–33. January 2019. doi:10.1038/s42255-018-0014-7. PMID 31032474.
- ↑ "Role of the methylcitrate cycle in Mycobacterium tuberculosis metabolism, intracellular growth, and virulence". Molecular Microbiology 60 (5): 1109–22. June 2006. doi:10.1111/j.1365-2958.2006.05155.x. PMID 16689789.
- ↑ "Multiple propionyl coenzyme A-supplying pathways for production of the bioplastic poly(3-hydroxybutyrate-co-3-hydroxyvalerate) in Haloferax mediterranei". Applied and Environmental Microbiology 79 (9): 2922–31. May 2013. doi:10.1128/AEM.03915-12. PMID 23435886. Bibcode: 2013ApEnM..79.2922H.
- ↑ "Peroxisomal metabolism of propionic acid and isobutyric acid in plants". The Journal of Biological Chemistry 282 (34): 24980–9. August 2007. doi:10.1074/jbc.M701028200. PMID 17580301.
- ↑ "Connection of propionyl-CoA metabolism to polyketide biosynthesis in Aspergillus nidulans". Genetics 168 (2): 785–94. October 2004. doi:10.1534/genetics.104.027540. PMID 15514053.
- ↑ "Lysine propionylation and butyrylation are novel post-translational modifications in histones". Molecular & Cellular Proteomics 6 (5): 812–9. May 2007. doi:10.1074/mcp.M700021-MCP200. PMID 17267393.
- ↑ 16.0 16.1 "Molecular characterization of propionyllysines in non-histone proteins". Molecular & Cellular Proteomics 8 (1): 45–52. January 2009. doi:10.1074/mcp.M800224-MCP200. PMID 18753126.
- ↑ "N-lysine propionylation controls the activity of propionyl-CoA synthetase". The Journal of Biological Chemistry 282 (41): 30239–45. October 2007. doi:10.1074/jbc.m704409200. PMID 17684016.
- ↑ "Structural basis for acyl-group discrimination by human Gcn5L2". Acta Crystallographica Section D 72 (Pt 7): 841–8. July 2016. doi:10.1107/S2059798316007907. PMID 27377381.
 |
