Biology:Prokaryote
A prokaryote (/proʊˈkærioʊt, -ət/, also spelled procaryote)[1] is a single-cell organism whose cell lacks a nucleus and other membrane-bound organelles.[2] The word prokaryote comes from the Ancient Greek πρό (Script error: The function "transl" does not exist.) 'before' and κάρυον (Script error: The function "transl" does not exist.) 'nut, kernel'.[3][4] In the two-empire system arising from the work of Édouard Chatton, prokaryotes were classified within the empire Prokaryota.[5] But in the three-domain system, based upon molecular analysis, prokaryotes are divided into two domains: Bacteria (formerly Eubacteria) and Archaea (formerly Archaebacteria). Organisms with nuclei are placed in a third domain, Eukaryota.[6]
Prokaryotes evolved before eukaryotes, and lack nuclei, mitochondria or most of the other distinct organelles that characterize the eukaryotic cell. It was once thought that prokaryotic cellular components were unenclosed within the cytoplasm except for an outer cell membrane, but bacterial microcompartments, which are thought to be quasi-organelles enclosed in protein shells (such as the encapsulin protein cages), have been discovered,[7][8] along with other prokaryotic organelles.[9] While being unicellular, some prokaryotes, such as cyanobacteria, may form colonies held together by biofilms, and large colonies can create multilayered microbial mats. Others, such as myxobacteria, have multicellular stages in their life cycles.[10] Prokaryotes are asexual, reproducing via binary fission without any fusion of gametes, although horizontal gene transfer may take place.
Molecular studies have provided insight into the evolution and interrelationships of the three domains of life.[11] The division between prokaryotes and eukaryotes reflects the existence of two very different levels of cellular organization; only eukaryotic cells have an enveloped nucleus that contains its chromosomal DNA, and other characteristic membrane-bound organelles including mitochondria. Distinctive types of prokaryotes include extremophiles and methanogens; these are common in some extreme environments.[2]
History
The distinction between prokaryotes and eukaryotes was firmly established by the microbiologists Roger Stanier and C. B. van Niel in their 1962 paper The concept of a bacterium[12] (though spelled procaryote and eucaryote there). That paper cites Édouard Chatton's 1937 book Titres et Travaux Scientifiques[13] for using those terms and recognizing the distinction. One reason for this classification was so that what was then often called blue-green algae (now called cyanobacteria) would not be classified as plants but grouped with bacteria.
Structure
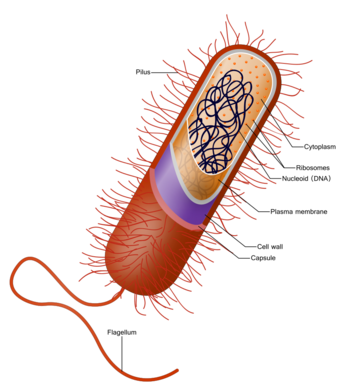
Prokaryotes have a prokaryotic cytoskeleton that is more primitive than that of the eukaryotes. Besides homologues of actin and tubulin (MreB and FtsZ), the helically arranged building-block of the flagellum, flagellin, is one of the most significant cytoskeletal proteins of bacteria, as it provides structural backgrounds of chemotaxis, the basic cell physiological response of bacteria. At least some prokaryotes also contain intracellular structures that can be seen as primitive organelles.
Membranous organelles (or intracellular membranes) are known in some groups of prokaryotes, such as vacuoles or membrane systems devoted to special metabolic properties, such as photosynthesis or chemolithotrophy. In addition, some species also contain carbohydrate-enclosed microcompartments, which have distinct physiological roles (e.g. carboxysomes or gas vacuoles).
Most prokaryotes are between 1 µm and 10 µm, but they can vary in size from 0.2 µm (Mycoplasma genitalium) to 750 µm (Thiomargarita namibiensis).
| Prokaryotic cell structure | Description |
|---|---|
| Flagellum (not always present) | Long, whip-like protrusion that aids cellular locomotion used by both gram-positive and gram-negative bacteria. |
| Cell membrane | Surrounds the cell's cytoplasm and regulates the flow of substances in and out of the cell. |
| Cell wall (except genera Mycoplasma and Thermoplasma) | Outer covering of most cells that protects the bacterial cell and gives it shape. |
| Cytoplasm | A gel-like substance composed mainly of water that also contains enzymes, salts, cell components, and various organic molecules. |
| Ribosome | Cell structures responsible for protein production. |
| Nucleoid | Area of the cytoplasm that contains the prokaryote's single DNA molecule. |
| Glycocalyx (only in some types of prokaryotes) | A glycoprotein-polysaccharide covering that surrounds the cell membranes. |
| Cytoplasmic inclusions | The inclusions such as ribosomes and larger masses scattered in the cytoplasmic matrix. |
Morphology
Prokaryotic cells have various shapes; the four basic shapes of bacteria are:[14]
- Cocci – A bacterium that is spherical or ovoid is called a coccus (Plural, cocci). e.g. Streptococcus, Staphylococcus.
- Bacilli – A bacterium with cylindrical shape called rod or a bacillus (Plural, bacilli).
- Spiral bacteria – Some rods twist into spiral shapes and are called spirilla (singular, spirillum).
- Vibrio – comma-shaped
The archaeon Haloquadratum has flat square-shaped cells.[15]
Reproduction
Bacteria and archaea reproduce through asexual reproduction, usually by binary fission. Genetic exchange and recombination still occur, but this is a form of horizontal gene transfer and is not a replicative process, simply involving the transference of DNA between two cells, as in bacterial conjugation.
DNA transfer
DNA transfer between prokaryotic cells occurs in bacteria and archaea, although it has been mainly studied in bacteria. In bacteria, gene transfer occurs by three processes. These are (1) bacterial virus (bacteriophage)-mediated transduction, (2) plasmid-mediated conjugation, and (3) natural transformation. Transduction of bacterial genes by bacteriophage appears to reflect an occasional error during intracellular assembly of virus particles, rather than an adaptation of the host bacteria. The transfer of bacterial DNA is under the control of the bacteriophage's genes rather than bacterial genes. Conjugation in the well-studied E. coli system is controlled by plasmid genes, and is an adaptation for distributing copies of a plasmid from one bacterial host to another. Infrequently during this process, a plasmid may integrate into the host bacterial chromosome, and subsequently transfer part of the host bacterial DNA to another bacterium. Plasmid mediated transfer of host bacterial DNA (conjugation) also appears to be an accidental process rather than a bacterial adaptation. File:Célula Procariota.ogv Natural bacterial transformation involves the transfer of DNA from one bacterium to another through the intervening medium. Unlike transduction and conjugation, transformation is clearly a bacterial adaptation for DNA transfer, because it depends on numerous bacterial gene products that specifically interact to perform this complex process.[16] For a bacterium to bind, take up and recombine donor DNA into its own chromosome, it must first enter a special physiological state called competence. About 40 genes are required in Bacillus subtilis for the development of competence.[17] The length of DNA transferred during B. subtilis transformation can be as much as a third to the whole chromosome.[18][19] Transformation is a common mode of DNA transfer, and 67 prokaryotic species are thus far known to be naturally competent for transformation.[20]
Among archaea, Halobacterium volcanii forms cytoplasmic bridges between cells that appear to be used for transfer of DNA from one cell to another.[21] Another archaeon, Sulfolobus solfataricus, transfers DNA between cells by direct contact. Frols et al. (2008) found[22] that exposure of S. solfataricus to DNA damaging agents induces cellular aggregation, and suggested that cellular aggregation may enhance DNA transfer among cells to provide increased repair of damaged DNA via homologous recombination.
Sociality
While prokaryotes are considered strictly unicellular, most can form stable aggregate communities.[23] When such communities are encased in a stabilizing polymer matrix ("slime"), they may be called "biofilms".[24] Cells in biofilms often show distinct patterns of gene expression (phenotypic differentiation) in time and space. Also, as with multicellular eukaryotes, these changes in expression often appear to result from cell-to-cell signaling, a phenomenon known as quorum sensing.
Biofilms may be highly heterogeneous and structurally complex and may attach to solid surfaces, or exist at liquid-air interfaces, or potentially even liquid-liquid interfaces. Bacterial biofilms are often made up of microcolonies (approximately dome-shaped masses of bacteria and matrix) separated by "voids" through which the medium (e.g., water) may flow easily. The microcolonies may join together above the substratum to form a continuous layer, closing the network of channels separating microcolonies. This structural complexity—combined with observations that oxygen limitation (a ubiquitous challenge for anything growing in size beyond the scale of diffusion) is at least partially eased by movement of medium throughout the biofilm—has led some to speculate that this may constitute a circulatory system[25] and many researchers have started calling prokaryotic communities multicellular (for example [26]). Differential cell expression, collective behavior, signaling, programmed cell death, and (in some cases) discrete biological dispersal[27] events all seem to point in this direction. However, these colonies are seldom if ever founded by a single founder (in the way that animals and plants are founded by single cells), which presents a number of theoretical issues. Most explanations of co-operation and the evolution of multicellularity have focused on high relatedness between members of a group (or colony, or whole organism). If a copy of a gene is present in all members of a group, behaviors that promote cooperation between members may permit those members to have (on average) greater fitness than a similar group of selfish individuals[28] (see inclusive fitness and Hamilton's rule).
Should these instances of prokaryotic sociality prove to be the rule rather than the exception, it would have serious implications for the way we view prokaryotes in general, and the way we deal with them in medicine.[29] Bacterial biofilms may be 100 times more resistant to antibiotics than free-living unicells and may be nearly impossible to remove from surfaces once they have colonized them.[30] Other aspects of bacterial cooperation—such as bacterial conjugation and quorum-sensing-mediated pathogenicity, present additional challenges to researchers and medical professionals seeking to treat the associated diseases.
Environment
Prokaryotes have diversified greatly throughout their long existence. The metabolism of prokaryotes is far more varied than that of eukaryotes, leading to many highly distinct prokaryotic types. For example, in addition to using photosynthesis or organic compounds for energy, as eukaryotes do, prokaryotes may obtain energy from inorganic compounds such as hydrogen sulfide. This enables prokaryotes to thrive in harsh environments as cold as the snow surface of Antarctica, studied in cryobiology, or as hot as undersea hydrothermal vents and land-based hot springs.
Prokaryotes live in nearly all environments on Earth. Some archaea and bacteria are extremophiles, thriving in harsh conditions, such as high temperatures (thermophiles) or high salinity (halophiles).[31] Many archaea grow as plankton in the oceans. Symbiotic prokaryotes live in or on the bodies of other organisms, including humans. Prokaryote have high populations in the soil - including the rhizosphere and rhizosheath. Soil prokaryotes are still heavily undercharacterized despite their easy proximity to humans and their tremendous economic importance to agriculture.[32]
Classification
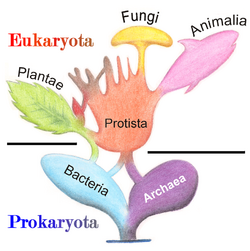
In 1977, Carl Woese proposed dividing prokaryotes into the Bacteria and Archaea (originally Eubacteria and Archaebacteria) because of the major differences in the structure and genetics between the two groups of organisms. Archaea were originally thought to be extremophiles, living only in inhospitable conditions such as extremes of temperature, pH, and radiation but have since been found in all types of habitats. The resulting arrangement of Eukaryota (also called "Eucarya"), Bacteria, and Archaea is called the three-domain system, replacing the traditional two-empire system.[33][34]
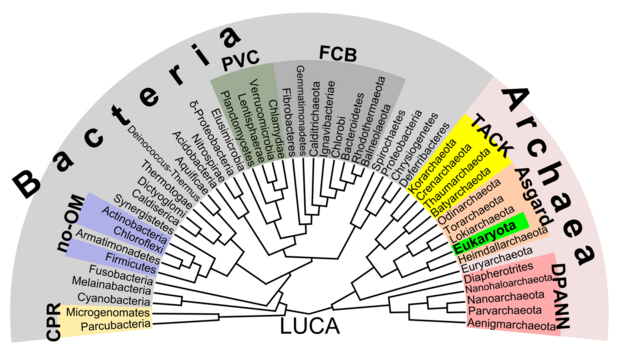
Phylogenetic tree
According to the phylogenetic analysis of Zhu (2019), the relationships could be the following:[35]
Evolution

A widespread current model of the evolution of the first living organisms is that these were some form of prokaryotes, which may have evolved out of protocells, while the eukaryotes evolved later in the history of life.[37] Some authors have questioned this conclusion, arguing that the current set of prokaryotic species may have evolved from more complex eukaryotic ancestors through a process of simplification.[38][39][40]
Others have argued that the three domains of life arose simultaneously, from a set of varied cells that formed a single gene pool.[41] This controversy was summarized in 2005:[42]
There is no consensus among biologists concerning the position of the eukaryotes in the overall scheme of cell evolution. Current opinions on the origin and position of eukaryotes span a broad spectrum including the views that eukaryotes arose first in evolution and that prokaryotes descend from them, that eukaryotes arose contemporaneously with eubacteria and archaebacteria and hence represent a primary line of descent of equal age and rank as the prokaryotes, that eukaryotes arose through a symbiotic event entailing an endosymbiotic origin of the nucleus, that eukaryotes arose without endosymbiosis, and that eukaryotes arose through a symbiotic event entailing a simultaneous endosymbiotic origin of the flagellum and the nucleus, in addition to many other models, which have been reviewed and summarized elsewhere.
The oldest known fossilized prokaryotes were laid down approximately 3.5 billion years ago, only about 1 billion years after the formation of the Earth's crust. Eukaryotes only appear in the fossil record later, and may have formed from endosymbiosis of multiple prokaryote ancestors. The oldest known fossil eukaryotes are about 1.7 billion years old. However, some genetic evidence suggests eukaryotes appeared as early as 3 billion years ago.[43]
While Earth is the only place in the universe where life is known to exist, some have suggested that there is evidence on Mars of fossil or living prokaryotes.[44][45] However, this possibility remains the subject of considerable debate and skepticism.[46][47]
Relationship to eukaryotes
The division between prokaryotes and eukaryotes is usually considered the most important distinction or difference among organisms. The distinction is that eukaryotic cells have a "true" nucleus containing their DNA, whereas prokaryotic cells do not have a nucleus.
Both eukaryotes and prokaryotes contain large RNA/protein structures called ribosomes, which produce protein, but the ribosomes of prokaryotes are smaller than those of eukaryotes. Mitochondria and chloroplasts, two organelles found in many eukaryotic cells, contain ribosomes similar in size and makeup to those found in prokaryotes.[48] This is one of many pieces of evidence that mitochondria and chloroplasts are descended from free-living bacteria. The endosymbiotic theory holds that early eukaryotic cells took in primitive prokaryotic cells by phagocytosis and adapted themselves to incorporate their structures, leading to the mitochondria and chloroplasts.
The genome in a prokaryote is held within a DNA/protein complex in the cytosol called the nucleoid, which lacks a nuclear envelope.[49] The complex contains a single, cyclic, double-stranded molecule of stable chromosomal DNA, in contrast to the multiple linear, compact, highly organized chromosomes found in eukaryotic cells. In addition, many important genes of prokaryotes are stored in separate circular DNA structures called plasmids.[3] Like Eukaryotes, prokaryotes may partially duplicate genetic material, and can have a haploid chromosomal composition that is partially replicated, a condition known as merodiploidy.[50]
Prokaryotes lack mitochondria and chloroplasts. Instead, processes such as oxidative phosphorylation and photosynthesis take place across the prokaryotic cell membrane.[51] However, prokaryotes do possess some internal structures, such as prokaryotic cytoskeletons.[52][53] It has been suggested that the bacterial phylum Planctomycetota has a membrane around the nucleoid and contains other membrane-bound cellular structures.[54] However, further investigation revealed that Planctomycetota cells are not compartmentalized or nucleated and, like other bacterial membrane systems, are interconnected.[55]
Prokaryotic cells are usually much smaller than eukaryotic cells.[3] Therefore, prokaryotes have a larger surface-area-to-volume ratio, giving them a higher metabolic rate, a higher growth rate, and as a consequence, a shorter generation time than eukaryotes.[3]
There is increasing evidence that the roots of the eukaryotes are to be found in (or at least next to) the archaean asgard group, perhaps Heimdallarchaeota (an idea which is a modern version of the 1984 eocyte hypothesis, eocytes being an old synonym for Thermoproteota, a taxon to be found nearby the then-unknown Asgard group)[56] For example, histones which usually package DNA in eukaryotic nuclei, have also been found in several archaean groups, giving evidence for homology. This idea might clarify the mysterious predecessor of eukaryotic cells (eucytes) which engulfed an alphaproteobacterium forming the first eucyte (LECA, last eukaryotic common ancestor) according to endosymbiotic theory. There might have been some additional support by viruses, called viral eukaryogenesis. The non-bacterial group comprising archaea and eukaryota was called Neomura by Thomas Cavalier-Smith in 2002.[57] However, in a cladistic view, eukaryota are archaea in the same sense as birds are dinosaurs because they evolved from the maniraptora dinosaur group. In contrast, archaea without eukaryota appear to be a paraphyletic group, just like dinosaurs without birds.
Prokaryotes may be split into two groups
Unlike the above assumption of a fundamental split between prokaryotes and eukaryotes, the most important difference between biota may be the division between bacteria and the rest (archaea and eukaryota).[56] For instance, DNA replication differs fundamentally between bacteria and archaea (including that in eukaryotic nuclei), and it may not be homologous between these two groups.[58] Moreover, ATP synthase, though common (homologous) in all organisms, differs greatly between bacteria (including eukaryotic organelles such as mitochondria and chloroplasts) and the archaea/eukaryote nucleus group. The last common antecessor of all life (called LUCA, last universal common ancestor) should have possessed an early version of this protein complex. As ATP synthase is obligate membrane bound, this supports the assumption that LUCA was a cellular organism. The RNA world hypothesis might clarify this scenario, as LUCA might have been a ribocyte (also called ribocell) lacking DNA, but with an RNA genome built by ribosomes as primordial self-replicating entities.[59] A Peptide-RNA world (also called RNP world) hypothesis has been proposed based on the idea that oligopeptides may have been built together with primordial nucleic acids at the same time, which also supports the concept of a ribocyte as LUCA. The feature of DNA as the material base of the genome might have then been adopted separately in bacteria and in archaea (and later eukaryote nuclei), presumably by help of some viruses (possibly retroviruses as they could reverse transcribe RNA to DNA).[60] As a result, prokaryota comprising bacteria and archaea may also be polyphyletic.
See also
References
- ↑ "Definition of PROCARYOTE" (in en). https://www.merriam-webster.com/dictionary/procaryote.
- ↑ 2.0 2.1 NC State University. "Prokaryotes: Single-celled Organisms". http://www.ncsu.edu/project/bio183de/Black/prokaryote/prokaryote1.html.
- ↑ 3.0 3.1 3.2 3.3 Campbell, N. "Biology:Concepts & Connections". Pearson Education. San Francisco: 2003.
- ↑ "prokaryote". Online Etymology Dictionary. http://www.etymonline.com/index.php?term=prokaryote&allowed_in_frame=0.
- ↑ Sapp, J. (2005). "The Prokaryote-Eukaryote Dichotomy: Meanings and Mythology". Microbiology and Molecular Biology Reviews 69 (2): 292–305. doi:10.1128/MMBR.69.2.292-305.2005. PMID 15944457.
- ↑ Coté, Gary; De Tullio, Mario (2010). "Beyond Prokaryotes and Eukaryotes: Planctomycetes and Cell Organization". Nature. http://www.nature.com/scitable/topicpage/beyond-prokaryotes-and-eukaryotes-planctomycetes-and-cell-14158971.
- ↑ "Protein structures forming the shell of primitive bacterial organelles". Science 309 (5736): 936–8. August 2005. doi:10.1126/science.1113397. PMID 16081736. Bibcode: 2005Sci...309..936K.
- ↑ "Cell biology of prokaryotic organelles". Cold Spring Harbor Perspectives in Biology 2 (10): a000422. October 2010. doi:10.1101/cshperspect.a000422. PMID 20739411.
- ↑ Murat, Dorothee; Byrne, Meghan; Komeili, Arash (2010-10-01). "Cell Biology of Prokaryotic Organelles". Cold Spring Harbor Perspectives in Biology 2 (10): a000422. doi:10.1101/cshperspect.a000422. PMID 20739411.
- ↑ "Coupling cell movement to multicellular development in myxobacteria". Nature Reviews. Microbiology 1 (1): 45–54. October 2003. doi:10.1038/nrmicro733. PMID 15040179.
- ↑ "Insights into the molecular evolution of HslU ATPase through biochemical and mutational analyses". PLOS ONE 9 (7): e103027. July 22, 2014. doi:10.1371/journal.pone.0103027. PMID 25050622. Bibcode: 2014PLoSO...9j3027S.
- ↑ "The concept of a bacterium". Archiv für Mikrobiologie 42 (1): 17–35. 1962. doi:10.1007/BF00425185. PMID 13916221.
- ↑ Chatton, Édouard (1937). Titres Et Travaux Scientifiques (1906-1937) De Edouard Chatton. Sète: Impr. E. Sottano..
- ↑ Microbiology. San Francisco: Pearson Benjamin Cummings. 2006. ISBN 978-0-8053-7693-7. https://archive.org/details/microbiology00robe_yj2.
- ↑ "Walsby's square bacterium: fine structure of an orthogonal procaryote". Journal of Bacteriology 148 (1): 352–60. October 1981. doi:10.1128/JB.148.1.352-360.1981. PMID 7287626.
- ↑ "DNA uptake during bacterial transformation". Nature Reviews. Microbiology 2 (3): 241–9. March 2004. doi:10.1038/nrmicro844. PMID 15083159.
- ↑ "Who's competent and when: regulation of natural genetic competence in bacteria". Trends in Genetics 12 (4): 150–5. April 1996. doi:10.1016/0168-9525(96)10014-7. PMID 8901420.
- ↑ "Incorporation of the whole chromosomal DNA in protoplast lysates into competent cells of Bacillus subtilis". Bioscience, Biotechnology, and Biochemistry 65 (4): 823–9. April 2001. doi:10.1271/bbb.65.823. PMID 11388459.
- ↑ "Fate of transforming bacterial genome following incorporation into competent cells of Bacillus subtilis: a continuous length of incorporated DNA". Journal of Bioscience and Bioengineering 101 (3): 257–62. March 2006. doi:10.1263/jbb.101.257. PMID 16716928.
- ↑ "Natural genetic transformation: prevalence, mechanisms and function". Research in Microbiology 158 (10): 767–78. December 2007. doi:10.1016/j.resmic.2007.09.004. PMID 17997281.
- ↑ "The mechanism of DNA transfer in the mating system of an archaebacterium". Science 245 (4924): 1387–9. September 1989. doi:10.1126/science.2818746. PMID 2818746. Bibcode: 1989Sci...245.1387R.
- ↑ "UV-inducible cellular aggregation of the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by pili formation". Molecular Microbiology 70 (4): 938–52. November 2008. doi:10.1111/j.1365-2958.2008.06459.x. PMID 18990182. https://www.rug.nl/research/portal/files/56956856/UV_inducible_cellular_aggregation_of_the_hyperthermophilic_archaeon_Sulfolobus_solfataricus_is_mediated_by_pili_formation.pdf.
- ↑ Brock biology of microorganisms (13th ed.). San Francisco: Benjamin Cummings. 2012. ISBN 9780321649638.
- ↑ Costerton, J. William (2007). "Direct Observations". The Biofilm Primer. Springer Series on Biofilms. 1. Berlin, Heidelberg: Springer. pp. 3–4. doi:10.1007/978-3-540-68022-2_2. ISBN 978-3-540-68021-5.
- ↑ "Microbial biofilms". Annual Review of Microbiology 49 (1): 711–45. October 1995. doi:10.1146/annurev.mi.49.100195.003431. PMID 8561477.
- ↑ "Thinking about bacterial populations as multicellular organisms". Annual Review of Microbiology 52 (1): 81–104. 1998. doi:10.1146/annurev.micro.52.1.81. PMID 9891794. http://www.sci.uidaho.edu/newton/math501/Sp05/Shapiro.pdf.
- ↑ "Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles". Nature Communications 5 (1): 4462. July 2014. doi:10.1038/ncomms5462. PMID 25042103. Bibcode: 2014NatCo...5.4462C.
- ↑ "The genetical evolution of social behaviour. II". Journal of Theoretical Biology 7 (1): 17–52. July 1964. doi:10.1016/0022-5193(64)90039-6. PMID 5875340. Bibcode: 1964JThBi...7...17H.
- ↑ "Introduction". Control of Biofilm Infections by Signal Manipulation. Springer Series on Biofilms. 2. Berlin, Heidelberg: Springer. 2008. pp. 1–11. doi:10.1007/7142_2007_006. ISBN 978-3-540-73852-7.
- ↑ "Bacterial biofilms: a common cause of persistent infections". Science 284 (5418): 1318–22. May 1999. doi:10.1126/science.284.5418.1318. PMID 10334980. Bibcode: 1999Sci...284.1318C. https://scholarworks.montana.edu/xmlui/handle/1/14368.
- ↑ "Extremophile". Encyclopedia of Earth. National Council of Science & the Environment. 2010. http://www.eoearth.org/article/Extremophile?topic=49540.
- ↑ Cobián Güemes, Ana Georgina; Youle, Merry; Cantú, Vito Adrian; Felts, Ben; Nulton, James; Rohwer, Forest (2016-09-29). "Viruses as Winners in the Game of Life". Annual Review of Virology (Annual Reviews) 3 (1): 197–214. doi:10.1146/annurev-virology-100114-054952. ISSN 2327-056X. PMID 27741409.
- ↑ "There must be a prokaryote somewhere: microbiology's search for itself". Microbiological Reviews 58 (1): 1–9. March 1994. doi:10.1128/MMBR.58.1.1-9.1994. PMID 8177167.
- ↑ "The prokaryote-eukaryote dichotomy: meanings and mythology". Microbiology and Molecular Biology Reviews 69 (2): 292–305. June 2005. doi:10.1128/MMBR.69.2.292-305.2005. PMID 15944457.
- ↑ Zhu, Qiyun; Mai, Uyen; Pfeiffer, Wayne; Janssen, Stefan; Asnicar, Francesco; Sanders, Jon G.; Belda-Ferre, Pedro; Al-Ghalith, Gabriel A. et al. (2019). "Phylogenomics of 10,575 genomes reveals evolutionary proximity between domains Bacteria and Archaea". Nature Communications 10 (1): 5477. doi:10.1038/s41467-019-13443-4. PMID 31792218. Bibcode: 2019NatCo..10.5477Z.
- ↑ "Primal eukaryogenesis: on the communal nature of precellular States, ancestral to modern life". Life 2 (1): 170–212. January 2012. doi:10.3390/life2010170. PMID 25382122. Bibcode: 2012Life....2..170E.
- ↑ "Origins. On the origin of eukaryotes". Science 325 (5941): 666–8. August 2009. doi:10.1126/science.325_666. PMID 19661396.
- ↑ "Ancient horizontal gene transfer". Nature Reviews. Genetics 4 (2): 121–32. February 2003. doi:10.1038/nrg1000. PMID 12560809.
- ↑ "Where is the root of the universal tree of life?". BioEssays 21 (10): 871–9. October 1999. doi:10.1002/(SICI)1521-1878(199910)21:10<871::AID-BIES10>3.0.CO;2-Q. PMID 10497338.
- ↑ "Early evolution: prokaryotes, the new kids on the block". BioEssays 21 (10): 880–9. October 1999. doi:10.1002/(SICI)1521-1878(199910)21:10<880::AID-BIES11>3.0.CO;2-P. PMID 10497339.
- ↑ "The universal ancestor". Proceedings of the National Academy of Sciences of the United States of America 95 (12): 6854–9. June 1998. doi:10.1073/pnas.95.12.6854. PMID 9618502. Bibcode: 1998PNAS...95.6854W.
- ↑ Martin, William (2005). "Woe is the Tree of Life". in Sapp, Jan. Microbial Phylogeny and Evolution: Concepts and Controversies. Oxford: Oxford University Press. pp. 139.
- ↑ Carl Woese, J Peter Gogarten, "When did eukaryotic cells (cells with nuclei and other internal organelles) first evolve? What do we know about how they evolved from earlier life-forms?" Scientific American, October 21, 1999.
- ↑ "Evidence for life in a martian meteorite?". GSA Today 7 (7): 1–7. July 1997. PMID 11541665.
- ↑ "Search for past life on Mars: possible relic biogenic activity in martian meteorite ALH84001". Science 273 (5277): 924–30. August 1996. doi:10.1126/science.273.5277.924. PMID 8688069. Bibcode: 1996Sci...273..924M.
- ↑ Crenson, Matt (2006-08-06). "After 10 years, few believe life on Mars". Associated Press (on space.com]). http://www.space.com/scienceastronomy/ap_060806_mars_rock.html.
- ↑ "Origin of carbonate-magnetite-sulfide assemblages in Martian meteorite ALH84001". Journal of Geophysical Research 104 (E2): 3803–13. February 1999. doi:10.1029/1998JE900034. PMID 11542931. Bibcode: 1999JGR...104.3803S.
- ↑ Bruce Alberts (2002). The Molecular Biology of the Cell (fourth ed.). Garland Science. pp. 808. ISBN 0-8153-3218-1. https://archive.org/details/molecularbiolog000wils.
- ↑ "The bacterial nucleoid: a highly organized and dynamic structure". Journal of Cellular Biochemistry 96 (3): 506–21. October 2005. doi:10.1002/jcb.20519. PMID 15988757.
- ↑ "Natural genetic transformation generates a population of merodiploids in Streptococcus pneumoniae". PLOS Genetics 9 (9): e1003819. 2013. doi:10.1371/journal.pgen.1003819. PMID 24086154.
- ↑ "Conservation and transformation of energy by bacterial membranes". Bacteriological Reviews 36 (2): 172–230. June 1972. doi:10.1128/MMBR.36.2.172-230.1972. PMID 4261111.
- ↑ "The bacterial cytoskeleton". Microbiology and Molecular Biology Reviews 70 (3): 729–54. September 2006. doi:10.1128/MMBR.00017-06. PMID 16959967.
- ↑ "Dynamic filaments of the bacterial cytoskeleton". Annual Review of Biochemistry 75 (1): 467–92. 2006. doi:10.1146/annurev.biochem.75.103004.142452. PMID 16756499. http://www2.mrc-lmb.cam.ac.uk/SS/Lowe_J/group/PDF/annrev2006.pdf.
- ↑ "Intracellular compartmentation in planctomycetes". Annual Review of Microbiology 59 (1): 299–328. 2005. doi:10.1146/annurev.micro.59.030804.121258. PMID 15910279.
- ↑ "Three-dimensional reconstruction of bacteria with a complex endomembrane system". PLOS Biology 11 (5): e1001565. 2013. doi:10.1371/journal.pbio.1001565. PMID 23700385.
- ↑ 56.0 56.1 56.2 "Major New Microbial Groups Expand Diversity and Alter our Understanding of the Tree of Life". Cell 172 (6): 1181–1197. March 2018. doi:10.1016/j.cell.2018.02.016. PMID 29522741. http://www.escholarship.org/uc/item/0299z0z4.
- ↑ Cavalier-Smith T (March 2002). "The phagotrophic origin of eukaryotes and phylogenetic classification of Protozoa". Int. J. Syst. Evol. Microbiol. 52 (Pt 2): 297–354. doi:10.1099/00207713-52-2-297. PMID 11931142. http://ijs.sgmjournals.org/cgi/pmidlookup?view=long&pmid=11931142.
- ↑ "DNA replication in the archaea". Microbiology and Molecular Biology Reviews 70 (4): 876–87. December 2006. doi:10.1128/MMBR.00029-06. PMID 17158702.
- ↑ The Vital Question – Energy, Evolution, and the Origins of Complex Life. W. W. Norton. 2015. p. 77. ISBN 978-0-393-08881-6. https://archive.org/details/vitalquestionene0000lane.
- ↑ Forterre, Patrick (2006). "Three RNA cells for ribosomal lineages and three DNA viruses to replicate their genomes: A hypothesis for the origin of cellular domain". PNAS 103 (10): 3669–3674. doi:10.1073/pnas.0510333103. PMID 16505372. Bibcode: 2006PNAS..103.3669F.
External links
- Prokaryote versus eukaryote, BioMineWiki
- The Taxonomic Outline of Bacteria and Archaea
- The Prokaryote-Eukaryote Dichotomy: Meanings and Mythology
- Quiz on prokaryote anatomy
- TOLWEB page on Eukaryote-Prokaryote phylogeny
![]() This article incorporates public domain material from the NCBI document "Science Primer".
This article incorporates public domain material from the NCBI document "Science Primer".
 |